Upon quiescence entry, yeast cells assemble telomere hyperclusters. These structures localize to the nuclear membrane in an Esc1-dependent manner and assemble through the combined action of the Sir complex, deacetylation of H4K16, the binding of the linker histone H1, and condensin.
Abstract
Like other eukaryotes, Saccharomyces cerevisiae spatially organizes its chromosomes within the nucleus. In G1 phase, the yeast’s 32 telomeres are clustered into 6–10 foci that dynamically interact with the nuclear membrane. Here we show that, when cells leave the division cycle and enter quiescence, telomeres gather into two to three hyperclusters at the nuclear membrane vicinity. This localization depends on Esc1 but not on the Ku proteins. Telomere hypercluster formation requires the Sir complex but is independent of the nuclear microtubule bundle that specifically assembles in quiescent cells. Importantly, mutants deleted for the linker histone H1 Hho1 or defective in condensin activity or affected for histone H4 Lys-16 deacetylation are impaired, at least in part, for telomere hypercluster formation in quiescence, suggesting that this process involves chromosome condensation. Finally, we establish that telomere hypercluster formation is not necessary for quiescence establishment, maintenance, and exit, raising the question of the physiological raison d’être of this nuclear reorganization.
INTRODUCTION
In yeast, just as in other eukaryotes, chromosomes are spatially organized (Taddei et al., 2010; Albert et al., 2012). This organization is thought to influence gene expression but also DNA repair, replication, and recombination (Cavalli and Misteli, 2013). In the G1 phase of the proliferating cycle, the yeast nucleus adopts a configuration called “Rabl,” in which centromeres are gathered together close to the spindle pole body (SPB), the yeast equivalent of the centrosome, through short nuclear microtubules (Guacci et al., 1997; Jin et al., 1998, 2000; Bystricky et al., 2004). The nucleolus containing the ribosomal DNA localizes at the SPB opposite pole (Yang et al., 1989). Constrained by the nuclear envelope, chromosomes undergo ATP-dependent undirected motion, each locus mobility being determined by its position along the arm (Marshall et al., 1997; Heun et al., 2001; Bystricky et al., 2004; Hajjoul et al., 2013). Finally, the 32 yeast telomeres cluster into 6–10 dynamic and mobile foci mostly found in the nuclear membrane proximity (Palladino et al., 1993; Gotta et al., 1996; Hediger et al., 2002; Bystricky et al., 2005).
Yeast telomeric sequences are short (∼350 base pairs) and made of T(G1-3) repeats with a 3′ G-rich extension. Repeats of X elements, and eventually of Y′ elements, compose yeast subtelomeric regions. Many proteins interact with telomeric DNA, including various telomerase-associated complexes, but also Rap1, which binds double-stranded DNA, the Ku proteins, and the Sirtuin (Sir) complex. The Sir complex is composed of the NAD+-dependent histone deacetylase Sir2 and the silencing proteins Sir3 and Sir4. This complex is recruited to telomeric DNA in part by Rap1, which interacts with Sir3 and Sir4. The Sir complex then associates with the hypoacetylated N-terminal tails of the histones H3 and H4 and spreads from telomeres into subtelomeric regions (for reviews, see Taddei et al., 2010; Wellinger and Zakian, 2012; Kupiec, 2014).
Telomere localization at the nuclear periphery is dependent on intricate and partially redundant pathways involving Esc1, the Ku complex, and Mps3, an essential integral nuclear membrane protein containing a SUN domain that is involved in SPB organization (Jaspersen et al., 2002; Nishikawa et al., 2003); Mps3, Esc1, and yKu80 interact with Sir4 (Taddei et al., 2010; Wellinger and Zakian, 2012). The nuclear pore proteins of the Nup84 subcomplex have also been involved in telomere tethering at the nuclear periphery (Therizols et al., 2006). In proliferating cells, telomere clustering into foci results from stochastic contacts generated by random motion, the chromosome arm length, and the nucleolus exclusion zone determining the telomere exploration radius (Schober et al., 2008; Therizols et al., 2010; Zimmer and Fabre, 2011; Wong et al., 2012). Telomere clusters are mobile and only transient (Schober et al., 2008; Therizols et al., 2010), their formation requiring intertelomere association through Sir3 oligomerization (Ruault et al., 2011). In fact, telomere clustering is thought to be generated via an aggregation/dissociation equilibrium, the association rate being dependent on geometrical parameters, and the dissociation rate being regulated by the Sir3 protein level (Hozé et al., 2013).
When cells cease to proliferate, they may enter a reversible nondividing state called quiescence (Coller, 2011; O’Farrell, 2011; De Virgilio, 2012). Many years ago, it was shown that, in nondividing yeast cells, chromatin adopts a more compact arrangement compared with proliferating cells (Piñon, 1978; Lohr and Ide, 1979). More recently, Patterson and coworkers have shown that chromatin is more condensed in glucose-starved quiescent cells than in proliferating cells, this compaction being partly due to an increase in chromatin-bound Hho1, the yeast equivalent of the linker histone H1 (Schäfer et al., 2008). Chromatin compaction generally involves the N-terminal tail of the histone H4, and more precisely, deacetylation of the Lys-16 (Wilkins et al., 2014). Importantly, while acetylated H4K16 is predominant in proliferating cells (Smith et al., 2003), no acetylated H4K16 has been detected in quiescent cells by nanoelectrospray tandem mass spectrometry (Ngubo et al., 2011). These data suggest that, upon quiescence establishment in yeast, chromatin adopts a compact conformation, maybe through an increased binding of Hho1 and a possible deacetylation of H4K16.
We have previously reported that, when quiescence is induced by glucose exhaustion, the yeast nucleus is drastically reshaped (Laporte and Sagot, 2014). Indeed yeast cells assemble a stable nuclear microtubule array that not only causes nucleolus relocalization but also centromere declusterization and redistribution along the microtubule array (Laporte et al., 2013). Very recently, centromere declustering in quiescence was confirmed by chromosome conformation capture (3C) in an elegant study clearly demonstrating that the overall quiescent cells’ genome organization is significantly different from the one observed in proliferating cells (Rutledge et al., 2015). In fact, Broach and coworkers have established that, in quiescent yeast cells, intrachromosomal interactions increase at longer distance and subtelomeric regions interact with each other much more frequently (both intra- and interchromosomally), these rearrangements being dependent on the condensin complex (Rutledge et al., 2015). The topological reorganizations reported by Rutledge and coworkers were confirmed by Guidi and colleagues using 3C (Guidi et al., 2015). In the later study, fluorescence microscopy analyses demonstrated that, in a subpopulation of dense quiescent cells selected on a Percoll gradient (Allen et al., 2006), telomeres regrouped in one or two hyperclusters in the center of the cell nucleus in a Sir3-dependent manner (Guidi et al., 2015). Astonishingly, telomere hyperclusters were not observed by Rutledge and colleagues, who instead reported an increased number of Rap1–green fluorescent protein (GFP) foci in quiescence (Rutledge et al., 2015). These discrepancies hence question the existence of telomere hyperclusters in the nucleus of live yeast quiescent cells.
In this study, we have investigated the behavior of telomeres in a whole population of live yeast cells that entered quiescence following carbon source exhaustion. Using various telomere-associated proteins fused to fluorescent proteins and fluorescence in situ hybridization (FISH), we show that telomeres form two to three hyperclusters that localize close to the nuclear membrane. This nuclear membrane proximity depends on Esc1 but not on the Ku complex. Additionally, we demonstrate that telomere hypercluster formation not only depends on the Sir complex but also on the chromatin condensation machinery, the hyperclustering being affected in hho1Δ or condensin mutants. We further reveal that deacetylation of the histone H4K16 is critical for the quiescence-induced telomere hyperclustering process. Importantly, upon quiescence exit, telomere hyperclusters slowly disassemble independently of actin and microtubule dynamics. Finally, we unambiguously establish that telomere hyperclustering is not required for cell survival in early quiescence, raising the question of the physiological raison d’être of this specific nuclear reorganization.
