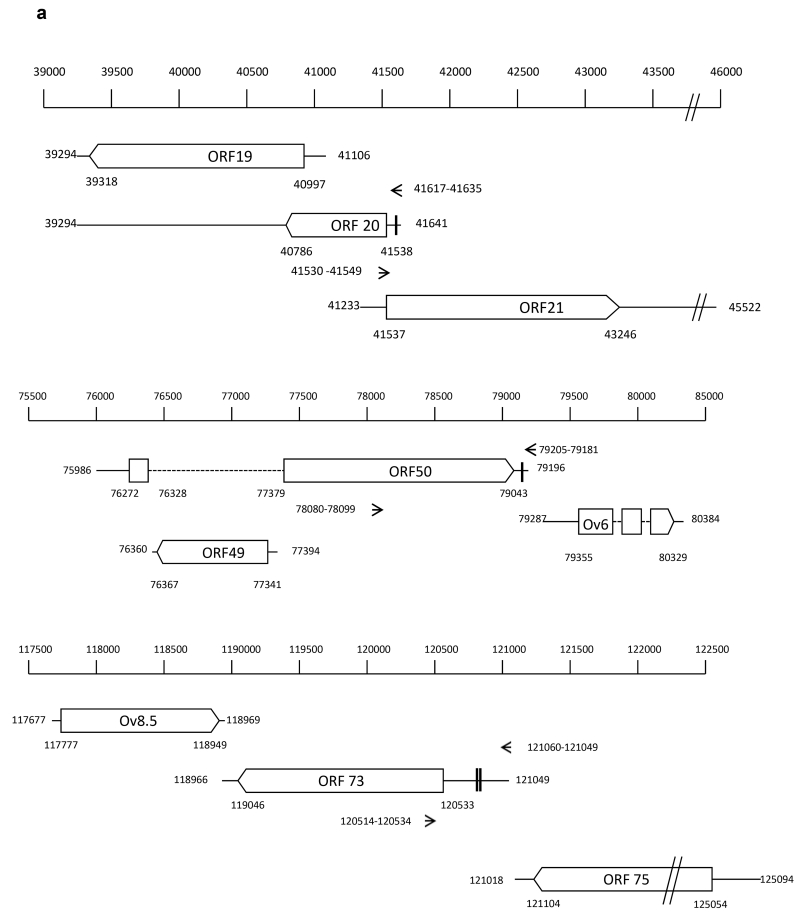
Fig. 1.
A. Genomic location of ORF 20 and ORF 50 relative to adjacent genes.
Genes are indicated by open boxes, arrow head represents direction of transcription. Dotted lines represent introns. The nucleotide positions representing the location of the predicted TATA box and poly A site for each gene are indicated. The position of primers used for PCR and cDNA priming (ORF 20) are indicated by arrows and nucleotide position. Predicted miRNA binding sequences are indicated by vertical bars in the respective UTRs.
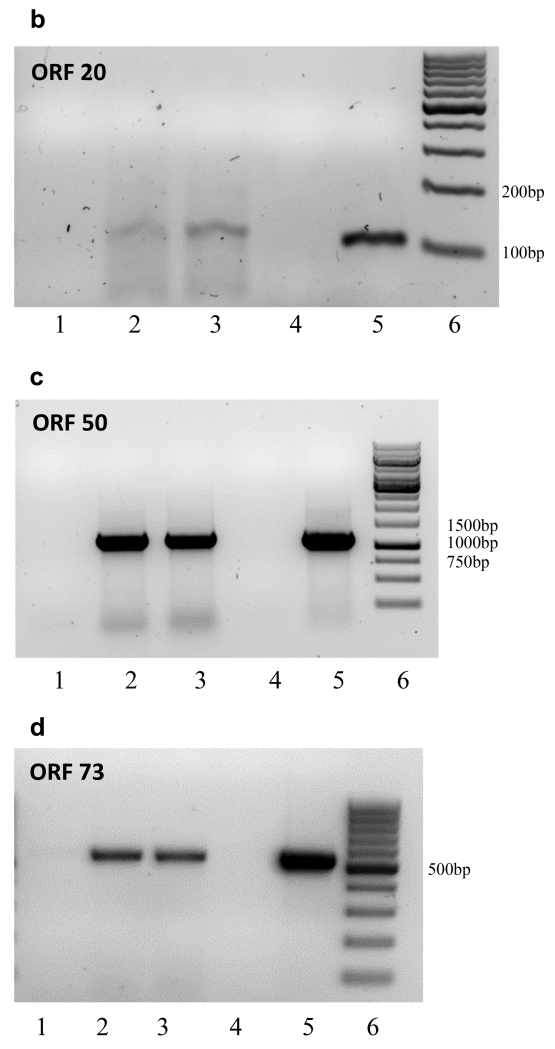
B. ORF 20 Lane 1, No RT; Lane 2, cDNA primed with 35 pm primer (250 ng); Lane 3, cDNA primed with 10 pm primer (66ng); Lane 4,: No template control; Lane 5, DNA positive control. Lane 6; Marker, Generuler 100bp
C. ORF 50 Lane 1, No RT, Lane 2, cDNA primed with Oligo dT; Lane 3, cDNA primed with random primers; Lane 4, No template control; Lane 5, DNA positive control; Lane 6, Marker, Generuler 1kb
D. ORF73 Lane 1, No RT, Lane 2, cDNA primed with Oligo dT; Lane 3, cDNA primed with random primers; Lane 4, No template control; Lane 5, DNA positive control; Lane 6, Marker, Generuler 100bp