Abstract
In addition to their canonical roles in the cell cycle, RB family proteins regulate numerous developmental pathways, although the mechanisms remain obscure. We found that Drosophila Rbf1 associates with genes encoding components of the highly conserved apical–basal and planar cell polarity pathways, suggesting a possible regulatory role. Here, we show that depletion of Rbf1 in Drosophila tissues is indeed associated with polarity defects in the wing and eye. Key polarity genes aPKC, par6, vang, pk, and fmi are upregulated, and an aPKC mutation suppresses the Rbf1-induced phenotypes. RB control of cell polarity may be an evolutionarily conserved function, with important implications in cancer metastasis.
The Retinoblastoma tumor suppressor protein (RB) is a conserved central regulator of eukaryotic cell cycle, a function that has been extensively documented1 (and references therein). RB restricts entry of cells into S phase through its activity as a corepressor of the E2F family of transcription factors. RB activity is functionally downregulated during cell cycle progression by G1 Cyclin/CDKs, whose activities result in extensive phosphorylation and allosteric changes that inactivate RB2. Steady-state levels of RB family members, including the p107 and p130 homologs, are additionally regulated at the level of protein stability via the ubiquitin-proteosome pathway3. In addition to well-characterized roles in cell cycle progression, RB family proteins play key roles in differentiation, apoptosis, genome stability, and senescence, although it is less well understood how RB family proteins influence these additional biological pathways4,5,6.
Essential features of RB regulation are recapitulated in the Drosophila system, which features two members, Rbf1 and Rbf2, providing a streamlined model to interrogate Rbf-E2F roles in developmental programs7. In genome-wide studies, we found that Rbf1 occupies a number of promoters for key genes linked to conserved cellular signaling pathways, including the insulin, Hippo, Jak /Stat and polarity pathways, in addition to canonical cell cycle genes8. Thus, the impact of RB proteins on diverse developmental processes may in part reflect evolutionarily conserved transcriptional regulation of signaling components. Indeed, Rbf1 and the Hippo tumor suppressor pathway are functionally linked during photoreceptor differentiation. In genetic tests, strong synergy was noted between rbf1 and the Hippo pathway kinase warts, resulting in increased proliferation, suppression of apoptosis and a failure of photoreceptor differentiation9. An RB-Hippo link has also been noted in mammals; partial knockdown of LATS2 compromises phosphorylated RB’s ability to suppress cell proliferation and induction of senescence in Saos2 cells, and triple knockout liver cells lacking RB and its homologs p107 and p130 showed elevated expression of the Hippo pathway transcriptional effector YAP, as well as YAP target genes10,11. Molecular studies have also linked RB to the insulin and mTOR signaling pathways12,13. Collectively, these studies suggest that RB plays a pivotal and evolutionarily conserved role in regulation of core signaling genes with pleiotropic roles in cell growth.
A large proportion of promoters for polarity and planar cell polarity (PCP) genes are bound by Rbf1, much more than would be expected by chance (Chi square test P < 0.001), raising the question of whether the corepressor might regulate polarity pathways14,15. Apical/basal polarity, a characteristic of all epithelial cells, allows for specialized functions such as directional transport of ions and regulation of asymmetric cell divisions. Proper establishment of apical/basal polarity is key to maintaining normal tissue architecture and tissue homeostasis, and provides a tumor suppressive function as a block against malignant invasion16. In addition, some epithelial cells display planar cell polarity, in which the vector for polarization is orthogonal to the epithelial plane. At a macroscopic level, the regimented orientation of scales in fish, feathers in birds and hairs in mammals are examples of PCP17. Much of our understanding of the genetics of this evolutionarily conserved process originates from studies in Drosophila, where PCP is visualized as distally pointing hairs on the wing, posterior pointing hairs on the abdomen and in the stereotypic orientation of ommatidia in the eye. Polarity genes show dynamic stage- and tissue-specific regulation, however little is known about mechanisms controlling transcription levels of these genes. Here, we report evidence for direct functional links between Rbf1 and conserved polarity genes, providing a novel molecular connection between regulation of cell cycle and cell polarity.
Results
Rbf1 depletion induces strong polarity defects in eyes and notum
The establishment of polarity in different tissue types commonly involves several core genes, including fz, dsh, pk, vang, dgo and fmi. To understand the biological relevance of Rbf1 interaction with cell polarity gene promoters, we tested whether depletion of Rbf1 phenocopies polarity defects, such as those observed in fz, vang, and other core polarity mutants18. In flies, polarity can be visualized in tissues of epithelial origin such as the eye, wing, and thorax. In the eye, planar polarity is apparent in the orderly arrangement of the ommatidia in the dorsal/ventral axis. Misrotation and loss of photoreceptor cells within the ommatidia were observed upon loss or gain of function of apical/basal polarity determinants19. We used the Gal4-UAS system to deplete Rbf1 in the eye and assayed for polarity defects. Analysis of tangential sections revealed that Rbf1 RNAi generated two phenotypes that showed disruption of photoreceptor morphology, and was apparent in all the eyes analyzed (Fig. 1A). First, in ommatidia with the full complement of photoreceptor cells, a specific polarity phenotype was apparent, whereby the eight cells were canonically disposed, but their rotation and chirality was random, similar to the phenotype of PCP mutants20. These rotation defects may reflect Rbf1 regulation of core polarity genes, and/or regulation of another Rbf1 target gene canoe, which is also physically bound by Rbf1. canoe is an effector of the EGFR pathway, important for regulating ommatidial rotation in the eye21,22. Second, a significant number of ommatidia showed fewer than eight photoreceptors. Consistent with this observation, earlier studies also found that rbf1 clones display disorganized ommatidia lacking one or more photoreceptors and cone cell defects23. The lack of photoreceptor cells may represent Rbf1 acting through polarity determinants as well as through control of genes involved cell differentiation and proliferation. Similar results were obtained with two different UASRbf1 RNAi lines, with stronger phenotypes observed using the Bloomington line shown in Fig. 1A. We observed similar phenotypes with ey-Gal4 and GMR-Gal4; combining the ey-Gal4 and GMR-Gal4 drivers produced a more severe phenotype (data not shown). We did not note polarity defects in eyes of rbf 120/14Δ heterozygotes, presumably because this reduction is less severe than in the RNAi background.