RESULTS AND DISCUSSION
Telomeres do form hyperclusters upon quiescence establishment
On carbon source exhaustion, budding yeast cells leave the cell cycle and enter quiescence. In these conditions, we have analyzed by FISH the localization of subtelomeric regions (Y′ subtelomere DNA sequences; Louis and Borts, 1995) in wild-type cells (WT). As previously described, 6–10 telomere clusters were detected in proliferating G1 cells (Palladino et al., 1993; Gotta et al., 1996). By contrast, in quiescent cells, only 2.3 ± 0.5 bright telomere clusters were observed (Figure 1A). These telomere hyperclusters contained Sir2 (Figure 1B), but also Sir3, Rap1, and Sir4 (Figure 1C, Supplemental Figure S1, A and B, and unpublished data), and form regardless of the culture temperature (for culture at 30°C, see Figure 1; at 25°C, Supplemental Figure S1F; and at 37°C, Figure 4D). Of note, in quiescent cells, one of the Sir2-GFP detected foci colocalized with the nucleolus (Supplemental Figure S1C). Kinetic analyses revealed that telomere hypercluster formation was a rather slow process that reached its plateau at ∼6–7 d of culture at 30°C (Figure 1D and Supplemental Figure S1D). A similar kinetic was observed when quiescence was induced by an abrupt carbon depletion, explaining why Rutledge and coworkers did not observe telomere hyperclusters after one day of carbon shortage when using this protocol (Rutledge et al., 2015; Supplemental Figure S1E). We also found that virgin and young mother cells were slightly more prone to form telomere hyperclusters than cells that have undergone more than four divisions (Supplemental Figure S1F). Our findings are consistent with chromosome conformation capture analyses demonstrating that intertelomeric interactions increase in quiescent cells (Guidi et al., 2015; Rutledge et al., 2015). They are also in agreement with Guidi and colleagues, who found that subtelomeric regions form one to two clusters in 7-d-old W303 cells (Guidi et al., 2015). Taken together, these data clearly establish that telomeres do form hyperclusters upon quiescence establishment following carbon source exhaustion.
FIGURE 1:
Telomeres form hyperclusters in quiescence. (A) FISH experiments on proliferating G1 and quiescent WT cells (7 d, DAPI: blue; Y′ sequences: green). (B) Telomere hyperclusters contain Sir2. Colocalization by immuno-FISH of Y′ sequences (red) and Sir2-GFP foci (green) in WT quiescent cells (7 d). (C) Telomere hyperclusters contain Sir3 and Rap1. Colocalization of Sir2-GFP and Sir3-RFP (top) or Rap1-RFP (bottom) in WT quiescent cells (7 d). White arrows point at telomere hyperclusters. Foci containing only Sir2-GFP correspond to the nucleolus (see Supplemental Figure S1C). Note that Sir2 nucleolus localization is lost for an unknown technical reason during the FISH procedure (compare B and C). (D) Telomere hypercluster formation kinetic upon quiescence entry. The number of telomere clusters per cell was scored over time in WT cells expressing Sir2-GFP. The mean number of telomere clusters per cell for each time point is indicated. ***, p < 1 × 10−5. Error bars are SD. Scale bars: 2 μm.
FIGURE 4:
Telomere hypercluster formation depends on the Sir complex and the chromatin condensation machinery. (A) Telomere hypercluster formation is affected in Sir mutants. Y′ sequence detection by FISH (green) in quiescent (7 d) WT, sir2Δ, sir4Δ, or sir3Δ cells stained with DAPI (blue). (B) Y′ sequence detection by FISH (green) in quiescent cells (6 d) with the indicated mutations in the histone H4 N-terminal tail stained with DAPI (blue). (C) Quiescent hho1Δ cells (7 d) expressing Sir2-GFP (green) and Bim1-RFP (red) and distribution of the number Sir2-GFP foci per cell in WT (red bars) and in hho1Δ (green bars) quiescent cell. (D) WT and smc2-8 cells expressing Sir2-GFP were grown 1 d at 25°C and then shifted for 2 d at 37°C. Representative cells and the distribution of Sir2-GFP foci per cell are shown. In A–C, the mean number of telomere clusters per cell is indicated. In C, the percentage of cells displaying a nuclear microtubule bundle in the population is indicated. Scale bars: 2 μm.
Telomere hyperclusters localize close to the nuclear membrane
In quiescent cells, we found that telomere hypercluster movements were confined (Figure 2A, red line), contrasting with their mobility in proliferating G1 cells (Figure 2A, green line). In fact, in quiescent cells, as in proliferating G1 cells, we mostly observed telomere hyperclusters close to the nuclear membrane (<250 nm, Figures 2B and 3C). This is in striking contrast with Guidi and coworkers, who described telomere hyperclusters in the inner zone of the quiescent cells nucleus (Guidi et al., 2015), and will be discussed in the following sections.
FIGURE 2:
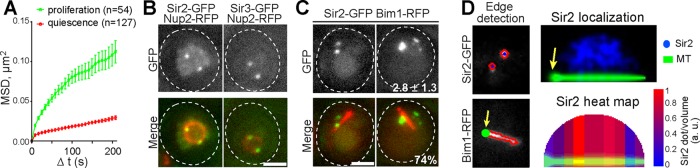
Telomere hyperclusters colocalize with the nuclear membrane. (A) Telomere hyperclusters (Sir2-GFP) display slower MSDs in WT quiescent cells (7 d, red line) than in proliferating G1 cells (green line). Error bars are SEM. (B) Telomere hyperclusters colocalize with the nuclear membrane. WT quiescent cells (7 d) expressing Nup2-RFP (red) and Sir2-GFP (green, left panel) or Sir3-GFP (green, right panel). (C and D) Telomere hyperclusters are distributed nonhomogenously around the quiescent cell nucleus periphery. (C) WT quiescent cells (7 d) expressing Sir2-GFP (green) and Bim1-RFP (red), a protein localized along the nuclear microtubule bundle, were used to analyze telomere hypercluster localization. The mean number of detected Sir2-GFP foci per cell and the percentage of cells displaying nuclear microtubule bundle in the population are indicated. (D) Left, schematics showing edge-based detection of telomere hyperclusters and nuclear microtubule bundle position. Right, Sir2-GFP localization (top, blue dots) and the corresponding heat map (bottom) are shown. In the two panels, a sum of nuclear volume 3D projections using the nuclear microtubule bundle (green) as an oriented symmetry axis reference (the SPB been the origin, yellow arrow) is shown. Scale bars: 2 μm.
FIGURE 3:

Telomere hypercluster localization at the nuclear membrane depends on Esc1. (A) Yku80 colocalizes with telomere hyperclusters. WT quiescent cells (5 d) expressing Yku80-GFP (green) and Sir3-RFP (red, left) or Sir2-RFP (red, right). (B) WT quiescent cells (6 d) expressing Spc42-RFP (red) and Esc1-GFP (green). Graph indicates Spc42 fluorescence intensity (red bar) and Esc1 fluorescence probability (green) along a line following the nuclear membrane (arrows). (C) Telomere hypercluster localization at the nuclear membrane depends on Esc1. Distance distributions between telomere hyperclusters (Sir3-GFP) and the nuclear membrane (Nup2-RFP) in WT, yku70Δ, yku80Δ, and esc1Δ quiescent cells (7 d). The orange zone corresponds to a distance smaller than the resolution limit (250 nm). The percentage of telomere hyperclusters localizing in this zone is indicated. WT, yku70Δ, yku80Δ, and esc1Δ quiescent cells expressing Sir3-GFP and Nup2-RFP are shown; the mean number of Sir3-GFP foci per cell is indicated. Scale bars: 2 μm.
