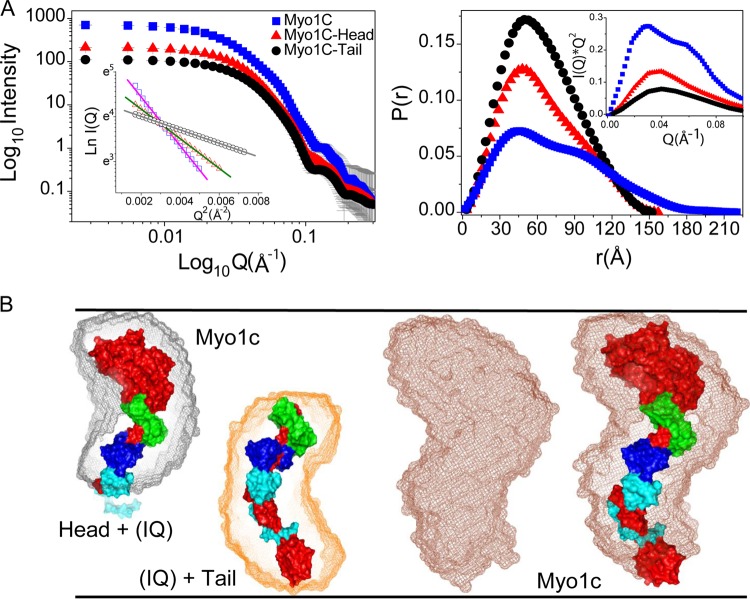
FIG 2.
SAXS data of unliganded protein samples. (A) (Left) SAXS intensity profiles acquired from the solutions of Myo1c-FL and its head and tail domains are presented. The inset shows linear fit to the Guinier region of the measured data sets. (Right) The P(r) curves computed for Myo1c-FL and its head and tail domains demonstrate the frequency distribution of interatomic vectors in the predominant scattering species. The inset shows the Kratky plots of the data sets. (B) Scattering shape of the proteins restored from dummy atom modeling using the SAXS data as a reference. The envelope shape (represented as mesh) of the predominant shape computed for full-length Myo1c and its head and tail domains is shown. Crystal structures of the head, tail, and full-length Myo1c (reconstructed by joining head and tail regions) with bound calmodulins (represented as surface) were overlaid by automated alignment of inertial axes with SAXS models. The structure of Myo1c is represented in red, while three calmodulins are represented in green, blue, and cyan.