Figure 1. Depletion of Rbf1 induces polarity defects in eye and notum.
(A) Tangential ommatidial sections were analyzed for orientation and number of photoreceptor cells in individual ommatidia. Control eyes showed normal number and dorsal/ventral orientation of cell clusters (duplicate eye section marked below original image; red trapezoids show dorsal and blue show ventral orientations). Rbf1 RNAi (center) and Rbf1 RNAi/UAS p35 (right) displayed similar phenotypes; ommatidia exhibited randomly oriented groups of photoreceptor cells (red), as well as many ommatidia lacking the full complement of cells (yellow). A few ommatidia with full complement of photoreceptor cells showed a symmetrical rather than trapezoidal arrangement (green). Anterior is to the right. 400 to 1000 ommatidia were analyzed for polarity defects in multiple sections from four to ten individuals. No equatorial midline could be defined either in Rbf1 RNAi or in Rbf1 RNAi/UAS p35 sections. (B) Bristle orientation defects seen on notum of Rbf1 RNAi individual (right panel). Anterior is to the top. ey-Gal4 was used for expression in the eye, and pen-Gal4 for expression in the notum. Similar but generally weaker phenotypes were observed with the GMR-Gal4 driver. The results shown were generated using Rbf1 RNAi line obtained from the Bloomington Stock Center; similar but overall weaker phenotypes were observed with a line obtained from VDRC (data not shown).
During eye development, in early third instar larval stages, the actual determination and specification of photoreceptor cells begin in the morphogenetic furrow (MF); loss of Rbf1 results in a pattern of cell death visualized in the morphogenetic furrow24,25. In order to understand if Rbf1 RNAi phenotypes in the eye are secondary to apoptosis, we coexpressed the p35 apoptotic inhibitor along with Rbf1 RNAi and analyzed polarity defects. p35 expression alone did not affect ommatidial architecture; the eyes showed mostly normal architecture, with a few ommatidia showing loss of photoreceptors (less than 5%, data not shown). However, we did not observe suppression of polarity defects when Rbf1 RNAi was carried out in this background. We observed similar loss of photoreceptors (84% of ommatidia affected) and of the ommatidia with full complement of photoreceptor cells, all showed orientation defects, suggesting that Rbf1 induced polarity defects are independent of apoptosis (Fig. 1A). We also depleted Rbf1 levels in the presumptive notum using the Pen-Gal4 driver. In these flies, defects were observed in the thoracic bristle pattern, whereby the normal anterior-posterior bristle orientation was completely disrupted on the notum, similar to the phenotype previously described for fz and fmi mutants26 (Fig. 1B).
Rbf1 depletion-induced polarity defects in Drosophila wings
There are significant tissue-specific differences in the requirement for some polarity genes, as well as in the specific downstream effectors needed for establishing polarity. In the wing, polarity is established largely independently of ft and ds, whereas these genes are essential for polarity in the eye27. Additional differences involve downstream effectors of polarity proteins; mwh, fy and in serve as downstream effectors in the wing whereas Delta serves as a downstream effector of fz in the eye, inducing Notch signaling in neighboring cells28,29. We therefore analyzed the effects of Rbf1 knockdown in developing wings.
Rbf1 RNAi resulted in subtle but reproducible wing hair polarity defects in all flies, manifested by the loss of highly stereotypic proximal–distal orientation of hairs in several sectors of the adult structure (Fig. 2A,B). Unlike the pleiotropic effects on all wing hairs seen in a fz null background, here we observed multiple, isolated sectors of disrupted polarity on affected wings.
Figure 2. Rbf1 RNAi causes polarity defects in Drosophila wings.

(A) Schematic representation of regions of adult wing as noted in Wong and Adler18. (B) Bristle orientation. Representative regions of wings showing orientation of wing hairs. In contrast to the normal proximal-distal orientation in control flies (left panels), Rbf1 RNAi expressing flies showed misorientation of wing hairs (right panels). The wings are oriented distal to the right, anterior to the top. (C) Ridge assay. To measure longitudinal ridges formed by edges of closely packed hexagonal cells, wild-type and Rbf1 RNAi wings were imaged as described32. Anterior regions (a) show normal A-P ridge orientation, while posterior regions (d,e) show P-D orientation in control wings (left panels). No clear ridge formation is apparent in Rbf1 RNAi expressing flies (right panels). Inserts in lower panels show magnified images to illustrate change in cell shape observed in Rbf1 RNAi wing. More than one hundred wings were analyzed for polarity and ridge defects. pen > Gal4 was used for expression in the wing. The results shown were generated using Rbf1 RNAi line obtained from the Bloomington Stock Center. Similar phenotypes were observed with a Bx > Gal4 driver; effects were restricted to posterior wing compartment using en-Gal4.