To more precisely localize telomere hyperclusters, we took advantage of the nuclear microtubule bundle that emanates from the SPB in quiescent cell nuclei (Laporte et al., 2013). We coimaged Sir2-GFP together with the nuclear microtubule bundle (Figure 2C). The nuclear microtubule bundle was automatically detected and used as an axis of reference, the SPB being the origin of the axis (yellow arrow in Figure 2D) on two-dimensional (2D) projections of z-sections (maximum projection). The nuclei were consequently reduced to hemi-disks. The relative position of each telomere hypercluster was then determined using a dedicated MatLab script (Figure 2D and Materials and Methods). We found that, statistically, telomere hyperclusters were excluded from both the SPB region and an SPB opposite zone that corresponds to the tip of the nuclear microtubule bundle (Figure 2D). The exclusion of telomeres from this later zone could not be due to the presence of the nucleolus, as we have previously shown that the nuclear microtubule bundle assembly causes nucleolus displacement toward one side of the nucleus in more than 80% of the cells (Laporte et al., 2013). It is interesting to note that the telomere reorganization in quiescence is distinct from the bouquet arrangement adopted by meiotic cells that gather their telomeres together close to the SPB (Trelles-Sticken et al., 1999, and references therein). Consistently, we found that neither Csm4 nor Ndj1, two proteins involved in telomere rearrangement during bouquet formation, were required for telomere hypercluster formation upon quiescence entry (Supplemental Figure S1G).
Telomere hyperclusters localize to nuclear membrane through Esc1 but not yKu
To get insight into the molecular relationships between telomere hyperclusters and the nuclear membrane in quiescent cells, we focused on yKu70/80 and Esc1, proteins known to play a role in telomere localization at the nuclear periphery in proliferating cells (Taddei et al., 2010; Wellinger and Zakian, 2012; Kupiec, 2014). In WT quiescent cells, we found that yKu80 colocalized with telomere hyperclusters (Figure 3A), while yKu70 could not be detected (unpublished data). Intriguingly, we observed that Esc1 localized as discrete zones all around the nuclear membrane of quiescent cells (Supplemental Figure S2A) with the exception of the SPB proximal zone and the region opposite to the SPB (Figure 3B), just as telomere hyperclusters do. Of note, in proliferating G1 cells, Esc1 was detected next to the SPB but was also excluded from the zone opposite to the SPB, probably because of the presence of the nucleolus in this region (Supplemental Figure S2B; Taddei et al., 2004). Importantly, we found that telomeres were still hyperclustered in esc1Δ cells, but their localization close to the nuclear membrane was strongly impaired. Indeed, telomere hyperclusters randomly localized inside the nucleus (for Sir3-GFP, see Figure 3C; for Sir2-GFP, see Supplemental Figure S2C). Yet no significant difference in telomere hypercluster motility was measured between esc1Δ and WT quiescent cells (Supplemental Figure S2D). This suggests that the slow motion of telomere hyperclusters observed in quiescent cells was not a consequence of a tight interaction with the nuclear membrane. Additionally, deletion of yKu protein–encoding genes had no effect either on telomere hypercluster formation or localization to the nuclear membrane vicinity (Figure 3C and Supplemental Figure S2C), and no additional defect was observed when combining esc1Δ with yku deletions (Supplemental Figure S2, C and E). Taken together, our data demonstrate that quiescent cell telomere hyperclusters localize close to the nuclear membrane through Esc1.
Telomere hypercluster formation requires the Sir complex
In proliferating cells, the Sir complex has been involved in telomere clustering (Palladino et al., 1993; Gotta et al., 1996). Using FISH, we found that deletion of sir2Δ, sir3Δ, or sir4Δ affected telomere hypercluster formation in quiescent cells (Figure 4A). This is in agreement with the findings of Guidi and colleagues, who described the absence of Rap1-GFP hyperclusters in sir3Δ quiescent cells (Guidi et al., 2015). Additionally, we found that the Sir3 signal detected using red fluorescent protein (RFP) or GFP increased in quiescent cells compared with G1 proliferating cells (Supplemental Figure S3, A and B), and Western blotting indicated a slight augmentation of Sir3 steady-state level in quiescence (Supplemental Figure S3C). In proliferating cells, Ruault and coworkers have shown that overexpression of Sir3 leads to telomere hypercluster formation and that nonacetylable Sir3 can promote telomere clustering independent of its spreading in subtelomeric regions (Ruault et al., 2011). Because chromatin immunoprecipitation experiments did not detect any significant changes in the spreading of Sir3 between exponentially growing cells and quiescent cells (Guidi et al., 2015), we propose that, in quiescence, increased Sir3–Sir3 interactions promote trans interaction between Sir3-bound telomeres, leading to the formation of hyperclusters. This increased Sir3 recruitment in quiescence may rely on posttranslational modifications that were shown to modulate Sir3 interaction with chromatin in actively dividing cells (Arnaudo et al., 2013).
Ruault and colleagues have shown that, when Sir3 is highly overexpressed in proliferating cells, telomere hyperclusters localize inside the nucleus rather than at the nuclear periphery, possibly because of a competition between Sir3 and Esc1 for the binding of Sir4 (Ruault et al., 2011). We found that the Sir3 steady-state level in quiescent BY strains was slightly lower than the one we detected in quiescent W303 (Supplemental Figure S3C), a difference that may explain why telomere hyperclusters were found in the nucleoplasm of quiescent W303 cells (Guidi et al., 2015).
Telomere hypercluster formation requires H4K16 deacetylation and the chromatin condensation machinery
In quiescent cells, the histone H4 Lys-16 (H4K16) is solely found in its deacetylated form (Ngubo et al., 2011). To address the role of this histone H4 posttranslational modification, we analyzed telomere hypercluster formation in histone H4 mutants. No telomere hyperclusters were detected in H4 mutants bearing N-terminal tail deletion or H4K16A or H4K16Q point mutations. Conversely, a mutation preventing H4K16 acetylation (H4K16R) had no effect on telomere hypercluster formation (Figure 4B). Thus deacetylation of H4K16 seems to be a key event that is needed for telomere hypercluster formation. Two nonexclusive hypotheses can be envisioned regarding the influence of H4K16 deacetylation on telomere hypercluster formation. First, as Sir3 is known to preferentially bind unacetylated H4K16 (Carmen et al., 2002; Onishi et al., 2007; Oppikofer et al., 2011), the extensive deacetylation of H4K16 might increase the recruitment of Sir3 and consequently participate in the Sir3-induced strengthening of telomere–telomere interaction in quiescent cells. Second, deacetylation of H4K16 has been associated with chromatin condensation, since it regulates the interaction between the histone H4 N-terminal tail and the H2A-H2B dimer within the nucleosomes (Wilkins et al., 2014). As mutations altering H4/H2A-H2B interaction such as H4K16A or H4K16Q also affected telomere hypercluster formation, we speculate that this process could be influenced by chromatin condensation.
In fact, in proliferating cells, telomere cluster formation is thought to be influenced by structural constraints (Hozé et al., 2013). The linker histone Hho1 has been involved in chromatin compaction during quiescence establishment (Schäfer et al., 2008). Interestingly, we found that telomere hypercluster formation was slightly impaired in quiescent hho1Δ cells (Figure 4C). Moreover, inactivating the condensin complex (smc2-8) drastically compromised telomere hypercluster formation (Figure 4D and Supplemental Figure S3D), in agreement with Rutledge and coworkers, who demonstrated that Smc2 is involved in chromosomes compaction in quiescent yeast cells (Rutledge et al., 2015). Therefore we could envision that, upon quiescence establishment, chromosome-arm condensation is strengthened, thereby reducing the telomere exploration area and indirectly triggering telomere hypercluster formation.
