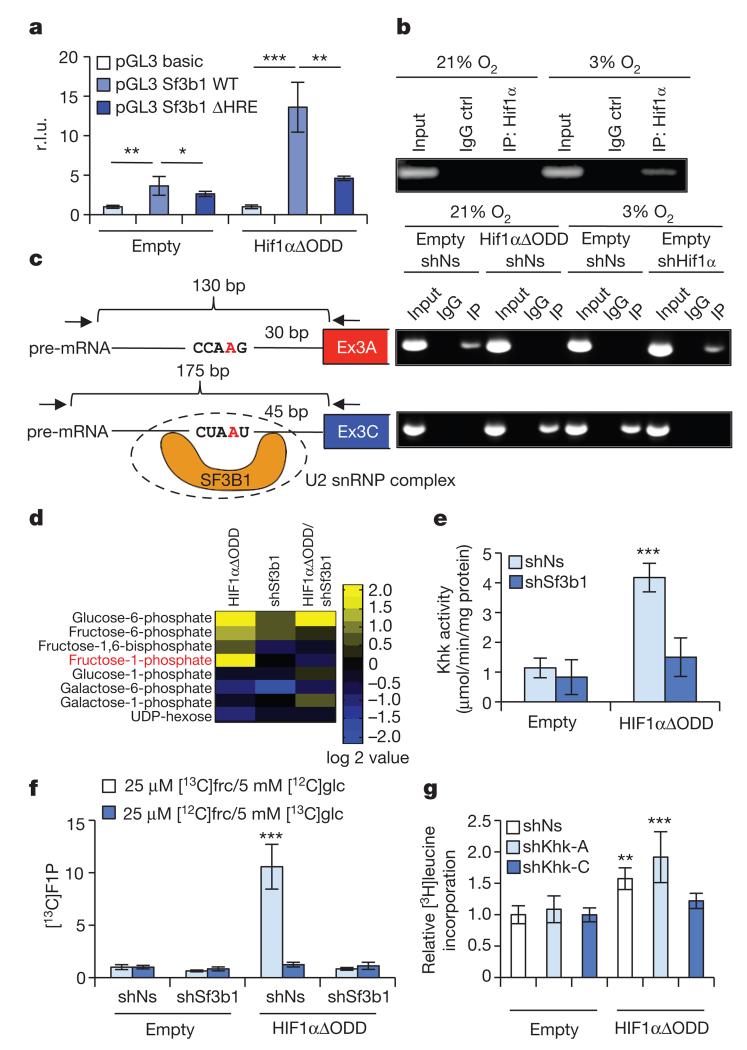
Figure 2. SF3B1 is a HIF1α-target gene that mediates splicing of KHK.
a, Co-transfection of wild type (WT) or HRE-mutated SF3B1 promoters fused to luciferase with either an empty vector control or HIF1αΔODD (values are relative light units (r.l.u.); n = 4 biological replicates per group). b, NMCs were cultured as indicated and processed for chromatin immunoprecipitation with a HIF1α-specific antibody (IP: HIF1α) or with a control isotype-matched antibody (IgG control). c, Schematic representing branch-point sequences located in front of exon 3A (upper left) or exon 3C (lower left). Branch point adenine is labelled in red. Location and orientation of primers (arrows), expected length of fragments to be amplified by RT–PCR and potential Sf3b1 binding sites are depicted. NMCs treated as indicated were processed for RNA-immunoprecipitation assay with antibodies specific for Sf3b1 (IP) or control-isotype matched antibody (IgG). Recovered RNA was purified and analysed by RT–PCR (upper right and lower right). d, Mass spectrometry-based metabolomic profiling of NMCs transduced with lentiviruses as denoted focused on hexose-phosphates. Metabolite values are compared to amounts in control transduced NMCs (n = 4 biological replicates per group). e, KHK activity in lysates of NMCs transduced with lentiviruses as indicated (n = 7 biological replicates per group). f, Relative amount of [13C]fructose-1-phosphate derived from [13C]fructose ([13C]frc) or [13C]glucose ([13C]glc) in NMCs transduced as denoted (n = 4 biological replicates per group). g, [3H]leucine incorporation in NMCs infected as indicated (n = 3 biological replicates per group, data show 1 of 3 representative experiments). For panels a, e–g, *P < 0.05; **P < 0.01; ***P < 0.001. Two-tailed unpaired t-test (a) one-way ANOVA followed by Dunnett’s multiple comparison post-test (e–g). Error bars are s.d. (a, g) or s.e.m. (e, f).