We additionally analyzed the orientation of adult wing cuticular ridges, prominent aligned edges of cells that extend across the wing. The formation of ridges reflects polarized cellular organization. The development of ridges requires planar cell polarization and hexagonal cell packing, and as with hair polarity, this process is regulated by the PCP signaling pathway, although with some differences30,31,32. Ridges are oriented anterior-posterior in the wing anterior, and proximal-distal in the wing posterior regions. Rbf1 RNAi wings showed greatly reduced ridge structure across the entire surface of the wing, both in the anterior and posterior regions (Fig. 2C). This phenotype is similar to the defect observed in pk mutants32. In addition, cells in Rbf1 RNAi wings exhibited a “cobblestone” pattern indicating irregular packing, a process which is influenced by PCP components31. To mitigate RNAi off target effects, we tested the two independent RNAi lines with various wing specific drivers for polarity defects. We observed similar hair orientation and cuticular defects using an alternative UASRbf1 RNAi line as well as alternative Gal4 drivers (en-Gal4 and Beadex-Gal4; data not shown).
Rbf1 represses transcription of key polarity genes in wing imaginal discs
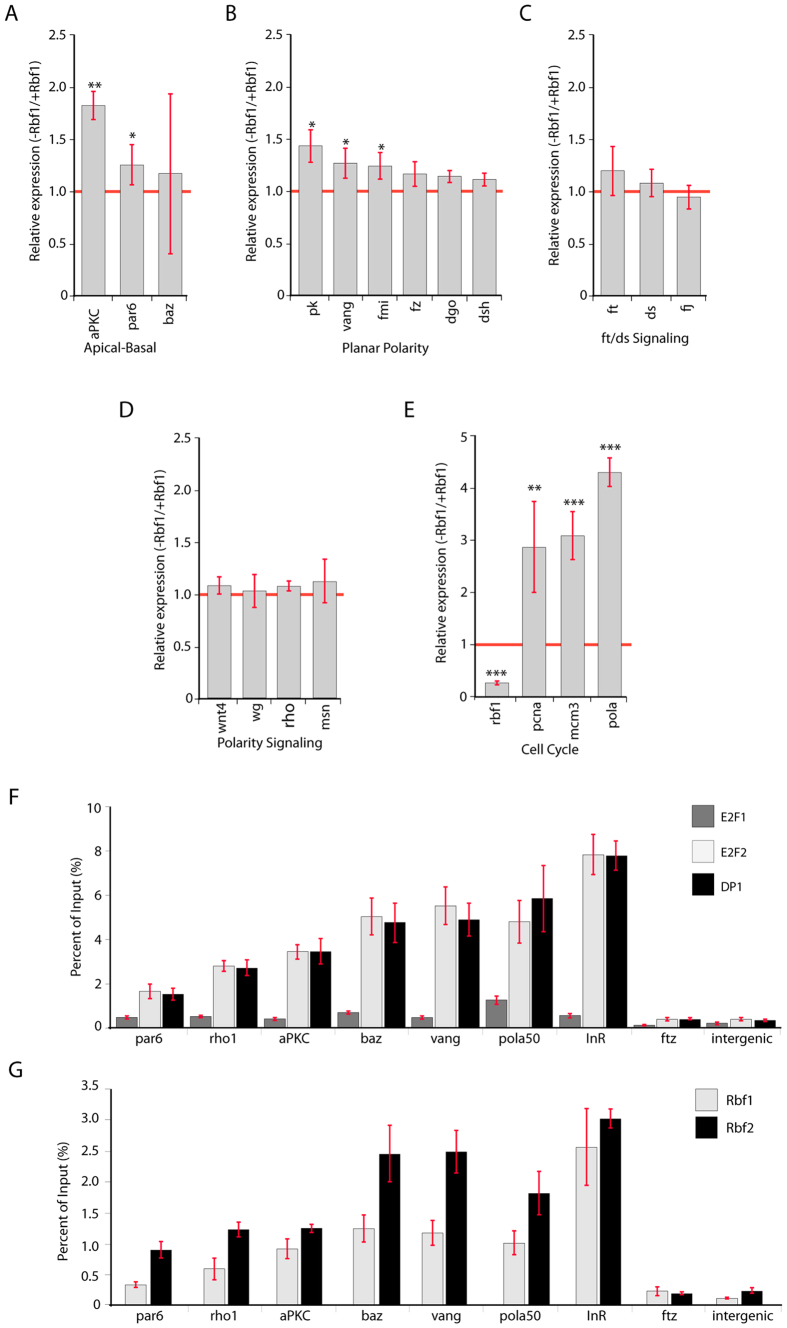
Planar polarity observed in the adult wing is established in the third instar disc33. To test whether the observed phenotypes reflect direct transcriptional regulation of key polarity genes by Rbf1, we depleted Rbf1 in third instar larval wing imaginal discs using RNAi and measured the expression levels of target genes. Members of core polarity genes demonstrated significant upregulation. The apical polarity determinant aPKC was upregulated 1.8 fold, and par6, which encodes a protein that forms a complex with aPKC, showed a smaller but reproducible increase (Fig. 3A). We were unable to determine if baz, encoding another aPKC complex member, showed a significant change in transcript levels. Half of the genes of fz planar polarity pathway (vang, pk and fmi) showed more modest but still highly reproducible upregulation (Fig. 3B). We did not observe significant changes in the expression of fz, dsh, dgo, or members of the ft/ds pathway (Fig. 3B,C). Transcript levels of additional genes associated with cell polarity signaling (wnt4, wg, rho1, msn) were not changed by this treatment (Fig. 3D). Thus, all genes with significant changes in expression showed upregulation, consistent with the role of Rbf1 as a transcriptional repressor. Three of the five upregulated genes (aPKC, vang, and par-6) were previously found to be bound by Rbf1 in our ChIP-seq data, however regulation was not entirely predicted by binding, as three core polarity genes (fz, dsh, ft) were not upregulated in this context. As controls, we measured transcript levels of the rbf1 gene itself and cell cycle genes regulated by Rbf1; RNAi significantly depleted levels of rbf1, and we observed corresponding upregulation of pcna, mcm3, and DNApol-α50 (Fig. 3E).
