Figure 5.
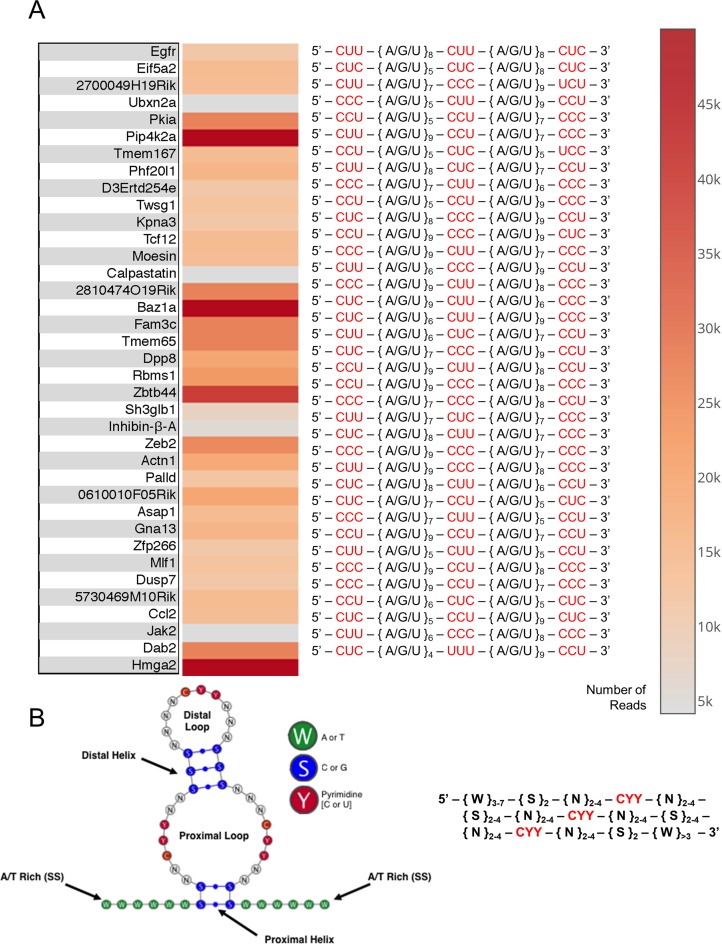
Genomic exonuclease ION torrent analysis reveals 37 consensus motifs with conserved pyrimidine residues interspaced by variable structural regions. (A) Genes that were identified to be bound to hnRNP E1 and protected from exonuclease digestion. Heat map indicates number of times a sequenced read was mapped to this parent mRNA. Annotated nucleic acid sequences show conserved pyrimidine residues highlighted in red, interspaced regions annotated to better show alignment. Parent genes were determined for each sequenced read by a custom Python script and custom 3′-UTR database containing mouse genomic 3′-UTR sequences from (UTRdb). (B) Structural depiction of consensus motif predicted by custom Python structural prediction algorithm, generalized hnRNP E1 specific binding motif descriptor. Descriptor sequence was inferred by a statistical comparison of all sequenced reads, followed by evolutionary conservation analysis, performed by a custom Python script.