Fig. 1.
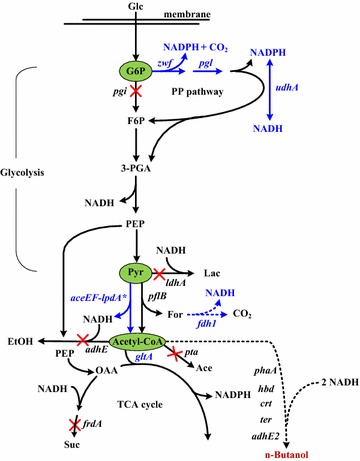
The central metabolic pathways leading to n-butanol in E. coli. The dotted lines denote the heterologous pathways. The CoA-dependent synthetic pathway of n-butanol is composed of heterologous phaA, hbd, crt, ter, and adhE2 genes as shown. Three metabolite nodes including G6P, pyruvate, and acetyl-CoA are targeted for engineering and marked. The genes involved in the metabolic pathways: aceEF-lpdA*, pyruvate dehydrogenase complex; adhE, aldehyde-alcohol dehydrogenase; adhE2, butyraldehyde-butanol dehydrogenase; crt, crotonese; gltA, citrate synthase; hbd, 3-hydroxybutyryl-CoA dehydrogenase; ldhA, lactate dehydrogenase; fdh1, formate dehydrogenase; frdA, subunit of fumarate reductase; pflB, pyruvate formate-lyase; pgi, phosphoglucose isomerase; pgl, lactonase; phaA, acetoacetyl-CoA thiolase; pta, phosphate acetyltransferase; ter, trans-enoyl-CoA reductase; udhA, transhydrogenase; zwf, glucose-6-phosphate dehydrogenase. The deleted genes are indicated by “X”. Abbreviations: Ace acetate; EtOH ethanol; F6P fructose-6-phosphate; Lac lactate; For formate; G6P glucose-6-phosphate; Glc glucose; OAA oxaloacetate; PEP phosphoenolpyruvate; 3-PGA 3-phosphoglyceraldehyde; Pyr pyruvate; Suc succinate
