Fig. 4.
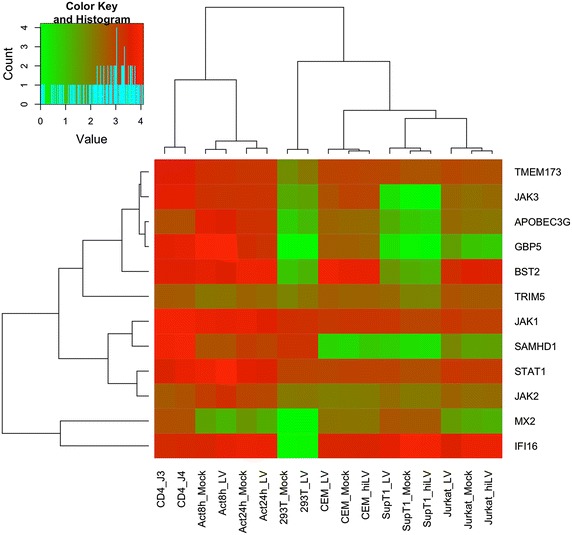
Heatmap of expression values of paradigmatic genes involved in antiretroviral defense and signaling relevant to HIV biology. The figure shows the expression values of antiretroviral genes (APOBEC3G, TRIM5, BST2, MX2, GBP5, and SAMHD1) and signaling genes (JAK, STAT1, IFI16 and STING/TMEM173) in RNA-seq libraries of resting CD4+ T cells, cell lines HEK293T, Jurkat, SupT1 and CEM -mock (Mock), heat-inactivated (hiLV) and HIV-infected (LV)- and activated CD4+ T cells at 8 and 24 h after TCR activation (Methods). The color-code scale in the inset represents the expression levels (log10 transformation of the number of library size-normalized reads per kilobase of exonic sequence), ranging from green (low) to red (high) expression. Complete hierarchical clustering of genes was based on Pearson correlation of the expression levels. Complete hierarchical clustering of samples was kept as assessed in Additional file 6: Figure S4. JAK1, JAK2, TMEM173 and GBP5 had a fold change higher than 2 between resting CD4+ T cells and permissive cell lines (Benjamini–Hochberg adjusted p value <0.01, Methods)
