Abstract
It has been proposed that the polymerase gene of barley yellow dwarf virus and related viruses is expressed by a ribosomal frameshift event during translation. The 5' end of this gene overlaps with the 3' end of an upstream gene that is in a different reading frame. The region of overlap is similar to sequences in retro- and coronaviruses that are known to express their polymerase genes by frameshifting. This overlap region includes a "shifty" heptanucleotide, followed by a highly structured region that may contain a pseudoknot. Sequences of 115 or 144 base pairs that span this region from barley yellow dwarf virus (PAV serotype) genomic RNA were introduced into a plasmid, so that a reporter gene could be expressed in plant cells only if a minus one (-1) frameshift event occurred. Frameshifting was detected at a rate of approximately 1%. This frameshifting was abolished when the stop codon at the 3' end of the upstream open reading frame was deleted. A sequence expected to form a strong stem-loop immediately upstream of the frameshift site was unnecessary for frameshifting, and initiation at AUG codons within the stem-loop appeared to be inhibited. Like viruses that infect hosts in other kingdoms, plant viruses also can induce frameshifting in translation of their genes.
Full text
PDF
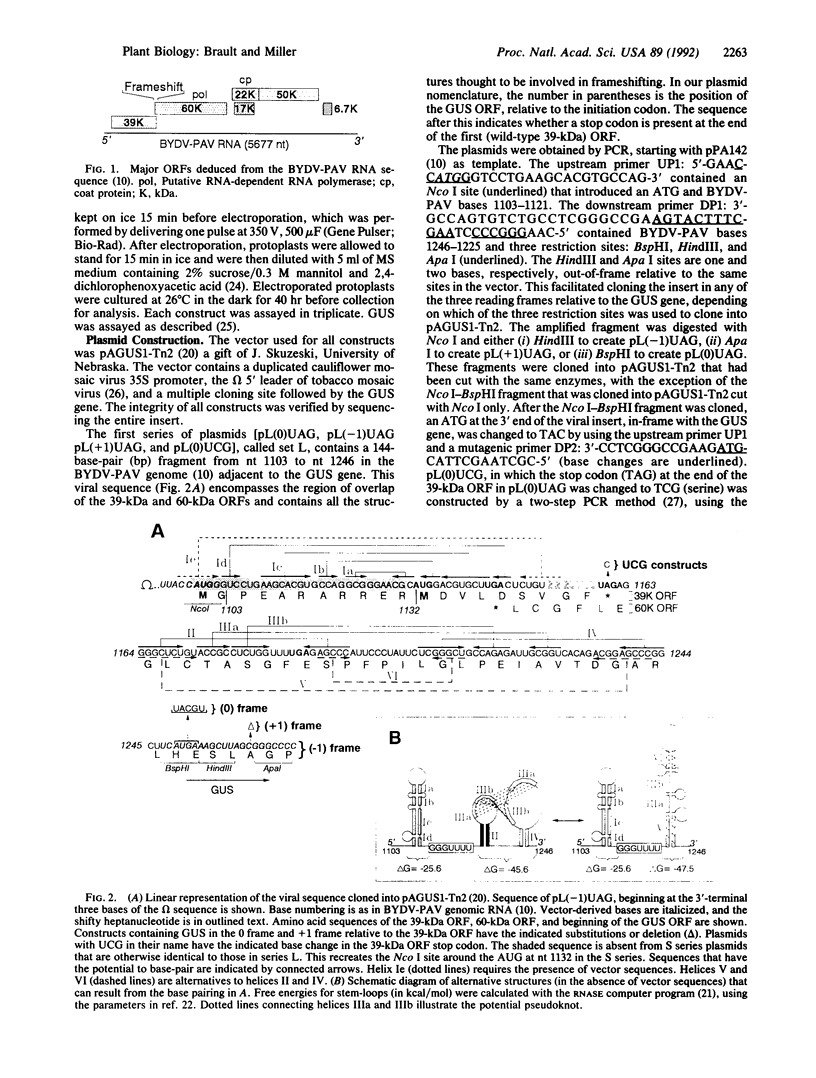




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Atkins J. F., Weiss R. B., Gesteland R. F. Ribosome gymnastics--degree of difficulty 9.5, style 10.0. Cell. 1990 Aug 10;62(3):413–423. doi: 10.1016/0092-8674(90)90007-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bredenbeek P. J., Pachuk C. J., Noten A. F., Charité J., Luytjes W., Weiss S. R., Spaan W. J. The primary structure and expression of the second open reading frame of the polymerase gene of the coronavirus MHV-A59; a highly conserved polymerase is expressed by an efficient ribosomal frameshifting mechanism. Nucleic Acids Res. 1990 Apr 11;18(7):1825–1832. doi: 10.1093/nar/18.7.1825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brierley I., Boursnell M. E., Binns M. M., Bilimoria B., Blok V. C., Brown T. D., Inglis S. C. An efficient ribosomal frame-shifting signal in the polymerase-encoding region of the coronavirus IBV. EMBO J. 1987 Dec 1;6(12):3779–3785. doi: 10.1002/j.1460-2075.1987.tb02713.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brierley I., Digard P., Inglis S. C. Characterization of an efficient coronavirus ribosomal frameshifting signal: requirement for an RNA pseudoknot. Cell. 1989 May 19;57(4):537–547. doi: 10.1016/0092-8674(89)90124-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cedergren R., Gautheret D., Lapalme G., Major F. A secondary and tertiary structure editor for nucleic acids. Comput Appl Biosci. 1988 Mar;4(1):143–146. doi: 10.1093/bioinformatics/4.1.143. [DOI] [PubMed] [Google Scholar]
- Clare J. J., Belcourt M., Farabaugh P. J. Efficient translational frameshifting occurs within a conserved sequence of the overlap between the two genes of a yeast Ty1 transposon. Proc Natl Acad Sci U S A. 1988 Sep;85(18):6816–6820. doi: 10.1073/pnas.85.18.6816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demler S. A., de Zoeten G. A. The nucleotide sequence and luteovirus-like nature of RNA 1 of an aphid non-transmissible strain of pea enation mosaic virus. J Gen Virol. 1991 Aug;72(Pt 8):1819–1834. doi: 10.1099/0022-1317-72-8-1819. [DOI] [PubMed] [Google Scholar]
