ABSTRACT
(MAMP)-triggered immunity (MTI) is the first layer of molecular defense encountered by pathogens. Genetic screens have contributed to our knowledge of MTI, but are limited to phenotype-causing mutations. Here we attempt to identify novel factors involved in the early event leading to plant MTI by comparing the nuclear proteomes of two Arabidopsis genotypes treated with chitosan. Our approach revealed that following chitosan treatment, cerk1 plants had many nuclear accumulating proteins in common, but also some unique ones, when compared with Col-0 plants. Analysis of the identified proteins revealed a nuclear accumulation of DNA-modifying enzymes, RNA-binding proteins and ribosomal proteins. Our results demonstrate that nuclear proteomic is a valid, phenotype-independent approach to uncover factor involved in cellular processes.
KEYWORDS: Arabidopsis, cerk1, chitosan, MAMP triggered immunity, nucleus, proteomic
Plants have evolved a multilayered system to detect and defend against potentially harmful pathogenic microbes. Beyond structural defenses, the first molecular layer is composed of transmembrane pattern recognition receptors (PRR) that detect slowly-evolving microbial components.1 These microbe-associated molecular patterns (MAMPs), also known as pathogen-associated molecular patterns (PAMPs) include, among many others, the bacterial flagellin (flg22) and elongation factor Tu.2 MAMP recognition by PRRs triggers ion fluxes, oxidative bursts3 and mitogen-activated protein kinase (MAPK) pathways activation4 leading to the transcriptional reprogramming of over 1,200 genes5 and to the induction of required basal defense responses.1 The importance of MTI is best illustrated by the pressure exerted by the pathogen to suppress it. One striking example is the HopF2 effector which directly supresses MTI at two different levels of the MAMP-activated MAPK cascades. It can directly target BAK1, which is required for the full elicitation of pathogen-induced defense responses,6 at the plasma membrane, thereby acting upstream of the MEKK1-MKK1/2- MPK4 pathway. It can also directly block MKK5 of the MEKK-MKK4/5-MPK3/6 cascade.6
The chitin receptor is one of the MAMP receptors that has been investigated with some success. Chitin, a major component of the fungal cell wall, is a β-1,4-linked N-acetyl-glucosamine polymer that has long been recognized as a potent MAMP in plant-fungus interactions.7 In Arabidopsis, it is mostly detected by the CHITIN-ELICITED RECEPTOR KINASE 1 (CERK1). cerk1 knock-out plants lose their response to chitin elicitor, including MAPK activation, reactive oxygen species (ROS) generation and induction of gene expression.8 Indeed, CERK1 phosphorylates after exposure to chitin or chitosan (acetylated chitin) and can homodimerize when binding to chitin monomers to activate its kinase domain.9 However, chitin signaling seems to require co-receptors: two additional LysM receptor kinases, AtLYK4 and AtLYK5, are also involved in chitin recognition.10,11 Supporting the co-receptor theory is the fact that AtLYK5 binds chitin with high affinity and can dimerize with CERK1 in a chitin-dependent manner.10 Other receptors may also be implicated but are masked by the dominant effect of CERK1.
Despite the importance of MTI, the intracellular modulation that takes place after MAMP recognition, which involves transcriptional reprogramming, is still somewhat unclear. More precisely, the chitin-elicited nuclear proteins involved in the establishment of basal defense responses are not fully known. Two MAPK pathways have been shown to be activated downstream of MAMP signaling. One elicits the activation of the MAPKs MPK3 and MPK64 and the second leads to MPK4 activation.12 Recently, MPK1, MPK11 and MPK13 were also found to be phosphorylated upon flg22 treatment.13 The absence of MTI defect in these three MAPKs knockout lines suggests functional redundancy, so many more components acting downstream of receptor activation may be missed in phenotype-based screening.
In the present study, we sought to discover proteins that participate in MTI but have escaped phenotype-based screening. Toward this end, we took an unbiased approach based on protein mass spectrometry (MS) of the nuclear proteome of young Arabidopsis plants subjected or not to chitosan treatment. Chitosan is known to also bind CERK19 and triggers a transcriptional response that overlaps with the response to chitin.14 Using high performance liquid chromatography-electrospray ionization tandem mass spectrometry (HPLC-ESI-MS-MS), we identified several plant proteins that accumulate in the nucleus exclusively after chitosan treatment of Arabidopsis Columbia-0 (Col-0) or cerk1 plants.
Before proceeding with the nuclear proteome MS analysis, we assessed if chitosan treatment was efficient in triggering a MAMP-like response. Three genes that are among the most up-regulated after chitin treatment15 were analyzed by RT-qPCR: At2g37430 (C2H2-ZF), At1g22810 (AP2/ERE) and At2g44840 (AP2/ERE). All three genes were upregulated after chitosan treatment of Col-0 plants, showing respectively 13-fold, 51-fold and 3-fold induction 15 min post-treatment (Fig. 1A). We also observed that At1g22810 was slightly upregulated following chitosan treatment of cerk1 plants albeit at much lower level than in Col-0 (5-fold).
Figure 1.