Figure 3. Alterations in Rbf1 levels affect the expression of apical/basal and planar polarity genes.
(A) Transcript levels of apical/basal polarity genes, from wing imaginal discs that were depleted for Rbf1 by RNAi. aPKC and par6 showed significant upregulation in the RNAi background. (B) Planar polarity genes; pk, vang and fmi were modestly but significantly upregulated, while other genes were not significantly affected. (C) ft/ds pathway genes were not significantly affected (D) Expression of polarity-interacting genes was similarly unaffected by Rbf1 depletion. (E) rbf1 mRNA levels were significantly depleted by RNAi treatment, and Rbf1-targeted cell cycle genes pcna, mcm5, and DNA polα-50 were upregulated. Red line indicates control expression (Gal4-RNAi). Values for RNAi represent average of five biological replicates (*P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001) and error bars represent standard deviations. (F) Interaction of E2F1, E2F2, and DP1 transcription factors with promoters of polarity-determining genes assayed by chromatin immunoprecipitation. E2F2 and DP1 were strongly enriched on aPKC, vang, baz, and rho1, as well as on positive controls (inr and pol alpha 50), but not on negative controls (ftz and intergenic locus on chromosome 3). (G) Rbf2 is also associated with Rbf1 bound promoters on polarity regulating genes. Error bars represent standard deviation. Chromatin was prepared from 12–18 hour embryos as described8.
RB family proteins interact with target promoters through a variety of mechanisms, chiefly but not exclusively via interaction with E2F transcription factor family members34. Some targeted promoters appear to involve additional proteins, such as those of the dREAM complex, or in certain cases non E2F proteins35,36. Previous ChIP-chip data on whole larvae indicated that aPKC, vang, par 6, baz, and rho1 are bound by Rbf1 and/or E2F215. We had previously noted that Rb targeting tends to be conserved across developmental stages. To provide further information on possible cooccupancy of Rbf and E2F factors on PCP genes, we performed ChIP analysis for these promoters on Rbf1, Rbf2, E2F1, E2F2, and the E2F cofactor DP1. We observed significant enrichment of E2F2 and DP1, but not E2F1, on all of these genes, but not on nonspecific loci (Fig. 3F). We also observed that the Rbf2 protein was also consistently associated with these promoters (Fig. 3G); this protein has a secondary role in regulation of many promoters, hence Rbf1 depletion may be sufficient to affect transcription of the target genes we observed. Here, we used embryonic chromatin; however, binding properties of these proteins appears to be similar in embryos and larvae15. We conclude that Rbf1 interaction with the regulated polarity gene promoters uses a canonical E2F-DP1 targeting mechanism, and the regulation of the promoters may in fact resemble that of other signaling pathway genes such as InR36. Another similarity of protein complexes observed on these polarity gene promoters is the presence of the dREAM complex, a set of DNA-binding factors often associated with RB-bound loci; ChIP-chip data indicates that aPKC, vang, and par- 6 are associated with dREAM components in Kc cells35.
