Fig. 3.
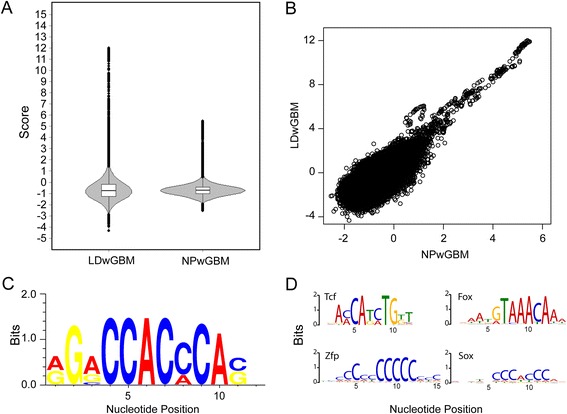
Assessment of genomic kmer-SVM predictions using classifiers trained on LDwGBM and NPwGBM datasets. a All genomic sequences matching the restricted 548 GBM 12-mers (wGMB) were identified and the 600 bp surrounding each GBM were assessed and scored using the kmer-SVM classifier that was trained on each of the two datasets. b Correlation plot depicting the relationship between LDwGBM and NPwGBM scores; scores >1 are highly correlated in the two datasets. c GLI motif generated from overlapping high weighted k-mers shared between LDwGBM and NPwGBM classifiers. d High weighted k-mers (identified by Tomtom) represented in either LDwGBM (Tcf and Zfp) or NPwGBM (Fox and Sox)