- Dinman J. D., Icho T., Wickner R. B. A -1 ribosomal frameshift in a double-stranded RNA virus of yeast forms a gag-pol fusion protein. Proc Natl Acad Sci U S A. 1991 Jan 1;88(1):174–178. doi: 10.1073/pnas.88.1.174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freier S. M., Kierzek R., Jaeger J. A., Sugimoto N., Caruthers M. H., Neilson T., Turner D. H. Improved free-energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci U S A. 1986 Dec;83(24):9373–9377. doi: 10.1073/pnas.83.24.9373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fromm M., Taylor L. P., Walbot V. Expression of genes transferred into monocot and dicot plant cells by electroporation. Proc Natl Acad Sci U S A. 1985 Sep;82(17):5824–5828. doi: 10.1073/pnas.82.17.5824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallie D. R., Lucas W. J., Walbot V. Visualizing mRNA expression in plant protoplasts: factors influencing efficient mRNA uptake and translation. Plant Cell. 1989 Mar;1(3):301–311. doi: 10.1105/tpc.1.3.301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallie D. R., Sleat D. E., Watts J. W., Turner P. C., Wilson T. M. The 5'-leader sequence of tobacco mosaic virus RNA enhances the expression of foreign gene transcripts in vitro and in vivo. Nucleic Acids Res. 1987 Apr 24;15(8):3257–3273. doi: 10.1093/nar/15.8.3257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatfield D., Feng Y. X., Lee B. J., Rein A., Levin J. G., Oroszlan S. Chromatographic analysis of the aminoacyl-tRNAs which are required for translation of codons at and around the ribosomal frameshift sites of HIV, HTLV-1, and BLV. Virology. 1989 Dec;173(2):736–742. doi: 10.1016/0042-6822(89)90589-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herlitze S., Koenen M. A general and rapid mutagenesis method using polymerase chain reaction. Gene. 1990 Jul 2;91(1):143–147. doi: 10.1016/0378-1119(90)90177-s. [DOI] [PubMed] [Google Scholar]
- Heus H. A., Pardi A. Structural features that give rise to the unusual stability of RNA hairpins containing GNRA loops. Science. 1991 Jul 12;253(5016):191–194. doi: 10.1126/science.1712983. [DOI] [PubMed] [Google Scholar]
- Huang W. M., Ao S. Z., Casjens S., Orlandi R., Zeikus R., Weiss R., Winge D., Fang M. A persistent untranslated sequence within bacteriophage T4 DNA topoisomerase gene 60. Science. 1988 Feb 26;239(4843):1005–1012. doi: 10.1126/science.2830666. [DOI] [PubMed] [Google Scholar]
- Jacks T., Madhani H. D., Masiarz F. R., Varmus H. E. Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell. 1988 Nov 4;55(3):447–458. doi: 10.1016/0092-8674(88)90031-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacks T., Varmus H. E. Expression of the Rous sarcoma virus pol gene by ribosomal frameshifting. Science. 1985 Dec 13;230(4731):1237–1242. doi: 10.1126/science.2416054. [DOI] [PubMed] [Google Scholar]
- Kozak M. Circumstances and mechanisms of inhibition of translation by secondary structure in eucaryotic mRNAs. Mol Cell Biol. 1989 Nov;9(11):5134–5142. doi: 10.1128/mcb.9.11.5134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayo M. A., Robinson D. J., Jolly C. A., Hyman L. Nucleotide sequence of potato leafroll luteovirus RNA. J Gen Virol. 1989 May;70(Pt 5):1037–1051. doi: 10.1099/0022-1317-70-5-1037. [DOI] [PubMed] [Google Scholar]