Telomere hyperclusters slowly disassemble upon quiescence exit
When quiescence exit was triggered by cell refeeding, the number of telomeric clusters per cell progressively increased, reaching 6.7 ± 1.7 upon entry into S phase (Figure 5A). As for other quiescent cell–specific structures such as actin bodies, proteasome storage granules, or nuclear microtubule bundle (Sagot et al., 2006; Laporte et al., 2008, 2013), telomere hyperclusters disassembled even if the de novo protein synthesis was inhibited by cycloheximide (CHX; Figure 5B). During Saccharomyces cerevisiae meiosis, telomere movements were shown to be driven by actin cables (Koszul et al., 2008). Figure 5B shows that telomere hypercluster disassembly upon quiescence exit was clearly unaffected by latrunculin A, a drug causing the depolymerization of all yeast actin filament–containing structures (Ayscough et al., 1997), including the actin cables assembled within seconds upon quiescent cell refeeding (Sagot et al., 2006). At the present time, we do not know what causes telomere hypercluster disassembly upon quiescence exit. We can speculate that it may involve a sequence of events mirroring those occurring upon quiescence establishment, that is, the H4K16 reacetylation, the subsequent decrease in Sir3 interaction with subtelomeric regions, and chromosome arm decondensation upon entry into S phase.
FIGURE 5:
Telomere reorganization upon quiescence exit and viability of cells unable to form telomere hypercluster. (A) WT quiescent cells (7 d) expressing Sir2-GFP were refed with YPDA, and the number of telomere clusters per cell was scored over time. The mean number of clusters per cell is indicated. (B) WT quiescent cells (7 d) expressing Sir2-GFP (green) and Bim1-RFP (red) were refed with YPDA or with YPDA + CHX, or nocodazole or latrunculin A (Lat-A) or DMSO as a control and imaged 120 min after refeeding. Scale bars: 2 μm. (C) Micromanipulation of 2-, 7-, or 14-d-old WT, sir2Δ, sir3Δ, sir4Δ, hho1Δ, yku70Δ, and esc1Δ cells. Graphs indicate the percentage of micromanipulated cells that produced a colony after 3 d of growth on YPDA plate. Representative images of micromanipulation plates are shown.
Telomeres hyperclusters are not required for cell survival in quiescence
On quiescence establishment following glucose exhaustion, yeast cells assemble a nuclear bundle of microtubules that not only causes nucleolus relocalization but also centromere declusterization and redistribution (Laporte et al., 2013). Intriguingly, telomere hyperclustering was only slightly affected in some (kar3Δ and dyn1Δ) but not all (jnm1Δ) mutants in which the nuclear microtubule bundle formation is compromised (Supplemental Figure S3, E and F). Therefore, if the nuclear microtubule bundle formation clearly participates in several nuclear reorganizations occurring upon quiescence establishment (Laporte et al., 2013; Laporte and Sagot, 2014), it is not strictly required for telomere hypercluster formation. Accordingly, nocodazole treatment did not affect telomere hypercluster disassembly upon quiescence exit (Figure 5B), demonstrating that microtubule dynamics is not required for this process. Besides, the nuclear microtubule bundle assembly was not affected in hho1Δ, sir3Δ, or esc1Δ mutants (Figure 4 and Supplemental Figure S3G), demonstrating that microtubule remodeling in quiescence does not rely on telomere hypercluster formation or telomere interaction with the nuclear membrane. Therefore the reorganization of microtubules and telomeres can be dissociated and, as such, can be considered as independent quiescence-specific events.
Importantly, while the assembly of the nuclear bundle of microtubules is critical for cell survival in quiescence (Laporte et al., 2013), none of the mutations impeding telomere hypercluster formation or localization had a significant effect on cell viability in quiescence (Supplemental Figure S3H) or quiescent cells’ ability to give rise to a viable progeny upon quiescence exit (Figure 5C and Supplemental Figure S3I). The formation of other quiescent cell–specific structures (actin bodies or nuclear microtubule bundle) testified to accurate quiescence establishment in those mutants (Supplemental Figure S3G). A number of studies have proposed the existence of a correlation between gene subnuclear positioning and transcriptional activity in both yeast and mammals (Taddei et al., 2010; Albert et al., 2012; Nguyen and Bosco, 2015). Our data demonstrate that, upon glucose exhaustion, telomere hypercluster formation and localization are not required for survival in quiescence. Thus, if telomere hypercluster formation and localization modulate gene expression, this transcriptional regulation seems to not be mandatory for quiescence establishment, maintenance, and exit.
In conclusion, if the active assembly of quiescent cell–specific structures such as actin bodies or nuclear microtubule bundle are necessary for cell survival in quiescence (Sagot et al., 2006; Laporte et al., 2013), other reorganizations, like telomere hypercluster formation, are not strictly required for facing chronological aging (Figure 5). Of course, drugs or physical conditions to which mutants unable to assemble telomere hyperclusters in quiescence are sensitive could potentially be found. However, their specific effect on the quiescent state will be difficult to tackle. Nevertheless, at the present time, we cannot rule out that telomere hyperclustering may have a role in the long-term maintenance of yeast quiescence or could be involved in the fitness of quiescent exit.
Hence either telomere hypercluster formation in quiescence does have a physiological function, but we have not appreciated it yet, and an extensive amount of work is needed to shed light on the biological significance of this specific reorganization, or telomere hyperclusterization does not provide any beneficial trait. Why would cells actively embark on rearranging telomeres if this reorganization does not influence their survival in quiescence? In fact, telomere hypercluster formation could be just a passive consequence of other quiescence-induced modifications that may be vital for cells, such as chromatin hypercondensation (Piñon, 1978; Lohr and Ide, 1979; Schäfer et al., 2008; Rutledge et al., 2015) and/or the increase of nucleoplasm molecular crowding that alters the biophysical properties of the nuclear environment (Joyner et al., 2016). Therefore, to shed light on the obscure molecular mechanisms involved in quiescence survival, one of the key issues will be to establish whether each structure specifically assembled in quiescent cells results from a dedicated active process or whether it is just a passive consequence of another quiescence-induced phenomenon.
MATERIALS AND METHODS
Yeast strains and growth conditions
All the S. cerevisiae strains used in this study are isogenic to BY4741 or BY4742 available from GE Healthcare (Chicago, IL) and are listed in Supplemental Table S1. Yeast strains carrying GFP fusions were obtained from Invitrogen (Carlsbad, CA). Histone H4 mutants and the corresponding control strain were obtained from Invitrogen. The smc2-8 mutant (Strunnikov et al., 1995) was obtained from C. Boone (University of Toronto, Canada). The RFP (tdimer 2(12); Campbell et al., 2002) sequence carried by plasmids p3695, p4589, p5041, p5043, and p5045 was integrated at the 3′ end of the NUP2, BIM1, RAP1, SIR2, and SIR3 endogenous loci, respectively. Three tandem copies of the GFP sequence carried by the plasmid p4587 were integrated at the 3′ end of the BIM1 endogenous locus (Laporte et al., 2013). Details of the constructions are available upon request. The Sir2-GFP fusion protein functionality was analyzed by measuring cell size using a Beckman Coulter Multi-sizer 4 (Beckman Coulter, Brea, CA). As expected sir2Δ cells were large, the median cell volume for sir2Δ being 53.4 μm3 (>40,000 cells counted). By contrast, the median cell volumes for WT, Sir2-GFP, and Sir2-RFP cells were 45.4, 45.5, and 43.8 μm3 (>25,000 cells counted), respectively. The functionality of the Sir3-GFP construct was attested by the viability of the Sir3-GFP pfd1Δ strain.