Genetic interaction between Rbf1 and aPKC
Our data suggests that misexpression of core polarity-regulating genes underlies the polarity defects found in the Rbf1 depleted flies. The largest effect was on the expression of aPKC; in the eye, this apical determinant regulates the PCP pathway. Previous work has indicated that upregulation of aPKC produces polarity defects: a constitutively active form of aPKC resulted in misrotated ommatidia and in loss of photoreceptors19. To test whether the upregulation of aPKC alone was important for the observed phenotypes, we reduced the dose Rbf1 using RNAi in an aPKC heterozygous background to test for genetic suppression. In this background, we found an increased number of ommatidia containing the complete complement of photoreceptors (68%). Most ommatidia displayed proper orientation with respect to their A/P and D/V planes with less than 5% percent showing orientation defects and only 28% showing loss of photoreceptors (Fig. 4). The ommatidia of flies that were heterozygous for aPKC alone had the normal number and orientation of photoreceptors, with very few ommatidia showing missing photoreceptors. As before, Rbf1 RNAi animals showed extensive loss of photoreceptor cells (82%) and rotational defects (18%). The genetic suppression suggests that much, if not all, of the effects observed here are influenced by misregulation of aPKC. The possible regulation of other genes by Rbf1 may be more important in other developmental settings, or play a modulatory role. We tried to rescue wing polarity defects in a similar manner by generating a combined aPKC heterozygote and Rbf1 RNAi using the pen-Gal4 driver, however this combination was synthetic lethal. We speculate that the lethality may be a consequence of disruption of neuroblast development, as both aPKC and Rbf1 are functional in this cell type, and the pen promoter is active in neuroblasts as well as wing imaginal disc cells37,38,39. Therefore we carried out Rbf1 RNAi using engrailed-Gal4 or Beadex-Gal4 drivers and observed strong wing polarity defects particularly with the Beadex driver. The aPKC heterozygous background did not rescue these polarity defects, however (data not shown). Misregulation of other polarity genes may play a more important role in this context. Indeed, tissue specific roles of polarity related genes have been previously observed: inturned and fuzzy are important for wing polarity but not for eye polarity while nemo is a key player in eye but not wing development40,41.
Figure 4. Genetic interaction between rbf1 and aPKC in eye development.
(A,A′) Section showing imaginal discs of wild-type eye, with markings to indicate dorsal and ventral polarity of photoreceptor cells (red and blue, respectively). (B,B′) Rbf1 RNAi eye section showing ommatidia displaying random orientation (red), chirality defects (green), and ommatidia lacking photoreceptor cells (yellow). No equatorial midline could be defined in these sections. (C,C′) Eye section of aPKC heterozygote showing normal number and orientation of photoreceptors within many ommatidia. A few ommatidia with missing photoreceptors are shown in yellow. (D,D′) Eye section of Rbf1 RNAi/aPKC heterozygote showing partial rescue. In comparison to rbf1 heterozygous phenotype (panel B), a large fraction of ommatidia show normal number and correct orientation of photoreceptors with respect to the dorsal/ventral axis. Orientation shows anterior to the right. At least 400 ommatidia were analyzed for polarity defects in multiple sections from four individuals (six for the rescue).
Discussion
An important question raised by our study is whether Rbf1 directly coordinates coupling between cell cycle and cell polarity pathways. These two processes are coordinated at some level in diverse systems. For example, in Drosophila neuroblasts, asymmetric division of mother and daughter cells is regulated by the mitotic kinases Polo and Aurora, which are required for both smooth progression through the cell cycle as well as correct localization of aPKC and Numb polarity determinants42. Similarly, although RB proteins are not conserved in basal eukaryotes, polarity and cell cycle are nonetheless linked in fission yeast43. Coordination can involve simultaneous targeting by transcription regulators, as in Arabidopsis, where the JAG transcription factor coordinately regulates cell cycle and polarity genes in the developing meristem44. Our results indicate that Rbf1 is directly influencing the function of polarity factors by regulating their expression levels, providing a mechanism for coordinating the regulation of this pathway with the cell cycle. In this model, Rbf1 would respond to cellular signaling to release from E2F complexes upon the start of S phase in response to cyclin/Cdk phosphorylation. Simultaneous upregulation of cell polarity genes by inactivation of Rbf1 might be important for cells to adjust to an altered surface to volume ratio following cytokinesis. Alternatively, this change in polarity determinants may even be necessary for successful progress through the cell cycle, as observed in neuroblasts, where apkc plays a role in both proliferation and polarity38. A similar regulatory role for polarity-associated genes lgl, dlg, and scrib on proliferative control has been reported45.
Previous studies have indirectly linked mammalian RB to cell polarity; ear-specific deletion of RB 1 resulted in a cell polarity defect characterized by aberrant inner ear hair cell proliferation and disorganized cilia in the mouse46. A role of mammalian RB family proteins in regulating transcription of cell polarity gene promoters has not been described, however, so the mechanism underlying this phenotype has been unknown. We note that among the reported targets of RB and p130 in human lung fibroblasts are the proximal promoter regions of aPKCΙ, fzd2, prickle1, and celsr16. Furthermore, among the genes found to be upregulated in these cells upon depletion of RB were fzd2 and prickle1, although the authors did not comment on these findings, and the relevance of this regulation to apical/basal or planar polarity is yet unanswered6. Resolving the intertwined roles of RB family proteins on polarity and cell cycle genes will yield a fresh perspective on the mechanisms employed by the RB family proteins.
Our results indicate that transcriptional regulation of polarity-determining genes by RB proteins is an important and evolutionarily conserved process, likely of importance for development in many contexts. Interestingly, a recent report from Knudsen shows that loss of RB in ErbB2- expressing breast cancer cells lead to changes in expression of genes important for maintaining epithelial architecture; mislocaliztion of E-cad and Laminin proteins were also observed in mammary cancer cell cultures, indicative of polarization defects47. Regulation of polarity genes by E2F family proteins appears to be another facet of this conserved regulatory mechanism: our analysis of published studies involving perturbations of E2F levels in Drosophila, C. elegans and human cells indicates that expression of PCP genes is affected48,49,50. Consistent with this model, null mutants for dDP, the partner of E2F proteins, exhibit dorsal-ventral polarity defects in egg chambers51. This subject will be a fruitful area for further inquiry, as disruption of RB-regulated polarity in cancer cells would potentially play an important role in development of metastasis.
