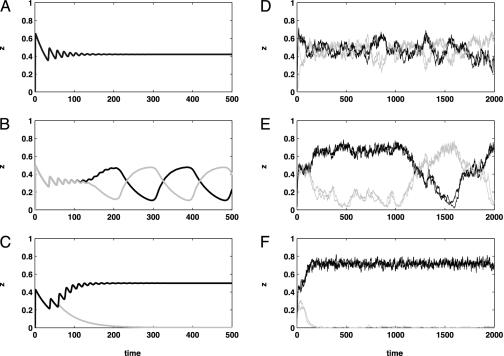
Fig. 2.
An illustration of the changing dynamics for both the original deterministic model and our stochastic mean-field approximation model for pathogen populations with two loci (i.e., four strains). Two strains comprising one discordant set are plotted in black; the other discordant set is plotted in gray. Plotted in all simulations is the proportion of the host population immune to each of the four strains. Parameter values [corresponding to the parameter notation of Gupta et al. (6)] used for the ODE simulations (A–C) were μ = 0.02, σ = 10, R0 = 2, α = 1, with the only difference in parameter values being the degree of cross-immunity γ (0.3 in A, 0.7 in B, 0.9 in C). The mean-field approximation of the stochastic IBM (D–F) used parameter values corresponding to our model's parameter notation in Table 1: C = 12, r = 0.0953, τ = 0.0042, β = 0.2472, 1/μ = 7, 1/σ = 23, P = 223, n = 2. The degree of cross-immunity γ was 0.01 in D, 0.95 in E, and 2.00 in F. (Note that γ is defined slightly differently in our model compared with its definition in ref. 6). A and D, with the lowest values ofγ, both have no strain structure, with the mean diversity in D being 0.9882 and the mean discordance being 0.6723. B and E have intermediate values of γ, and both exhibit cyclical strain dynamics. Mean diversity in E is 0.8733, and mean discordance is 0.7496. C and F have high values of γ, and both exhibit strong strain structure, with one discordant set being dominant. Mean diversity in F is 0.5480, and mean discordance is 0.9720. Simulations were run for 2,000 time steps for the IBM and 500 time steps for the ODE.