All experiments were done at least in duplicate and, unless specified, more than 200 cells were scored. Yeast cells were grown in liquid yeast–peptone–dextrose–adenine (YPDA) medium at 30°C in flasks, as described previously (Sagot et al., 2006), except for Figure 4D and Supplemental Figure S3D, for which cells were grown at 37°C and 25°C, respectively.
For quiescence exit in the presence of different drugs (Figure 5B), cells were preincubated 30 min in the presence of the drug before quiescence exit. Drugs used were CHX (180 μM; Sigma-Aldrich, St. Louis, MO), nocodazole (7.5 μM; Sigma-Aldrich), and latrunculin A (200 μM; Enzo Life Sciences, Farmingdale, NY).
Cell staining
For identification of mother and daughter cells (Supplemental Figure S1F), 7-d-old WT cells expressing Sir2-GFP were incubated 5 min with Calcofluor white (20 μg/ml), then washed twice in phosphate-buffered saline (PBS) and imaged.
Cell viability (Supplemental Figure S3H) was scored after 5 min incubation in a solution containing 0.2% of methylene blue (Sigma-Aldrich) and 2% sodium citrate solution, pH 7 (Sigma-Aldrich).
For actin phalloidin staining (Supplemental Figure S3G), cells were fixed with freshly made paraformaldehyde (PFA) prepared as follows: PFA (3.8% final; Sigma-Aldrich) was resuspended in PEM (0.1 M PIPES, 1 mM EGTA, 1 mM MgSO4, pH 6.9, with NaOH 5 N), vortexed every 20 min for 1 h at 70°C, and then centrifuged 5 min at 3000 rpm. After 1 h fixation at 30°C, cells were washed twice with PEM, resuspended in 1.5 ml of PEM + 1% Triton X-100, and incubated 3 min. Samples were then washed twice in 1.5 ml of PEM, resuspended in PEM containing 1/10 volume of Alexa Fluor phalloidin (Invitrogen) and incubated 24 h at 4°C. Finally, cells were washed twice, resuspended in a mounting solution containing 70% glycerol and 5 mg/l para-phenylenediamine, and imaged.
The probe for FISH experiments was obtained by PCR with a template plasmid containing ∼5 kb of Y′ element (pEL42H10; Louis and Borts, 1995) and using the primer pair GAAGAATTGGCCTGCTCTTG/CCGTAAGCTCGTCAATTATT. PCR purification was followed by a nick translation labeling reaction using the Nick Translation kit (DIG-Nick Translation Mix; Roche, Mannheim, Germany; ref. 11745816910). After 5 min at 98°C, the probe was purified by ethanol precipitation and resuspended at the concentration of 25 ng/μl in a hybridization mix (50% formamide, 2×SSC, 10% dextran-sulfate, 0.5 mg/ml single-stranded salmon sperm DNA).
Spheroplasts were created as follows. Quiescent cells were fixed 15 min in 1.9% PFA (Sigma-Aldrich; diluted in PEM), centrifuged 1 min at room temperature, and washed twice in PEMS (PEM + 1.2 M sorbitol filter sterilized). Cells were then incubated 10 min in SH buffer (β-mercaptoethanol 0.5 M, Tris-HCl 0.1 M, pH 9.3) and washed twice in KCl Tris-HCl buffer (5×, KCl 2.5 M, Tris-HCl 50 mM, pH 7). Cells were washed twice in PEMS, resuspended in 1 ml PEMS containing 27 mg/ml Zymo 20T (MP Biomedical, Santa Ana, CA; ref. 32092), and incubated at 37°C for 90 min (>60% of the cells were spheroplasted). Spheroplasts were resuspended in 1 ml PEMS + 1% Triton and incubated ∼5 min at room temperature. After four washes in 1 ml PEM, cells were incubated 1 h at 37°C in 1 ml PEM + 25 μl RNaseA (10 mg/ml).
For FISH experiments (Figures 1A and 4, A and B, and Supplemental Figure S2E), 200 μl of the spheroplast preparation described above was centrifuged and washed sequentially with 150 μl 2×SSC, 150 μl 2×SSC 10% formamide, 150 μl 2×SSC 20% formamide, and 150 μl 2×SSC 40% formamide. Between each wash, cells were incubated 15 min. Then, 35 μl of the probe mix (150 ng of probe, 30 μl hybridization mix, adjusted to 38 μl with H2O) incubated at 75°C for 15 min was added. After resuspension, cells were incubated at 75°C for 10 min and then at 37°C overnight. Cells were then washed three times in 150 µl 2xSSC (with an incubation of 30 min at 37°C between each wash), resuspended in 150 µl PBS-BAG (PBS + 1% bovine serum albumin [BSA; Sigma-Aldrich; A-7511] + 0.1% Na azide) and incubated 1 h at room temperature. After centrifugation, 100 µl PBS-BAG + anti-digitonine fluorescein isothiocyanate (FITC)–conjugated antibodies (1/50) was added and cells were incubated overnight. The next day, cells were washed twice in 100 µl PBS-BAG and once in 100 µl PBS. Cells were then mounted on poly-l-lysine–coated slides, with anti-fading reagent containing 4′,6-diamidino-2-phenylindole (DAPI) solution (Fluoroshield; Sigma-Aldrich, ref: F6057) before imaging.
For immuno-FISH experiments (Figure 1B), spheroplasts were washed twice for 30 min in 100 μl PEM-BAL (PEM + 1% BSA [Sigma, ref. B4287], 0.1% NaN3, 100 mM lysine hydrochloride). After addition of GFP antibody (anti-GFP from mouse; Roche, ref. 11814460001, 1/50), cells were incubated overnight and then washed three times in PEM-BAL. The secondary antibody (sheep anti-mouse CY3 1/400; ThermoFisher Scientific, Waltham, MA) was added in PEM-BAL, and cells were incubated overnight and then washed once in PEM-BAL (300 μl) and twice in PEM (500 μl). For the last wash, 25 μl RNaseA (10 mg/ml) was added and incubated 3 h at 37°C. Finally, spheroplasted cells were fixed using 3% PFA for 30 min and then washed three times in 500 μl PEM. The FISH protocol described above was then applied.
Cell viability
Quiescent cells (n > 120 cells) were micromanipulated as described previously (Laporte et al., 2011). Plates were incubated 3 d at 30°C before colony scoring.
Colony-forming capacity was addressed after 2, 7, and 14 d at 30°C by plating 200 cells, measured using a Beckman Coulter Multi-sizer 4, on YPDA. Each strain was tested in duplicate, and each plating was done in triplicate.
Fluorescence microscopy
Cells were observed in a fully automated Zeiss 200M inverted microscope (Carl Zeiss, Thornwood, NY) equipped with an MS-2000 stage (Applied Scientific Instrumentation, Eugene, OR), a Lambda LS 175-W xenon light source (Sutter, Novato, CA), a 100× 1.4 NA Plan-Apochromat objective, and a 5-position filter turret. For GFP imaging, we used a FITC filter (Ex: HQ487/25–Em: HQ535/40–BS: Q505lp). For RFP imaging, we used a Cy3 filter (Ex: HQ535/50–Em: HQ610/75–BS: Q565lp). For calcofluor imaging, we used a DAPI filter (Ex: 360/40–Em: 460/50–BS: 400). All the filters are from Chroma Technology. Images were acquired using a CoolSnap HQ camera (Roper Scientific, Tucson, AZ). The microscope, camera, and shutters (Uniblitz, Rochester, NY) were controlled by SlideBook software 5.0 (Intelligent Imaging Innovations, Denver, CO). Images are, unless specified, 2D maximal projection of z-stacks performed using a 0.25 μm step. For live-cell imaging, 2 μl of the cell culture was spotted onto a glass slide and immediately imaged at room temperature.