Methods
Fly stocks and genetics
The aPKCk06403, UASRbf1 RNAi (Transgenic RNAi Project [TRiP]: HMS03004, Control dsRNA (TRiP: valium 20-Gal4.1) fly lines and pen > Gal4 (NP6333), Bx > Gal4 (stock number 8860), and en > Gal4 (stock number 30564) stocks were obtained from the Bloomington Stock Center (Bloomington, Indiana). A description of the alleles can be found in Flybase. A second UASRbf1 RNAi line was obtained from the Vienna Drosophila Resource Center (VDRC ID 10696). The ey-Gal4, GMR-Gal4, ey-Gal4; GMR-Gal4 and UAS-p35 stocks were kindly provided by Helen McNeill. Males from UASRbf1 RNAi were crossed to pen > Gal4 or ey > Gal4 and the progeny were analyzed for PCP defects. More than 100 wings were analyzed for wing defects and for ommatidial rotation defects n > 400 ommatidia were analyzed. Cuticle refraction microscopy was performed to obtain images of wing membrane topography32. Adult eye sections were prepared as described in Wolff, 200052. A minimum of 4 individuals were sectioned from each genotype and analyzed for ommatidial defects. For the genetic suppression studies, we generated flies of the genotype aPKCk06403/Gla-Bc; UASRbf1 RNAi or Control Gal4.1/TM6 Tb1 and crossed them to homozygous pen > Gal4 or en > Gal4/SM2Cyo or Bx > Gal4 or ey > Gal4/SM2 Cyo flies. aPKCk06403/pen > Gal4; UASRbf1 RNAi/TM6 Tb1 was lethal, precluding analysis of adult wing, however we were able to recover aPKCk06403/ey > Gal4; UASRbf1 RNAi/TM6 Tb1 animals, which were analyzed for ommatidial rescue. Five to ten individuals from at least two completely separate genetic crosses and more than 400 ommatidia were analyzed for all experiments. We analyzed two individuals from GMR-Gal4 > UASRbf1 RNAi (RNAi line obtained from Bloomington) and counted greater than 300 ommatidia for polarity defects.
Quantitative PCR
Wing imaginal discs were dissected from third-instar larvae, and total RNA was isolated according to the protocol53 using TRIzol (Invitrogen) followed by RNeasy Mini kit (Qiagen) for cleanup. 300 ng of total RNA was converted to cDNA using High Capacity cDNA Reverse Transcription kit (Applied Biosystems). The resulting cDNA was diluted 1:15 and 3 μl was used for PCR in a 20 μl reaction mixture using SYBR green PCR Master Mix (Applied Biosystems). qPCR was performed on five biological replicates and the fold change in gene expression was calculated based on the 2−ΔΔCT method and normalized to rp49 gene expression. Sequences for primers used for gene expression analysis are available upon request.
Chromatin Immunoprecipitation
Chromatin was prepared from 12–18 hour embryos following the protocol described in Acharya et al. 20128. Immunoprecipitations were carried out using Rbf1 and Rbf2 antibodies43. E2F and DP antibodies were gifts from Dr. Nicholas Dyson (Harvard Medical School). qPCR using SYBR green PCR Master Mix (Applied Biosystems) was performed on two biological replicates, and results in Fig. 3F,G show averages of these experiments. Sequences for primers used for PCR were situated under predicted Rbf1 binding areas in promoter proximal regions, except for negative control genes, and are available upon request. The intergenic negative control represents region on Drosophila chromosome 3L (chr 3L: 13736222–13736322).
Statistical Analysis
Data represents mean ± SD. P values were determined by unpaired Student’s t test.
Additional Information
How to cite this article: Payankaulam, S. et al. Regulation of cell polarity determinants by the Retinoblastoma tumor suppressor protein. Sci. Rep. 6, 22879; doi: 10.1038/srep22879 (2016).
Acknowledgments
We thank the Bloomington Stock Center (supported by NIH grant P40OD018537) for Gal4 and RNAi lines. We thank Dr. Nicholas Dyson for the E2F and DP antibodies, Dr. Tony J. C. Harris for providing the aPKC mutant flies, Y. Wei for embryonic chromatin, and A. Gjidoda for assistance in analyzing ChIP-Seq data. This work is supported by NIH GM079098 to RWH and DNA, GM056976 to DNA, and MOP-102656 and MOP-97933 to H.M.
Footnotes
Author Contributions The project was conceived by S.P., D.N.A. and S.P. analyzed wings for polarity defects, measured the transcript levels and performed genetic interaction assay. K.Y. performed sectioning of fly eyes and analyzed ommatidial defects. R.W.H. and S.P. prepared the figures. The manuscript written by S.P., D.N.A., H.M. and R.W.H. All authors reviewed the manuscript.
