Figure 2.
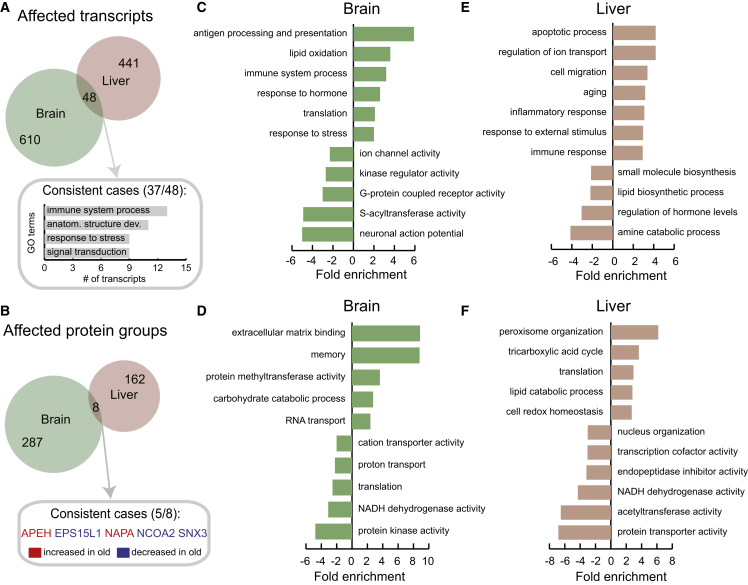
Common and Organ-Specific Alterations of the Proteome in Old Rats
(A and B) We identified only 48 transcripts affected at the level of translation output (A) and eight proteins affected at the level of protein abundance (B) that were altered in both brain and liver. The most represented gene ontology (GO) terms among consistently affected transcripts are shown in (A) while the five consistently affected proteins are indicated in (B).
(C–F) Functional enrichment was performed on the list of quantified transcripts and proteins that were ranked according to the level of differential expression (fold change) using GOrilla (Eden et al., 2009). Displayed GO terms are representative cases selected from among those significantly enriched (cut-off for transcripts: q value < 0.05, minimum number of transcript >4, fold enrichment ≥2; for proteins the same criteria were applied with the exception of q value < 0.2). The fold enrichments are plotted using positive values for terms enriched in transcripts/proteins that are increased in old animals, or using negative values for terms enriched in transcripts/proteins that are decreased in old animals.