For fluorescence intensity measurement of Esc1-GFP (Figure 3B and Supplemental Figure S2B), a line scan (i1) of 5 pixel width was drawn all along nuclear membrane using ImageJ software to simultaneously measure Esc1-GFP and Spc42-RFP fluorescence. Background was subtracted from these fluorescence intensities. The maximum intensity in Spc42 data (i.e., the SPB location) was used to align the two data sets. For Spc42 graph representation, the fluorescence intensity of the different measurements was summed. For Esc1 graph representation, data were processed as follows: Esc1 intensities <0 were set to zero, while intensities >0 were set to 1. This binary approach allows the construction of a probability map (sum of 0 and 1 divided by the number of measurements at a specific distance from the SPB) and avoids the overrepresentation of strong fluorescence signals.
For fluorescence intensity measurement of Sir3, Sir2, and Sir4 (Supplemental Figure S3, A and B), a circle (i1) containing both RFP or GFP signal and background was drawn around a telomeric focus (Supplemental Figure S3A) or around all telomeric signals (Supplemental Figure S3B) using ImageJ software. A circle two times larger at the same location was drawn to calculate the intensity of the surrounding background (i2). The real intensity (ir) was calculated as follows: ib = (i2 × area i2) − (i1 × area i1) and ir = i1 − [ib/(area i2 − area i1)]. Measures displayed on graphs were obtained by multiplying ir by area i1. Sir3-GFP/RFP signals measured in proliferation and in quiescence were multiplied by an arbitrary factor 2 to be on the same scale as Sir2-GFP/RFP signals.
For distance measurement (Figure 3C and Supplemental Figure S2C), distances between Sir3-GFP or Sir2-GFP foci and the nuclear membrane were determined using ImageJ. Given that the position of the nuclear membrane cannot be set precisely at the top or bottom of the cell, only telomere hyperclusters located in the equatorial plane of the nuclear membrane were scored.
Edge detection and heat-map construction
Individual cells were marked out manually directly from the raw data, and the corresponding z-stacks were exported as separate TIFF files using ImageJ. The detection and localization of telomere clusters and microtubule bundle were performed in each z-stacks using the Matlab (Mathworks) scripts summarized below. For each type of detection, the approaches used and the choice of analysis parameters were confirmed by comparison of the detected shapes with visual inspection. Individual telomere clusters were isolated and counted as follows: First, the averaged intensity (background) was subtracted from each plane of the z-stack independently and the resulting signal was smoothed with a 3 × 3 pixel Wiener filter and then normalized to its absolute maximum value over the entire stack. Then intensity thresholds were used to convert the z-stack to binary images. Matlab’s Image Processing Toolbox shape-detection and morphological operation algorithms were next applied to detect and isolate 2D shapes in each plane of the z-stack, yielding a list of GFP spots with their three-dimensional (3D) coordinates and area. To prevent double counting of the same telomere clusters, we removed spots within a neighborhood of 7 pixels. This step eliminates the trace of a unique telomere cluster in adjacent planes of the z-stack (due to the blur along the z-axis), and ensures that the z coordinate of a cluster is attributed to the plane where its projection has the largest area (equatorial plane). Similarly, nuclear microtubule bundle signal (Bim1-RFP signal) was first normalized to its stack-wide maximum value and binarized using an intensity threshold. Then shape detection and morphological operations were performed to find the contour of the bundle in each plane. The extremity close to the brightest region of the bundle was identified as the SPB. For determination of the localization of the bundle extremities along the Z axis, the signal intensity was averaged over a 25-pixel window centered on the extremities of the trace of the bundle in each plane. The variations of these signals over the planes showed clear maxima in specific planes, indicating the most likely localization of each bundle’s extremity along the z-axis. Finally, we calculated the distance of each telomere cluster to the two extremities of the bundle in 3D and normalized this distance to the length of the bundle to allow compilation of telomere cluster localization relative to nuclear microtubule bundle axis over multiple cells (Figure 2D, top right).
For heat-map construction, the nucleus was “sliced” along the nuclear microtubule bundle. The number of telomere clusters within each slice was normalized to the 3D volume of the slice, computed under the assumption of a spherical nucleus. The resulting heat map, shown in Figure 2D, bottom right, represents the probability density to find a telomere cluster in various regions of the nucleus. Note that the assumption of a spherical nucleus tends to underestimate the cluster density close to the equator. Indeed, for nuclei that tend to stretch along the microtubule bundle (as observed in vivo in some quiescent cells), the equatorial slices are smaller than for the spherical model, and the volumetric factor is overestimated.
Mean square displacement (MSD) measurement
For individual spot dynamics analysis (Figure 2A and Supplemental Figure S2D), Sir2-GFP spots were tracked over time in ImageJ to extract spot coordinates along 2D trajectories spanning 30–50 time points (4–7 min movies). For each trajectory, the MSD was computed as a function of the time frame between trajectory points (Michalet and Berglund, 2012). The presented MSD curve is averaged over multiple telomere trajectories (WT G1 cells: n = 54; WT quiescent cells: n = 127; esc1Δ quiescent cells: n = 62).
Western blot
Strains were grown overnight in YPDA medium and then diluted to OD600 = 0.1. Cells (108) were harvested in exponential growth (12 h) or after 7 d, and 200 μl of TCA 20%, 500 μl of acid-washed glass beads, and 200 μl of TCA buffer (20 mM Tris-HCl, pH 8, 50 mM ammonium acetate, 2 mM EDTA, 1 mM phenylmethylsulfonyl fluoride (PMSF), 1 μl of protease inhibitor cocktail [Sigma-Aldrich]) was added. Two 1-min rounds of vortexing were done to disrupt the cells. Samples were centrifuged at 14,000 rpm for 30 min at 4°C, and pellets were resuspended in 85 μl of SDS–PAGE sample buffer (120 mM Tris base, 3.5% SDS, 8 mM EDTA, 5% β-mercaptoethanol, 1 mM PMSF, 15% glycerol, 0.01% bromophenol blue). Extracts were boiled for 10 min and centrifuged at 14,000 rpm for 10 min. For immunoblotting, we used anti-Sir3 polyclonal antibody (1/200 dilution; yN-20; Santa Cruz Biotechnology, Dallas, TX). Loading was normalized according to anti-Ade13 antibodies (1:300,000; Escusa et al., 2006).
Supplementary Material
Acknowledgments
We express our profound gratitude to J. P. Javerzat for valuable comments about our work and his great help with FISH experiments. We thank A. Devin for her technical assistance with spheroplasts. We are grateful to Edward Louis for providing us the plasmid pEL42H10 for Y′-FISH experiments and to C. Boone for providing a yeast strain. We thank A. Taddei for sharing results and J. P. Javerzat, B. Daignan-Fornier, and O. Gadal for their critical comments on the manuscript. S.T. was funded by the Wellcome Trust Centre for Cell Biology. This work was supported by the University of Bordeaux and the Centre National de la Recherche Scientifique.
Abbreviations used:
- 2D
two-dimensional
- 3D
three-dimensional
- BSA
bovine serum albumin
- CHX
cycloheximide
- DAPI
4′,6-diamidino-2-phenylindole
- FISH
fluorescence in situ hybridization
- GFP
green fluorescent protein
- MSD
mean square displacement
- PBS
phosphate-buffered saline
- PMSF
phenylmethylsulfonyl fluoride
- RFP
red fluorescent protein
- SPB
spindle pole body
- WT
wild type.
Footnotes
This article was published online ahead of print in MBoC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E16-01-0069) on April 27, 2016.
