Fig. 7.
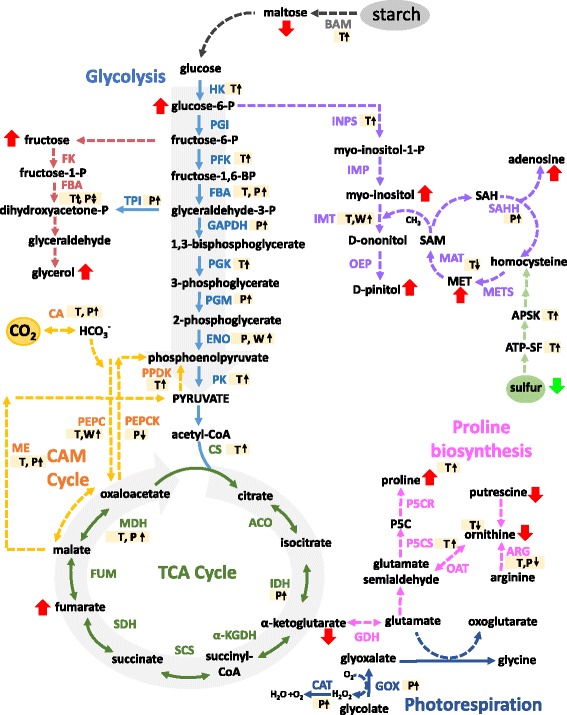
Integration of transcript, protein, metabolite and ionome data into EBC metabolic pathways. Data from all studies was obtained from 6-week-old plants treated for 2 weeks with 200 mM NaCl. Bladder cell extract was collected at the end of the dark period. Transcriptomic data from Oh et al., [24] and metabolomics data from Barkla and Vera-Estrella, [28]. T, transcript; P, protein and W, western blot analysis. Red arrows indicate changes in metabolites. Enzyme abbreviations: BAM – ϐ-amylase, HK – hexokinase, PGI – glucose-6P-isomerase, PFK – phosphofructokinase, FBA – aldolase, TPI – triose-P-isomerase, G3PD – glyceraldehyde-3P-dehydrogenase, PGK – phosphoglycerate kinase, PGM – phosphoglycerate mutase, ENO – enolase, PK- pyruvate kinase, FK – fructokinase, CS – citrate synthase, ACO – aconitase, IDH – isocitrate dehydrogenase, α-KGDH - α-ketoglutarate dehydrogenase, SCS – succinyl-CoA synthetase, SDH – succinate dehydrogenase, FUM – fumerase, MDH – malate dehydrogenase, PEPCK – PEP carboxykinase, ME – malic enzyme, PEPC – phosphoenolpyruvate carboxylase, PPDK – pyruvate-Pi-dikinase, CA – carbonic anhydrase, GDH – glutamate dehydrogenase, P5CS - pyrroline-5-carboxylate synthase, P5CR - pyrroline-5-carboxylase reductase, OAT - ornithine aminotransferase, ARG – arginase, INSP - myo-inositol 1-phosphate synthase, IMP – myo-inositol monophosphatase, IMT – inositol methyl transferase, OEP - ononitol epimerase, MAT – methionine adenosyltransferase, SAM - S-adenosyl methionine, SAH - S-adenosylhomocysteine, SAHH – S-adenosylhomocysteine hydrolase, MET – methionine, METS – methionine synthase, ATP-SF - ATP-sulfurylase, APSK - 5’-adenylylsulfate kinase, GOX - glucose oxidase, CAT – catalase