References
- Burkhart D. L. & Sage J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat. Rev. Cancer 8, 671–682 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mittnacht S. Control of pRB phosphorylation. Curr Opin Genet Dev. 8, 21–27 (1998). [DOI] [PubMed] [Google Scholar]
- Sengupta S. et al. The Evolutionarily Conserved C-terminal Domains in the Mammalian Retinoblastoma Tumor Suppressor Family Serve as Dual Regulators of Protein Stability and Transcriptional Potency. J. Biol. Chem. 290, 14462–14475 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lipinski M. M. & Jacks T. The retinoblastoma gene family in differentiation and development. Oncogene 18, 7873–7882 (1999). [DOI] [PubMed] [Google Scholar]
- Zheng L. & Lee W. H. Retinoblastoma tumor suppressor and genome stability. Adv. Cancer Res. 85, 13–50 (2002). [DOI] [PubMed] [Google Scholar]
- Chicas A. et al. Dissecting the unique role of the retinoblastoma tumor suppressor during cellular senescence. Cancer Cell 17, 376–387 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du W., Vidal M., Xie J. E. & Dyson N. RBF, a novel RB-related gene that regulates E2F activity and interacts with cyclin E in Drosophila. Genes Dev. 10, 1206–1218 (1996). [DOI] [PubMed] [Google Scholar]
- Acharya P. et al. Evidence for autoregulation and cell signaling pathway regulation from genome-wide binding of the Drosophila retinoblastoma protein. G3 (Bethesda). 2, 1459–1472 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicolay B. N., Bayarmagnai B., Moon N. S., Benevolenskaya E. V. & Frolov M. V. Combined inactivation of pRB and hippo pathways induces dedifferentiation in the Drosophila retina. PLoS Genet. 6, e1000918 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehmer U. et al. Organ size control is dominant over Rb family inactivation to restrict proliferation in vivo. Cell Rep. 8, 371–381 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tschöp K. et al. A kinase shRNA screen links LATS2 and the pRB tumor suppressor. Genes Dev. 25, 814–830 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Annicotte J.-S. et al. The CDK4-pRB-E2F1 pathway controls insulin secretion. Nat. Cell Biol. 11, 1017–1023 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon G. M. & Du W. Targeting Rb inactivation in cancers by synthetic lethality. Am. J. Cancer Res. 1, 773–786 (2011). [PMC free article] [PubMed] [Google Scholar]
- Wei Y. et al. Genome-wide analysis of Drosophila Rbf2 protein highlights diversity of RB family targets and possible role in regulation of ribosome biosynthesis. G3 (Bethesda). 5, 1503–1515 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korenjak M., Anderssen E., Ramaswamy S., Whetstine J. R. & Dyson N. J. RBF binding to both canonical E2F targets and noncanonical targets depends on functional dE2F/dDP complexes. Mol. Cell. Biol. 32, 4375–4387 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Royer C. & Lu X. Epithelial cell polarity: a major gatekeeper against cancer? Cell Death Differ. 18, 1470–1477 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNeill H. Planar cell polarity: keeping hairs straight is not so simple. Cold Spring Harb. Perspect. Biol. 2, a003376 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong L. L. & Adler P. N. Tissue polarity genes of Drosophila regulate the subcellular location for prehair initiation in pupal wing cells. J. Cell Biol. 123, 209–221 (1993). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Djiane A., Yogev S. & Mlodzik M. The apical determinants aPKC and dPatj regulate Frizzled-dependent planar cell polarity in the Drosophila eye. Cell 121, 621–631 (2005). [DOI] [PubMed] [Google Scholar]
- Strutt H. & Strutt D. Polarity determination in the Drosophila eye. Curr. Opin. Genet. Dev. 9, 442–446 (1999). [DOI] [PubMed] [Google Scholar]
- Gaengel K. & Mlodzik M. Egfr signaling regulates ommatidial rotation and cell motility in the Drosophila eye via MAPK/Pnt signaling and the Ras effector Canoe/AF6. Development 130, 5413–5423 (2003). [DOI] [PubMed] [Google Scholar]
- Strutt H. & Strutt D. EGF signaling and ommatidial rotation in the Drosophila eye. Curr. Biol. 13, 1451–1457 (2003). [DOI] [PubMed] [Google Scholar]
- Popova M. K., He W., Korenjak M., Dyson N. J. & Moon N.-S. Rb deficiency during Drosophila eye development deregulates EMC, causing defects in the development of photoreceptors and cone cells. J. Cell Sci. 124, 4203–4212 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiung F. & Moses K. Retinal development in Drosophila: specifying the first neuron. Hum Mol Genet. 11, 1207–1214 (2002). [DOI] [PubMed] [Google Scholar]
- Moon N. S., Di Stefano L. & Dyson N. A Gradient of Epidermal Growth Factor Receptor Signaling Determines the Sensitivity of rbf1 Mutant Cells to E2F-Dependent Apoptosis. Molecular and Cellular Biology 26, 7601–7615 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu B., Usui T., Uemura T., Jan L. & Jan Y. N. Flamingo controls the planar polarity of sensory bristles and asymmetric division of sensory organ precursors in Drosophila. Curr. Biol. 9, 1247–1250 (1999). [DOI] [PubMed] [Google Scholar]
- Peng Y. & Axelrod J. D. Asymmetric protein localization in planar cell polarity: mechanisms, puzzles, and challenges. Curr. Top. Dev. Biol. 101, 33–53 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper M. T. & Bray S. J. Frizzled regulation of Notch signalling polarizes cell fate in the Drosophila eye. Nature 397, 526–530 (1999). [DOI] [PubMed] [Google Scholar]