REFERENCES
- Albert B, Léger-Silvestre I, Normand C, Gadal O. Nuclear organization and chromatin dynamics in yeast: biophysical models or biologically driven interactions. Biochim Biophys Acta. 2012;1819:468–481. doi: 10.1016/j.bbagrm.2011.12.010. [DOI] [PubMed] [Google Scholar]
- Allen C, Büttner S, Aragon AD, Thomas JA, Meirelles O, Jaetao JE, Benn D, Ruby SW, Veenhuis M, Madeo F, et al. Isolation of quiescent and nonquiescent cells from yeast stationary-phase cultures. J Cell Biol. 2006;174:89–100. doi: 10.1083/jcb.200604072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnaudo N, Fernández IS, McLaughlin SH, Peak-Chew SY, Rhodes D, Martino F. The N-terminal acetylation of Sir3 stabilizes its binding to the nucleosome core particle. Nat Struct Mol Biol. 2013;20:1119–1121. doi: 10.1038/nsmb.2641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ayscough KR, Stryker J, Pokala N, Sanders M, Crews P, Drubin DG. High rates of actin filament turnover in budding yeast and roles for actin in establishment and maintenance of cell polarity revealed using the actin inhibitor latrunculin-A. J Cell Biol. 1997;137:399–416. doi: 10.1083/jcb.137.2.399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bystricky K, Heun P, Gehlen L, Langowski J, Gasser SM. Long-range compaction and flexibility of interphase chromatin in budding yeast analyzed by high-resolution imaging techniques. Proc Natl Acad Sci USA. 2004;101:16495–16500. doi: 10.1073/pnas.0402766101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bystricky K, Laroche T, van Houwe G, Blaszczyk M, Gasser SM. Chromosome looping in yeast: telomere pairing and coordinated movement reflect anchoring efficiency and territorial organization. J Cell Biol. 2005;168:375–387. doi: 10.1083/jcb.200409091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell RE, Tour O, Palmer AE, Steinbach PA, Baird GS, Zacharias DA, Tsien RY. A monomeric red fluorescent protein. Proc Natl Acad Sci USA. 2002;99:7877–7882. doi: 10.1073/pnas.082243699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carmen AA, Milne L, Grunstein M. Acetylation of the yeast histone H4 N terminus regulates its binding to heterochromatin protein SIR3. J Biol Chem. 2002;277:4778–4781. doi: 10.1074/jbc.M110532200. [DOI] [PubMed] [Google Scholar]
- Cavalli G, Misteli T. Functional implications of genome topology. Nat Struct Mol Biol. 2013;20:290–299. doi: 10.1038/nsmb.2474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coller HA. Cell biology. The essence of quiescence. Science. 2011;334:1074–1075. doi: 10.1126/science.1216242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Virgilio C. The essence of yeast quiescence. FEMS Microbiol Rev. 2012;36:306–339. doi: 10.1111/j.1574-6976.2011.00287.x. [DOI] [PubMed] [Google Scholar]
- Escusa S, Camblong J, Galan J-M, Pinson B, Daignan-Fornier B. Proteasome- and SCF-dependent degradation of yeast adenine deaminase upon transition from proliferation to quiescence requires a new F-box protein named Saf1p. Mol Microbiol. 2006;60:1014–1025. doi: 10.1111/j.1365-2958.2006.05153.x. [DOI] [PubMed] [Google Scholar]
- Gotta M, Laroche T, Formenton A, Maillet L, Scherthan H, Gasser SM. The clustering of telomeres and colocalization with Rap1, Sir3, and Sir4 proteins in wild-type Saccharomyces cerevisiae. J Cell Biol. 1996;134:1349–1363. doi: 10.1083/jcb.134.6.1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guacci V, Hogan E, Koshland D. Centromere position in budding yeast: evidence for anaphase A. Mol Biol Cell. 1997;8:957–972. doi: 10.1091/mbc.8.6.957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guidi M, Ruault M, Marbouty M, Loïodice I, Cournac A, Billaudeau C, Hocher A, Mozziconacci J, Koszul R, Taddei A. Spatial reorganization of telomeres in long-lived quiescent cells. Genome Biol. 2015;16:206. doi: 10.1186/s13059-015-0766-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajjoul H, Mathon J, Ranchon H, Goiffon I, Mozziconacci J, Albert B, Carrivain P, Victor JM, Gadal O, Bystricky K, Bancaud A. High-throughput chromatin motion tracking in living yeast reveals the flexibility of the fiber throughout the genome. Genome Res. 2013;23:1829–1838. doi: 10.1101/gr.157008.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hediger F, Neumann FR, Van Houwe G, Dubrana K, Gasser SM. Live imaging of telomeres: yKu and Sir proteins define redundant telomere-anchoring pathways in yeast. Curr Biol. 2002;12:2076–2089. doi: 10.1016/s0960-9822(02)01338-6. [DOI] [PubMed] [Google Scholar]
- Heun P, Laroche T, Shimada K, Furrer P, Gasser SM. Chromosome dynamics in the yeast interphase nucleus. Science. 2001;294:2181–2186. doi: 10.1126/science.1065366. [DOI] [PubMed] [Google Scholar]
- Hozé N, Ruault M, Amoruso C, Taddei A, Holcman D. Spatial telomere organization and clustering in yeast Saccharomyces cerevisiae nucleus is generated by a random dynamics of aggregation-dissociation. Mol Biol Cell. 2013;24:1791–1800. doi: 10.1091/mbc.E13-01-0031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaspersen SL, Giddings TH, Winey M. Mps3p is a novel component of the yeast spindle pole body that interacts with the yeast centrin homologue Cdc31p. J Cell Biol. 2002;159:945–956. doi: 10.1083/jcb.200208169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin Q, Trelles-Sticken E, Scherthan H, Loidl J. Yeast nuclei display prominent centromere clustering that is reduced in nondividing cells and in meiotic prophase. J Cell Biol. 1998;141:21–29. doi: 10.1083/jcb.141.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin QW, Fuchs J, Loidl J. Centromere clustering is a major determinant of yeast interphase nuclear organization. J Cell Sci. 2000;113:1903–1912. doi: 10.1242/jcs.113.11.1903. [DOI] [PubMed] [Google Scholar]
- Joyner RP, Tang JH, Helenius J, Dultz E, Brune C, Holt LJ, Huet S, Müller DJ, Weis K. A glucose-starvation response regulates the diffusion of macromolecules. eLife. 2016;5:e09376. doi: 10.7554/eLife.09376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koszul R, Kim KP, Prentiss M, Kleckner N, Kameoka S. Meiotic chromosomes move by linkage to dynamic actin cables with transduction of force through the nuclear envelope. Cell. 2008;133:1188–1201. doi: 10.1016/j.cell.2008.04.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kupiec M. Biology of telomeres: lessons from budding yeast. FEMS Microbiol Rev. 2014;38:144–171. doi: 10.1111/1574-6976.12054. [DOI] [PubMed] [Google Scholar]
- Laporte D, Coffman VC, Lee I-J, Wu J-Q. Assembly and architecture of precursor nodes during fission yeast cytokinesis. J Cell Biol. 2011;192:1005–1021. doi: 10.1083/jcb.201008171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laporte D, Courtout F, Salin B, Ceschin J, Sagot I. An array of nuclear microtubules reorganizes the budding yeast nucleus during quiescence. J Cell Biol. 2013;203:585–594. doi: 10.1083/jcb.201306075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laporte D, Sagot I. Microtubules move the nucleus to quiescence. Nucleus. 2014;5:113–118. doi: 10.4161/nucl.28538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laporte D, Salin B, Daignan-Fornier B, Sagot I. Reversible cytoplasmic localization of the proteasome in quiescent yeast cells. J Cell Biol. 2008;181:737–745. doi: 10.1083/jcb.200711154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohr D, Ide G. Comparison on the structure and transcriptional capability of growing phase and stationary yeast chromatin: a model for reversible gene activation. Nucleic Acids Res. 1979;6:1909–1927. doi: 10.1093/nar/6.5.1909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Louis EJ, Borts RH. A complete set of marked telomeres in Saccharomyces cerevisiae for physical mapping and cloning. Genetics. 1995;139:125–136. doi: 10.1093/genetics/139.1.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall WF, Straight A, Marko JF, Swedlow J, Dernburg A, Belmont A, Murray AW, Agard DA, Sedat JW. Interphase chromosomes undergo constrained diffusional motion in living cells. Curr Biol. 1997;7:930–939. doi: 10.1016/s0960-9822(06)00412-x. [DOI] [PubMed] [Google Scholar]
