Fig. 3.
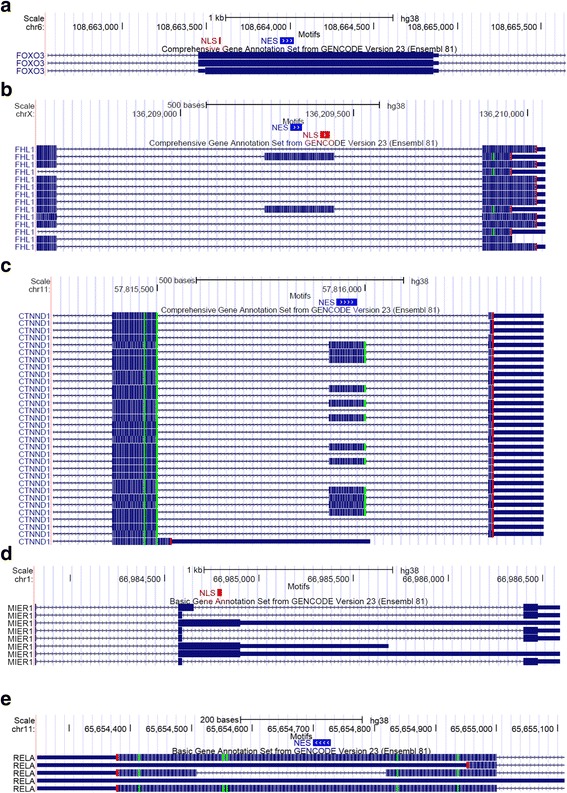
Examples of constitutive and alternative splicing-regulated NLSs and NESs. Screenshots of the UCSC Genome Browser displaying portions of transcripts encoding protein targeting motifs. NLSs and NESs are represented by red and blue blocks respectively, in the Motifs track. The Comprehensive Gene Annotation Set from GENCODE Version 23 (Ensembl 81) track shows the position of exons (blocks) and introns (lines with arrows showing direction of transcription) in the chromosome window considered. Coding exons are represented by thick blocks whereas non-coding exons (UTRs or non-coding transcripts) are represented by thin blocks. a Constitutive NLS and NES in FOXO3 gene. b FHL1 cassette exon containing NES and NLS. c Cassette exon of CTNND1 gene, containing NES. d MIER1 NLS encoded in exon regulated by alternative 5′ splice site. e RELA NES encoded in portion of exon that can be spliced out (intronic exon)
