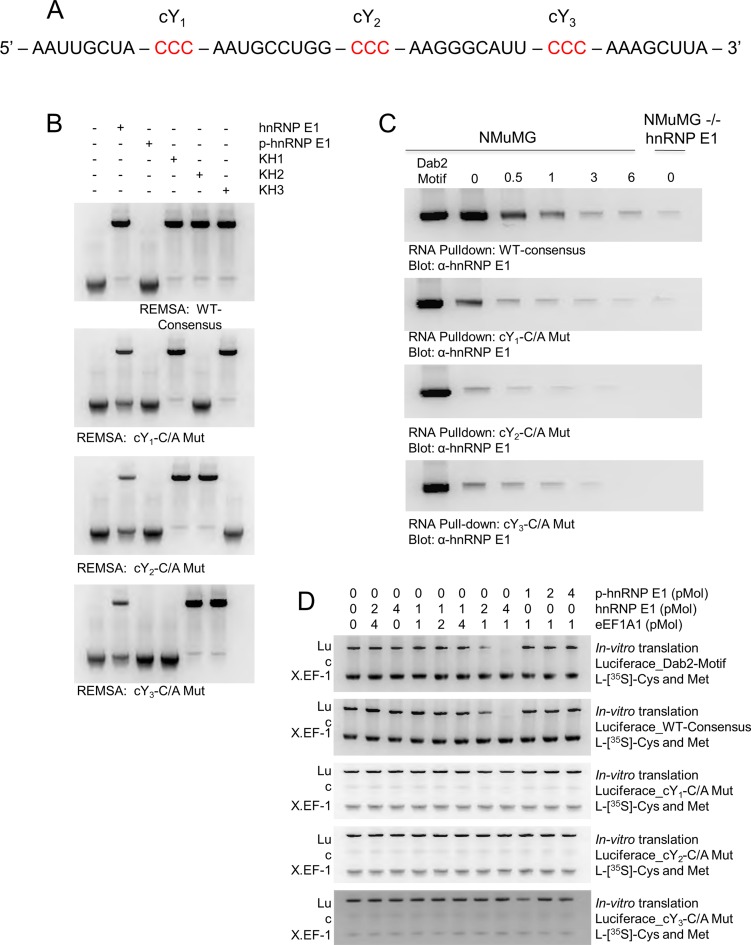
Figure 6.
Conservation of descriptor motif features in randomized synthetic RNA maintains regulatory motif properties in protein translation. (A) A randomized hnRNP E1-specific consensus motif generated by a custom Python script capable of randomizing oligonucleotides based on descriptor constraints. This sequence was used as the ‘WT-Consensus’ RNA for experiments contained in this figure. (B) RNA electromobility shift assay (REMSA) for WT-Consensus and subsequent mutations at conserved pyrimidine residues. Recombinant K-homology (KH) domains of hnRNP E1 were used to determine interactions with conserved motif regions. The REMSA data presented are typical of three independent experiments demonstrating similar binding trends (data not shown). (C) RNA pull-down analysis of WT-consensus motif compared to a bonafide Dab2 Motif element. Cellular lysate from NMuMG cells were treated with 5 ng/ml TGFβ for up to 6 h, hnRNP E1 interactions were measured through pull-down by various synthetic RNA constructs. Pull-down assays were performed a minimum of three times, similar binding trends were noted in each replicate (data not shown). D)In vitro translation assay of luciferase mRNA coupled to various 3′-UTR regulatory elements. Luciferase_Dab2-Motif mRNA was used as a control to compare strength of translation inhibition amongst WT-consensus and its various mutations. These assays were performed a minimum of three times, measurements were taken using luminescent units and similar trends were noted among replicates. For visual interpretation, we present radiolabeled samples analyzed by polyacrylamide gel electrophoresis.