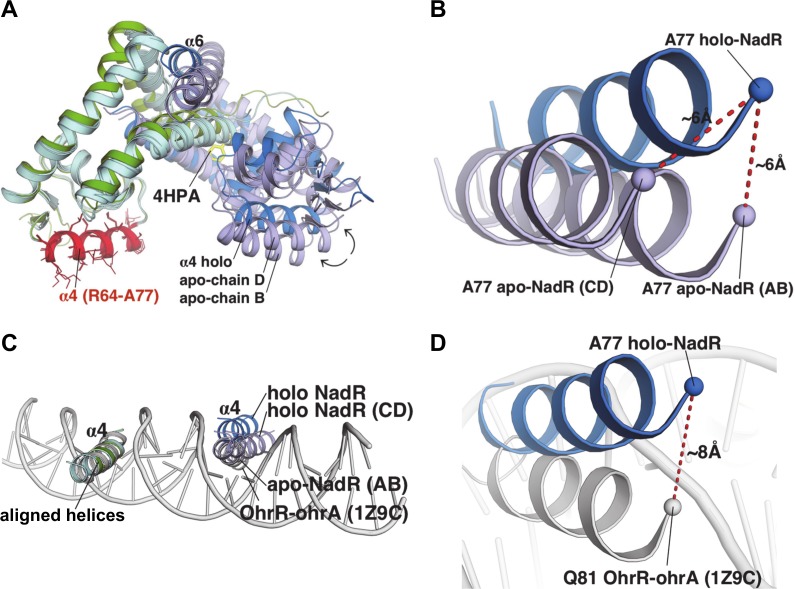
Fig 8. Structural comparisons of NadR and modelling of interactions with DNA.
(A) The holo-homodimer structure is shown as green and blue cartoons, for chain A and B, respectively, while the two homodimers of apo-NadR are both cyan and pale blue for chains A/C and B/D, respectively. The three homodimers (chains AB holo, AB apo, and CD apo) were overlaid by structural alignment exclusively of all heavy atoms in residues R64-A77 (shown in red, with side chain sticks) of chains A holo, A apo, and C apo, belonging to helix α4 (left). The α4 helices aligned closely, Cα rmsd 0.2Å for 14 residues. (B) The relative positions of the α4 helices of the 4-HPA-bound holo homodimer chain B (blue), and of apo homodimers AB and CD (showing chains B and D) in pale blue. Dashes indicate the Ala77 Cα atoms, in the most highly shifted region of the ‘non-fixed’ α4 helix. (C) The double-stranded DNA molecule (grey cartoon) from the OhrR-ohrA complex is shown after superposition with NadR, to highlight the expected positions of the NadR α4 helices in the DNA major grooves. The proteins share ~30% amino acid sequence identity. For clarity, only the α4 helices are shown in panels (B) and (C). (D) Upon comparison with the experimentally-determined OhrR:ohrA structure (grey), the α4 helix of holo-NadR (blue) is shifted ~8Å out of the major groove.