- Michalet X, Berglund AJ. Optimal diffusion coefficient estimation in single-particle tracking. Phys Rev E Stat Nonlin Soft Matter Phys. 2012;85:061916. doi: 10.1103/PhysRevE.85.061916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ngubo M, Kemp G, Patterton HG. Nano-electrospray tandem mass spectrometric analysis of the acetylation state of histones H3 and H4 in stationary phase in Saccharomyces cerevisiae. BMC Biochem. 2011;12:34. doi: 10.1186/1471-2091-12-34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nguyen HQ, Bosco G. Gene positioning effects on expression in eukaryotes. Annu Rev Genet. 2015;49:627–646. doi: 10.1146/annurev-genet-112414-055008. [DOI] [PubMed] [Google Scholar]
- Nishikawa S-I, Terazawa Y, Nakayama T, Hirata A, Makio T, Endo T. Nep98p is a component of the yeast spindle pole body and essential for nuclear division and fusion. J Biol Chem. 2003;278:9938–9943. doi: 10.1074/jbc.M210934200. [DOI] [PubMed] [Google Scholar]
- O’Farrell PH. Quiescence: early evolutionary origins and universality do not imply uniformity. Philos Trans R Soc Lond B Biol Sci. 2011;366:3498–3507. doi: 10.1098/rstb.2011.0079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Onishi M, Liou G-G, Buchberger JR, Walz T, Moazed D. Role of the conserved Sir3-BAH domain in nucleosome binding and silent chromatin assembly. Mol Cell. 2007;28:1015–1028. doi: 10.1016/j.molcel.2007.12.004. [DOI] [PubMed] [Google Scholar]
- Oppikofer M, Kueng S, Martino F, Soeroes S, Hancock SM, Chin JW, Fischle W, Gasser SM. A dual role of H4K16 acetylation in the establishment of yeast silent chromatin. EMBO J. 2011;30:2610–2621. doi: 10.1038/emboj.2011.170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palladino F, Laroche T, Gilson E, Axelrod A, Pillus L, Gasser SM. SIR3 and SIR4 proteins are required for the positioning and integrity of yeast telomeres. Cell. 1993;75:543–555. doi: 10.1016/0092-8674(93)90388-7. [DOI] [PubMed] [Google Scholar]
- Piñon R. Folded chromosomes in non-cycling yeast cells: evidence for a characteristic G0 form. Chromosoma. 1978;67:263–274. doi: 10.1007/BF02569039. [DOI] [PubMed] [Google Scholar]
- Ruault M, De Meyer A, Loïodice I, Taddei A. Clustering heterochromatin: Sir3 promotes telomere clustering independently of silencing in yeast. J Cell Biol. 2011;192:417–431. doi: 10.1083/jcb.201008007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rutledge MT, Russo M, Belton J-M, Dekker J, Broach JR. The yeast genome undergoes significant topological reorganization in quiescence. Nucleic Acids Res. 2015;43:8299–8313. doi: 10.1093/nar/gkv723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sagot I, Pinson B, Salin B, Daignan-Fornier B. Actin bodies in yeast quiescent cells: an immediately available actin reserve. Mol Biol Cell. 2006;17:4645–4655. doi: 10.1091/mbc.E06-04-0282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schäfer G, McEvoy CRE, Patterton H-G. The Saccharomyces cerevisiae linker histone Hho1p is essential for chromatin compaction in stationary phase and is displaced by transcription. Proc Natl Acad Sci USA. 2008;105:14838–14843. doi: 10.1073/pnas.0806337105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schober H, Kalck V, Vega-Palas MA, Van Houwe G, Sage D, Unser M, Gartenberg MR, Gasser SM. Controlled exchange of chromosomal arms reveals principles driving telomere interactions in yeast. Genome Res. 2008;18:261–271. doi: 10.1101/gr.6687808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith CM, Gafken PR, Zhang Z, Gottschling DE, Smith JB, Smith DL. Mass spectrometric quantification of acetylation at specific lysines within the amino-terminal tail of histone H4. Anal Biochem. 2003;316:23–33. doi: 10.1016/s0003-2697(03)00032-0. [DOI] [PubMed] [Google Scholar]
- Strunnikov AV, Hogan E, Koshland D. SMC2, a Saccharomyces cerevisiae gene essential for chromosome segregation and condensation, defines a subgroup within the SMC family. Genes Dev. 1995;9:587–599. doi: 10.1101/gad.9.5.587. [DOI] [PubMed] [Google Scholar]
- Taddei A, Hediger F, Neumann FR, Bauer C, Gasser SM. Separation of silencing from perinuclear anchoring functions in yeast Ku80, Sir4 and Esc1 proteins. EMBO J. 2004;23:1301–1312. doi: 10.1038/sj.emboj.7600144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taddei A, Schober H, Gasser SM. The budding yeast nucleus. Cold Spring Harb Perspect Biol. 2010;2:a000612. doi: 10.1101/cshperspect.a000612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Therizols P, Duong T, Dujon B, Zimmer C, Fabre E. Chromosome arm length and nuclear constraints determine the dynamic relationship of yeast subtelomeres. Proc Natl Acad Sci USA. 2010;107:2025–2030. doi: 10.1073/pnas.0914187107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Therizols P, Fairhead C, Cabal GG, Genovesio A, Olivo-Marin J-C, Dujon B, Fabre E. Telomere tethering at the nuclear periphery is essential for efficient DNA double strand break repair in subtelomeric region. J Cell Biol. 2006;172:189–199. doi: 10.1083/jcb.200505159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trelles-Sticken E, Loidl J, Scherthan H. Bouquet formation in budding yeast: initiation of recombination is not required for meiotic telomere clustering. J Cell Sci. 1999;112:651–658. doi: 10.1242/jcs.112.5.651. [DOI] [PubMed] [Google Scholar]
- Wellinger RJ, Zakian VA. Everything you ever wanted to know about Saccharomyces cerevisiae telomeres: beginning to end. Genetics. 2012;191:1073–1105. doi: 10.1534/genetics.111.137851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilkins BJ, Rall NA, Ostwal Y, Kruitwagen T, Hiragami-Hamada K, Winkler M, Barral Y, Fischle W, Neumann H. A cascade of histone modifications induces chromatin condensation in mitosis. Science. 2014;343:77–80. doi: 10.1126/science.1244508. [DOI] [PubMed] [Google Scholar]
- Wong H, Marie-Nelly H, Herbert S, Carrivain P, Blanc H, Koszul R, Fabre E, Zimmer C. A predictive computational model of the dynamic 3D interphase yeast nucleus. Curr Biol. 2012;22:1881–1890. doi: 10.1016/j.cub.2012.07.069. [DOI] [PubMed] [Google Scholar]
- Yang CH, Lambie EJ, Hardin J, Craft J, Snyder M. Higher order structure is present in the yeast nucleus: autoantibody probes demonstrate that the nucleolus lies opposite the spindle pole body. Chromosoma. 1989;98:123–128. doi: 10.1007/BF00291048. [DOI] [PubMed] [Google Scholar]
- Zimmer C, Fabre E. Principles of chromosomal organization: lessons from yeast. J Cell Biol. 2011;192:723–733. doi: 10.1083/jcb.201010058. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.