Fig. 3.
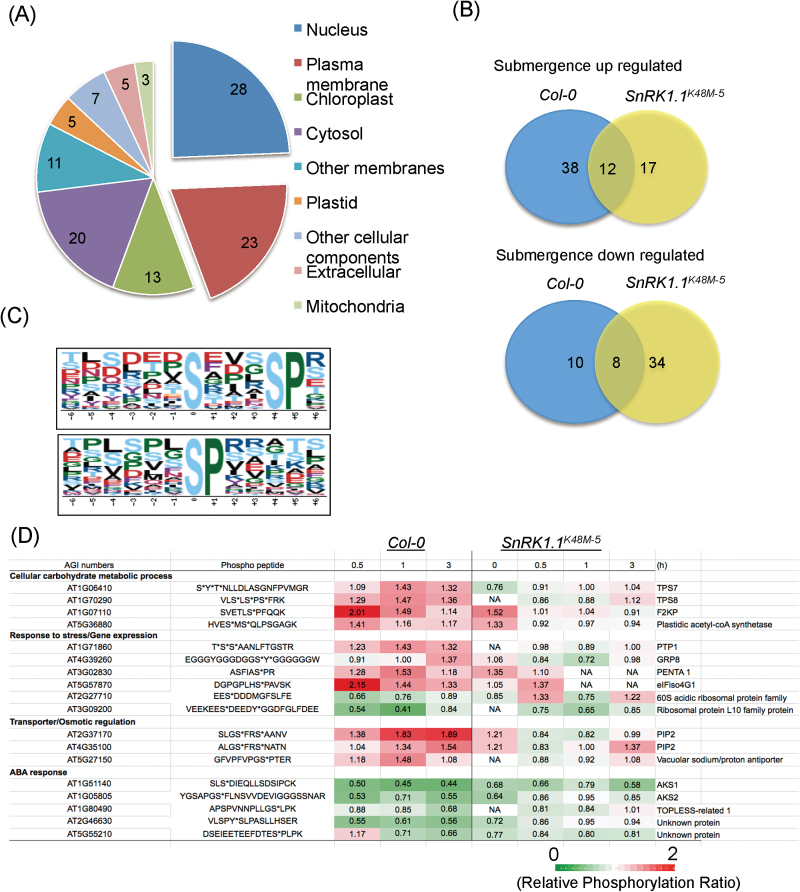
Phosphoproteomic profiles of the SnRK1.1 K48M mutant and Col-0 under submergence. The proteins with phosphorylation levels that were significantly different at any time point under submergence (at room temperature in the dark) were selected to for analysis. (A) Gene ontology (GO) catalogues of the submergence-changed phosphorylated proteins in Col-0. (B) Venn diagram illustrating the overlap of up-regulated (upper diagram) and down-regulated (lower diagram) phosphorylated proteins identified in Col-0 and SnRK1 K48M. (C) Sequence logos of the phosphorylation sites seen in submergence up-regulated phosphopeptides. (D) The four gene ontology groups in the SnRK1.1 phosphorylation list, where the statistical differences (P value) of protein phosphorylation in treatment and control of Col-0 was smaller than 0.05, but that in SnRK1 K48M-5 was not;. ‘*’ represents the phosphorylation site. The numbers indicate the average value of the phosphorylation fold change of all the biological repeats. The time point data not detected in the three batches or only detected once in three batches is marked as NA (not available).