Fig. 1.
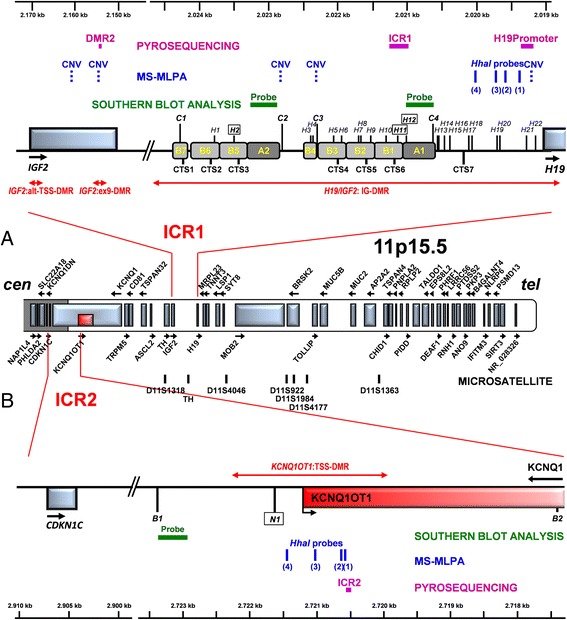
Schematic of the 11p15 imprinted region. ICR1 comprising H19/IGF2:IG-DMR, organized into the two CTS1-3 and CTS 4–6 clusters, plus the proximal CTS7, IGF2:ex9-DMR, and IGF2:alt-TSS-DMR (a) and ICR2 comprising KCNQ1OT1:TSS-DMR (b) are depicted. Red lines ending with arrowheads point to the DMRs. The HpaII sites, widely distributed across H19/IGF2:IG-DMR, are indicated by the vertical H bars; letters in the boxes represent CpG sites analyzed by SB. The Csp6I (a), BamHI, and NotI (b) sites are indicated by the vertical C, B, and N bars, respectively. Green horizontal bars indicate the probes for SB, blue vertical bars indicate the MLPA probes (dashed for CNVs and solid for methylation status), and solid purple bars indicate the pyrosequencing target regions [12]. The numbers in brackets of MLPA methylation probes refer to those indicated with the relative code in Fig. 2f, g
