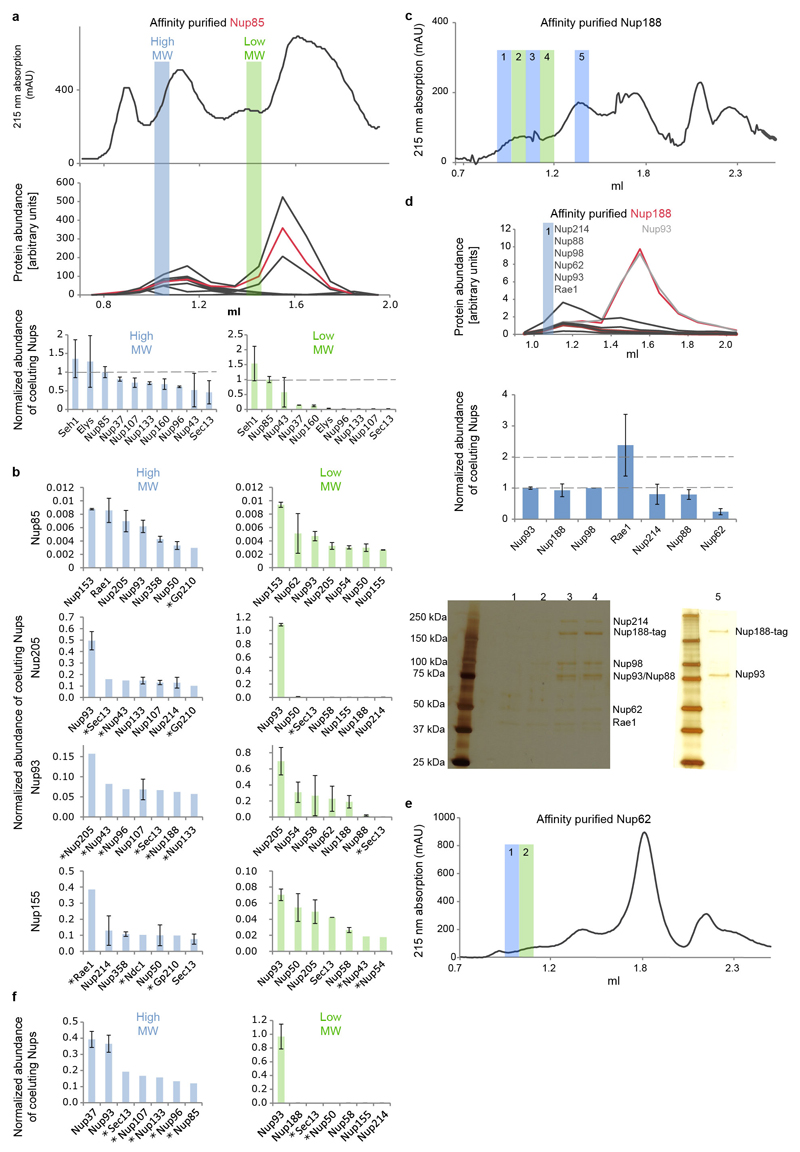
Extended Data Figure 8. Co-elution analysis to detect weak nucleoporin interactions.
To detect weak interactions of scaffold nucleoporins we combined rapid affinity isolation with gel filtration and quantitative targeted proteomics to measure absolute protein abundances. Hek293 cells expressing various affinity-tagged Nups (in contrast to Fig. 3c without nocodazole arrest) were lysed using mild conditions and sonication for protein solubilization. Affinity isolates were subjected to gel-filtration and all fractions were analyzed using targeted mass spectrometry, as previously described 24,29 to measure protein abundances in the high and low molecular weight fractions. The high-molecular-weight fractions will be indicative of potential outgoing interactions of large molecular species. The low molecular weight fraction will highlight smaller fragments that occur after sonication. In case of affinity tagged Nup85 (a) the top panel shows the 215 nm absorption curve of the gel-filtration experiment. The middle panel shows the arbitrary protein abundance units of Y-complex members in all fractions (red for Nup85, black for all other Y-complex members). Protein abundances (normalized to the affinity tagged protein) in the high-molecular-weight fractions (blue bar) and low-molecular-weight fraction (green bar) are shown as bar charts in the bottom panel. The low molecular weight peak corresponds to the small arm proteins (Nup85, Seh1 and Nup43) the high molecular weight peak to the intact Y-complex. (b) The same approach was applied to Nup205, Nup93 and Nup155. The seven most abundant co-eluting Nups are shown for the high-molecular-weight fractions (blue bar plots) and low-molecular weight fraction (green bar plots). In case of Nup85 (top) the seven most abundant proteins apart from the Y-complex members are shown. Weak interactions of Y-complex members with Nups 205 and 93 are apparent. In case of Nup155, weak interactions are detected with CR and NR members, as well as Sec13 that localizes to the proximity of the C-term domain of Nup155 when fitted into the density connecting the IR with CR/NR. Tpr was excluded from this analysis since it was present in all fractions. Protein abundances based on single reference peptides are marked with an asterisk. (c) Same as (a, top panel) but for Nup188 affinity purified from nocodazole arrested cells. (d) Same as Fig. 3c but for Nup188 affinity purified from nocodazole arrested cells. Co-eluting species in the high molecular weight fraction are similar to the ones observed for affinity-purified Nup62 (Fig 3c). Isostoichiometric amounts of Nups 188, 98, 93, 62 and an enrichment of Nups 214, 88 and Rae1 were detected. The Nup188–Nup93 heterodimer thus binds to Nups that are well-established components of the CR, which is consistent with the systematic fitting approach. (e) Same as (c) but corresponding to Fig. 3c. (f) Same as (b; second panel for Nup205) but for nocodazole-arrested cells. In case of Nup205, the co-purifying species are overall similar in nocodazole-arrested as compared to untreated cells.