- Lu Q., Yan J. & Adler P. N. The Drosophila planar polarity proteins inturned and multiple wing hairs interact physically and function together. Genetics 185, 549–558 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hogan J., Valentine M., Cox C., Doyle K. & Collier S. Two frizzled planar cell polarity signals in the Drosophila wing are differentially organized by the Fat/Dachsous pathway. PLoS Genet. 7, e1001305 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Classen A.-K., Anderson K. I., Marois E. & Eaton S. Hexagonal packing of Drosophila wing epithelial cells by the planar cell polarity pathway. Dev. Cell 9, 805–817 (2005). [DOI] [PubMed] [Google Scholar]
- Doyle K., Hogan J., Lester M. & Collier S. The Frizzled Planar Cell Polarity signaling pathway controls Drosophila wing topography. Dev. Biol. 317, 354–367 (2008). [DOI] [PubMed] [Google Scholar]
- Sagner A. et al. Establishment of global patterns of planar polarity during growth of the Drosophila wing epithelium. Curr. Biol. 22, 1296–1301 (2012). [DOI] [PubMed] [Google Scholar]
- Sellers W. R. et al. Stable binding to E2F is not required for the retinoblastoma protein to activate transcription, promote differentiation, and suppress tumor cell growth. Genes Dev. 12, 95–106 (1998). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georlette D. et al. Genomic profiling and expression studies reveal both positive and negative activities for the Drosophila Myb MuvB/dREAM complex in proliferating cells. Genes Dev. 21, 2880–2896 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raj N., Zhang L., Wei Y., Arnosti D. N. & Henry R. W. Ubiquitination of retinoblastoma family protein 1 potentiates gene-specific repression function. J. Biol. Chem. 287, 41835–41843 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kussel P. & Frasch M. Pendulin, a Drosophila protein with cell cycle-dependent nuclear localization, is required for normal cell proliferation. J. Cell Biol. 129, 1491–1507 (1995). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rolls M. M., Albertson R., Shih H.-P., Lee C.-Y. & Doe C. Q. Drosophila aPKC regulates cell polarity and cell proliferation in neuroblasts and epithelia. J. Cell Biol. 163, 1089–1098 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keller S. A., Ullah Z., Buckley M. S., Henry R. W. & Arnosti D. N. Distinct developmental expression of Drosophila retinoblastoma factors. Gene Expr. Patterns 5, 411–421 (2005). [DOI] [PubMed] [Google Scholar]
- Adler P. N., Charlton J. & Park W. J. The Drosophila tissue polarity gene inturned functions prior to wing hair morphogenesis in the regulation of hair polarity and number. Genetics 137, 829–836 (1994). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi K. W. & Benzer S. Rotation of photoreceptor clusters in the developing Drosophila eye requires the nemo gene. Cell 78, 125–136 (1994). [DOI] [PubMed] [Google Scholar]
- Noatynska A., Tavernier N., Gotta M. & Pintard L. Coordinating cell polarity and cell cycle progression: what can we learn from flies and worms? Open Biol. 3, 130083 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vaggi F. et al. Linkers of cell polarity and cell cycle regulation in the fission yeast protein interaction network. PLoS Comput. Biol. 8, e1002732 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiessl K., Muino J. M. & Sablowski R. Arabidopsis JAGGED links floral organ patterning to tissue growth by repressing Kip-related cell cycle inhibitors. Proc. Natl. Acad. Sci. USA 111, 2830–2835 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bilder D., Li M. & Perrimon N. Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science 289, 113–116 (2000). [DOI] [PubMed] [Google Scholar]
- Sage C. et al. Proliferation of functional hair cells in vivo in the absence of the retinoblastoma protein. Science 307, 1114–1118 (2005). [DOI] [PubMed] [Google Scholar]
- Witkiewicz A. K. et al. The retinoblastoma tumor suppressor pathway modulates the invasiveness of ErbB2-positive breast cancer. Oncogene 33, 3980–3991 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buttitta L. A., Katzaroff A. J. & Edgar B. A. A robust cell cycle control mechanism limits E2F-induced proliferation of terminally differentiated cells in vivo. JCell Biol. 189, 981–996 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chi W. & Reinke V. Promotion of oogenesis and embryogenesis in the C. elegans gonad by EFL-1/DPL-1 (E2F) does not require LIN-35 (pRB). Development 133, 3147–3157 (2006). Erratum in: Development 133, 3495 (2006). [DOI] [PubMed] [Google Scholar]
- Müller H. et al. E2Fs regulate the expression of genes involved in differentiation, development, proliferation, and apoptosis. Genes & Development 15, 267–285 (2001). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Myster D. L., Bonnette P. C. & Duronio R. J. A role for the DP subunit of the E2F transcription factor in axis determination during Drosophila oogenesis. Development 127, 3249–3261 (2000). [DOI] [PubMed] [Google Scholar]
- Wolff T. Histological techniques for the Drosophila eye part II: Adult, in Drosophila Protocols, Sullivan W., Ashburner M., Hawley R. S., Eds. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NewYork, 2000), chap. 13. [Google Scholar]
- Bin He. & Kreitman M. RNA extraction from small amount of samples (imaginal discs). OpenWetWare, 2012, 09/04/2012. http://openwetware.org/wiki/Kreitman:RNA_extraction_from_small_amount_of_samples_(imaginal_discs).