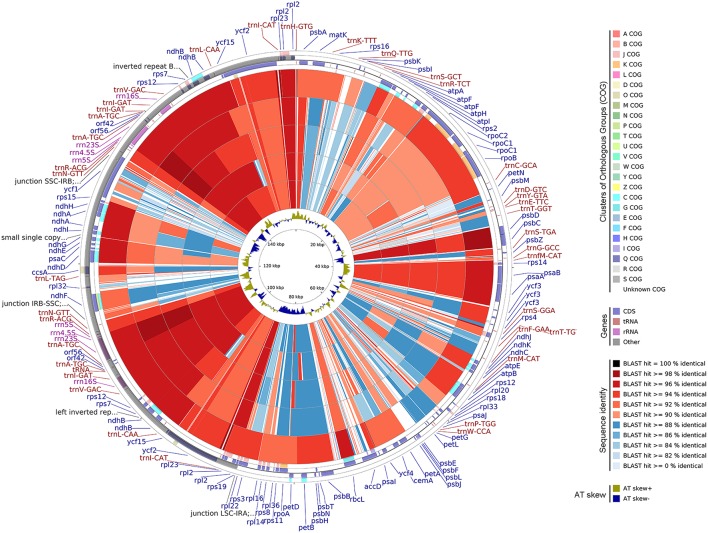
Figure 6.
Genome comparison of five CP genomes of Malvales to A.sinensis. From the outer to the inner color ring: Gonystylus bancanus, Theobroma cacao, Gossypium longicalyx, Hibiscus syriacus, and Gossypium bickii. BLAST was used to align other genomes to A. sinensis, and the results are shown with a circular map. The color codes are based on the similarity score, that is, dark red and blue depict similarity scores of 100%, above 90%, and below 90%, respectively. The four outer narrow rings are the protein-coding gene positions based on the A. sinensis cp genome. The color codes are based on Clusters of Orthologous Groups. The innermost ring is AT skew in the A. sinensis. AT skew+ indicate A>T, AT skew- indicate A < T.