- Miller W. A., Waterhouse P. M., Gerlach W. L. Sequence and organization of barley yellow dwarf virus genomic RNA. Nucleic Acids Res. 1988 Jul 11;16(13):6097–6111. doi: 10.1093/nar/16.13.6097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore R., Dixon M., Smith R., Peters G., Dickson C. Complete nucleotide sequence of a milk-transmitted mouse mammary tumor virus: two frameshift suppression events are required for translation of gag and pol. J Virol. 1987 Feb;61(2):480–490. doi: 10.1128/jvi.61.2.480-490.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moosmayer D., Reil H., Ausmeier M., Scharf J. G., Hauser H., Jentsch K. D., Hunsmann G. Expression and frameshifting but extremely inefficient proteolytic processing of the HIV-1 gag and pol gene products in stably transfected rodent cell lines. Virology. 1991 Jul;183(1):215–224. doi: 10.1016/0042-6822(91)90134-W. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skuzeski J. M., Nichols L. M., Gesteland R. F. Analysis of leaky viral translation termination codons in vivo by transient expression of improved beta-glucuronidase vectors. Plant Mol Biol. 1990 Jul;15(1):65–79. doi: 10.1007/BF00017725. [DOI] [PubMed] [Google Scholar]
- Veidt I., Bouzoubaa S. E., Leiser R. M., Ziegler-Graff V., Guilley H., Richards K., Jonard G. Synthesis of full-length transcripts of beet western yellows virus RNA: messenger properties and biological activity in protoplasts. Virology. 1992 Jan;186(1):192–200. doi: 10.1016/0042-6822(92)90073-x. [DOI] [PubMed] [Google Scholar]
- Veidt I., Lot H., Leiser M., Scheidecker D., Guilley H., Richards K., Jonard G. Nucleotide sequence of beet western yellows virus RNA. Nucleic Acids Res. 1988 Nov 11;16(21):9917–9932. doi: 10.1093/nar/16.21.9917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walbot V. RNA editing fixes problems in plant mitochondrial transcripts. Trends Genet. 1991 Feb;7(2):37–39. doi: 10.1016/0168-9525(91)90225-F. [DOI] [PubMed] [Google Scholar]
- Weiss R. B., Dunn D. M., Atkins J. F., Gesteland R. F. Ribosomal frameshifting from -2 to +50 nucleotides. Prog Nucleic Acid Res Mol Biol. 1990;39:159–183. doi: 10.1016/s0079-6603(08)60626-1. [DOI] [PubMed] [Google Scholar]
- Weiss R. B., Dunn D. M., Shuh M., Atkins J. F., Gesteland R. F. E. coli ribosomes re-phase on retroviral frameshift signals at rates ranging from 2 to 50 percent. New Biol. 1989 Nov;1(2):159–169. [PubMed] [Google Scholar]
- Wilson W., Braddock M., Adams S. E., Rathjen P. D., Kingsman S. M., Kingsman A. J. HIV expression strategies: ribosomal frameshifting is directed by a short sequence in both mammalian and yeast systems. Cell. 1988 Dec 23;55(6):1159–1169. doi: 10.1016/0092-8674(88)90260-7. [DOI] [PubMed] [Google Scholar]
- Xiong Z., Lommel S. A. The complete nucleotide sequence and genome organization of red clover necrotic mosaic virus RNA-1. Virology. 1989 Aug;171(2):543–554. doi: 10.1016/0042-6822(89)90624-7. [DOI] [PubMed] [Google Scholar]
- Young M. J., Kelly L., Larkin P. J., Waterhouse P. M., Gerlach W. L. Infectious in vitro transcripts from a cloned cDNA of barley yellow dwarf virus. Virology. 1991 Jan;180(1):372–379. doi: 10.1016/0042-6822(91)90042-a. [DOI] [PubMed] [Google Scholar]
- de Smit M. H., van Duin J. Secondary structure of the ribosome binding site determines translational efficiency: a quantitative analysis. Proc Natl Acad Sci U S A. 1990 Oct;87(19):7668–7672. doi: 10.1073/pnas.87.19.7668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ten Dam E. B., Pleij C. W., Bosch L. RNA pseudoknots: translational frameshifting and readthrough on viral RNAs. Virus Genes. 1990 Jul;4(2):121–136. doi: 10.1007/BF00678404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Steeg H., van Oostrom C. T., Hodemaekers H. M., van Kreyl C. F. Cloning and functional analysis of the rat ornithine decarboxylase-encoding gene. Gene. 1990 Sep 14;93(2):249–256. doi: 10.1016/0378-1119(90)90232-g. [DOI] [PubMed] [Google Scholar]