Figure 4.
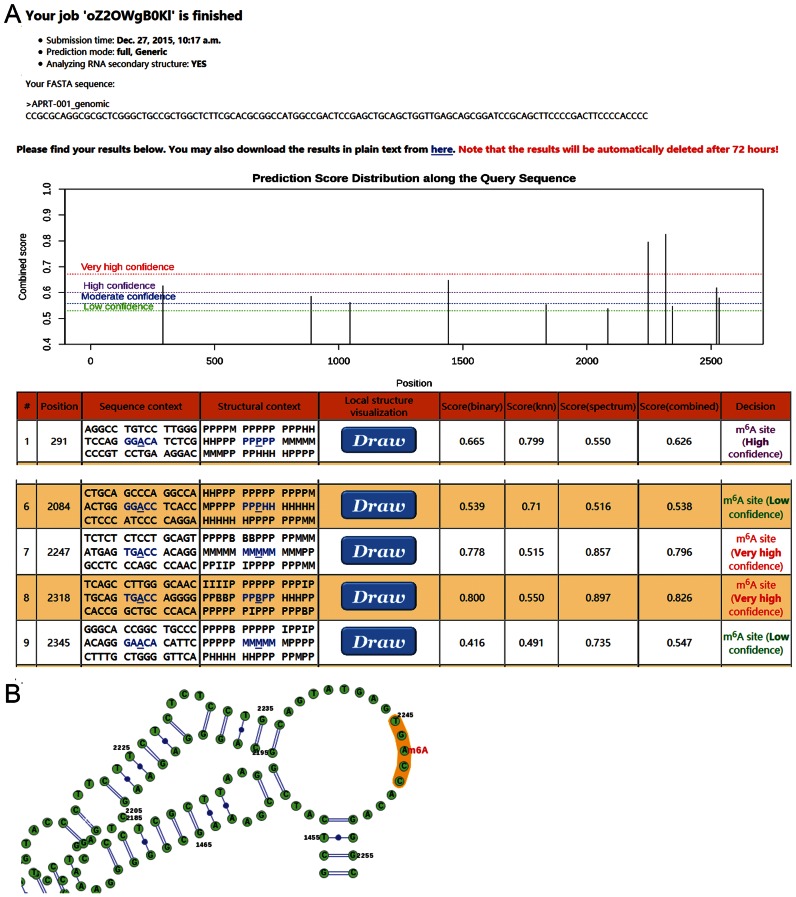
A sample SRAMP prediction result. The genomic sequence of a representative APRT transcript (ENST00000378364) was used as the input to the full transcript mode predictor, and the analysis of secondary structures was enabled. (A) The overview of result page. The exhibition of the query sequences is truncated, and only the detailed results for the first predicted m6A site and those near the pathogenic mutation site (G2246->C) are shown. The H, M, I, B, P in the secondary structure strings mean hairpin loop, multiple loop, interior loop, bulged loop and paired residues, respectively. In addition to such string, a graphical representation of the local secondary structure will be generated when click on the ‘draw’ button. (B) A graphical representation of the local secondary structure context around the mutation site. This graphical representation was generated by SRAMP server exploiting the VARNA structural visualization tool. We focused on the local secondary structure in proximal to the mutation site for clarity.