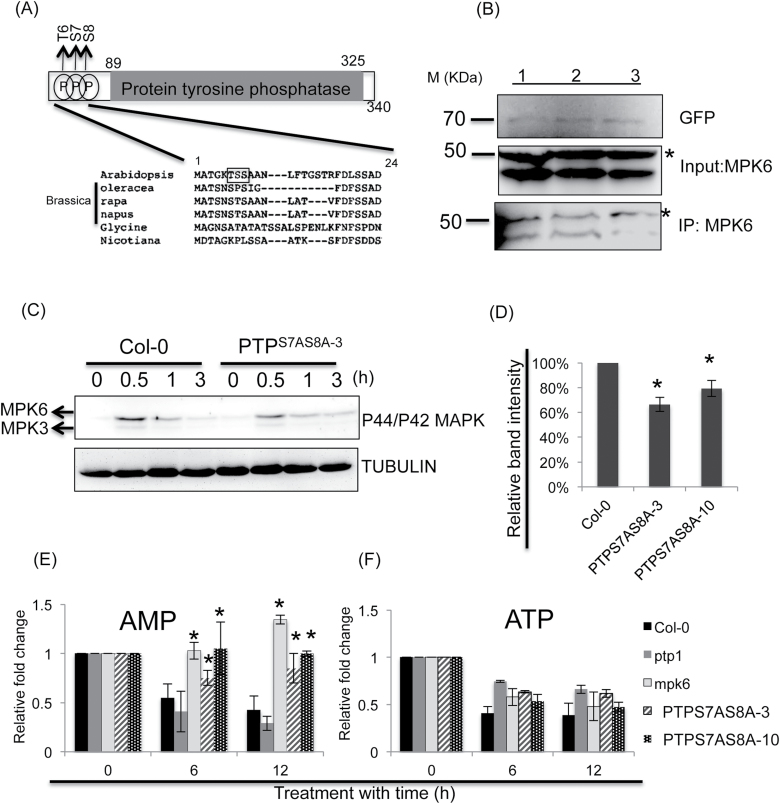
Fig. 6.
SnRK1.1 regulated MPK6 signalling by phosphorylating PTP1 under submergence. (A) Comparison of three phosphorylation sites (Thr6, Ser7, and Ser8) of Arabidopsis PTP1 with orthologues in other Brassica species, Glycine max and Nicotiana tomentosiformis by ClustalW. The black frame in the amino acid alignment indicates the phosphorylation sites of Arabidopsis PTP1. (B) The interaction of PTP1 and MPK6 was disrupted by PTP1S7DS8D. Upper panel: after immune precipitation, the abundance of (1) PTP1–GFP, (2) PTPS7AS8A–GFP, and (3) PTP1–S7DS8D–GFP conjugated in the beads of each transient samples. Middle panel: the abundance of MPK6 in the three PTP1 transient expression samples. Lower panel: the pull-down MPK6 were detected in PTP1–GFP and PTPS7AS8A–GFP transient expression samples, but not in PTP1–S7DS8D–GFP. *: Non-specific band. At least three biological repeats were performed and showed similar patterns. (C) Phosphorylation of MPK3/6 in Col-0, and PTP1 S7AS8A-3 under submergence. (D) The quantification of band intensity of (C) Western blot by GelEval software (FrogDance). The band intensity of 0.5h submergence in Col-0 and PTP1 S7AS8A lines was quantified, and TUBULIN was used as an internal control. The AMP (E) and ATP (F) profiles in Col-0, ptp1, mpk6, and two PTP1 S7AS8A lines under submergence. Statistical differences between Col-0 and mutant samples were determined by Student’s t test. *, P <0.05.