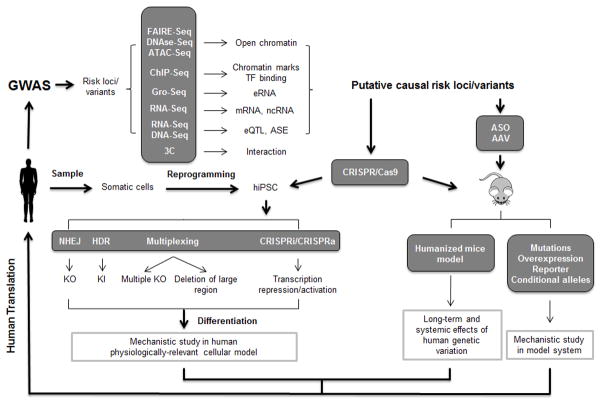
Figure 2. Experimental tools for GWAS functional follow-up studies.
GWAS findings can be functionally annotated using genome-wide methods, which help prioritize loci with likely biological function. These putative risk loci can be further interrogate by genome editing using the CRISPR/Cas9 system in vitro and in vivo. Additionally, adeno-associated virus (AAV) and antisense oligos (ASO) can be employed to study candidate gene knockdown and overexpression in the mouse model. TF: transcription factor; eRNA: enhancer RNA; ncRNA: non-coding RNA; eQTL: expression quantitative trait loci; ASE: allele-specific expression; iPSC: induced pluripotent stem cells; NHEJ: non-homologous end joining; HDR: homology-directed repair; KO: knockout; KI: knockin.