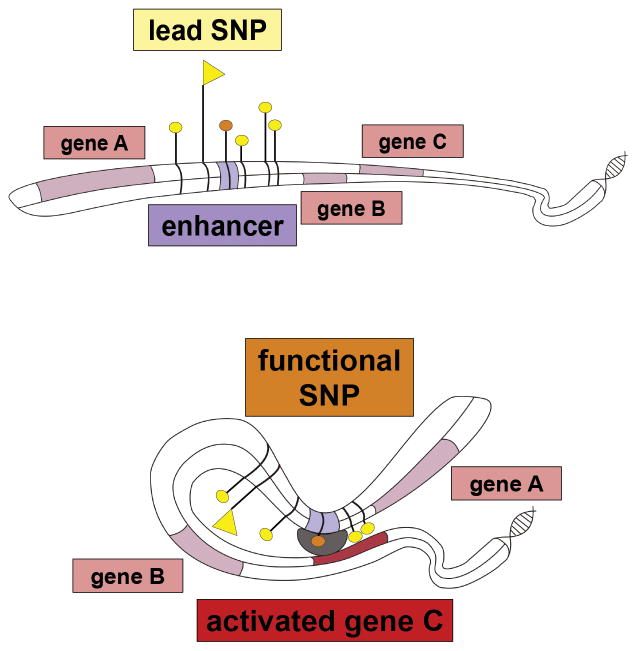
Figure 1. Mechanism by which non-coding risk SNP can affect phenotype.
Top: multiple SNPs associated with disease are located in the intergenic region proximal to genes A, B and C. One of the SNPs with genome-wide significance is situated within a cis- regulatory element (orange tag). The lowest P-value SNP (‘lead SNP’, flag tag) lies outside the regulatory element.
Bottom: Through bending of the DNA molecule the regulatory element gets into physical contact with the promoter of its target gene, in this case gene C which is not the gene in closest proximity, leading to regulation of its expression (‘activation’ or upregulation in case of an enhancer element). The SNP located within the regulatory element (‘functional SNP’, orange tag) can now affect transcription by for instance altering transcription factor (TF) binding affinity based on genotype via disruption of a TF binding motif