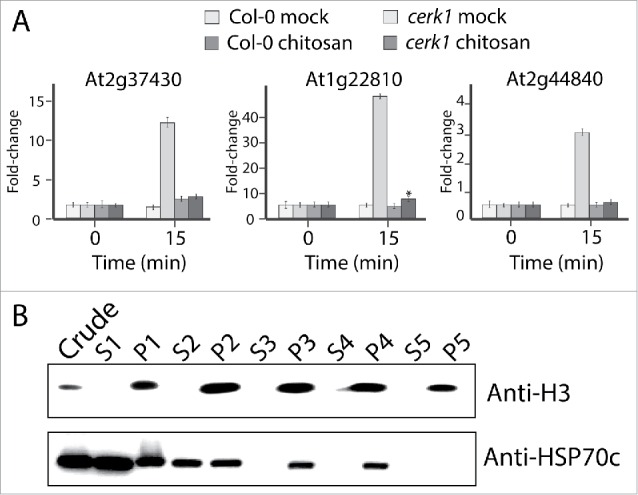
Chitosan treatment elicits MTI responsive gene in planta. (A) Expression of the MAMP-triggered immunity responsive marker genes At2g37430, At1g22810, At2g44840 in Col-0 and cerk1 following chitosan or mock treatment. Q-RT-PCR was performed on soil grown three-weeks-old plants. ACT1 was used to normalize the transcript levels. (B) Quality control of the fractionation procedure by western blotting using HSP70c and histone H3 as cytosolic and nuclear markers respectively. Crude indicates crude extract, S = supernatant, P = pellet and number indicate the wash number.
We assessed the purity of our nuclear fractions by using the cytosolic marker HSP70c and nuclear marker histone H3. HSP70c could not be detected by Western blotting in the nuclear fraction corresponding to pellet five, while the nuclear marker anti-histone H3 was still clearly visible, hence this nuclear fraction was sent for mass spectrometry analysis. Tandem MS identified 1,372 different Arabidopsis proteins among a total of 31,416 spectra from our eight samples (duplicates of cerk1 or Col-0 plants treated or not with chitosan) (PRIDE repository with the dataset identifier PXD003821 and 10.6019/PXD003821). We set very conservative criteria for our analyses: all proteins identified needed a minimum of two spectra to be considered, and all proteins that were present in only one of the duplicates were also rejected.
Our first analysis of the proteomic results was to compare the functional categorization of the 232 proteins found in the nucleus after chitosan treatment (in Col-0 and cerk1) with the 182 proteins from the nuclear proteome of cold-treated plants, one of the few studies of Arabidopsis nuclear proteomes that can relate to our investigation.16 In parallel, we performed similar analysis with the SUBA database using only proteins predicted to be nuclear by SUBA bioinformatics tools or confirmed to be nuclear by GFP-tagging (total of 4,421 proteins). Finally, we compared our data to findings on the cytosolic proteome published by Ito et al. (2011) (Fig. 2A). The first observation from this categorization based on predicted cellular components is that only 26% of the nuclear proteins from the SUBA data set were annotated as nuclear proteins by TAIR's gene ontology (GO) annotator (Fig. 2A). In other words, the remaining 74% may be nuclear at some point, but the nucleus was not deemed to be their primary localization in GO. This reflects the fact that proteins may have several putative locations and underlines the weakness of bioinformatic to predict protein localization. The nuclear proteomes of chitosan and cold-treated plants contained only 11% and 16% of predicted nuclear proteins while the cytosolic experimental proteome still showed 9% of nuclear predicted proteins (Fig. 2A). Based on the discrepancies observed with the SUBA dataset, we can assume that a significant proportion of proteins annotated as non-nuclear by GO in these three experimental data sets were indeed at some point nuclear.
Figure 2.
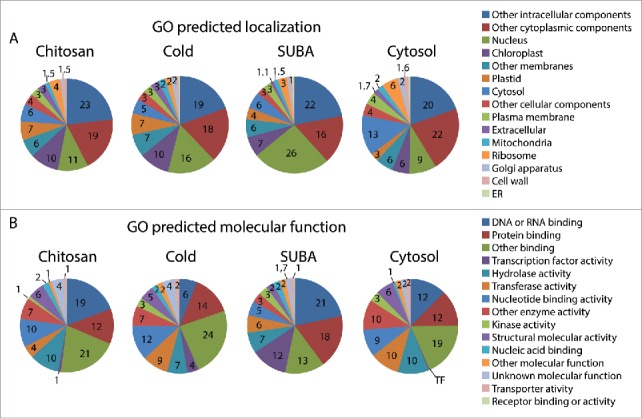
Gene ontologies in chitosan treated plants compared to other datasets. (A) GO predicted subcellular localization. (B) GO predicted molecular function.
In the search for proteins that participate in MTI, categorization by molecular function (Fig. 2B) enables us to identify proteins that have the capacity to modulate transcription or translation during defense responses. Our chitosan-induced nuclear proteome contains 19% of DNA- or RNA-binding proteins, which could alter gene expression through DNA-binding, mRNA-processing, and mRNA-export, or could impact translation through mRNA nuclear segregation. Fewer of these proteins (12%) were found in the cytosolic dataset.17 Proteins with transcription factor activity were most abundant in the SUBA nuclear data set (12%) but still represented 1%, 4% and 0.2% of proteins in chitosan, cold and cytosolic proteomes respectively, confirming that nuclear enrichment does indeed enrich transcription factors. It should also be noted that empirically-obtained proteomes are biased toward abundant proteins which could mask less abundant proteins. Therefore, signaling components such as transcription factors may be under-represented in LC-MS-MS proteomes, as demonstrated by their abundance in the SUBA dataset relative to the three other data sets.
We constructed a Venn diagram comparing the proteins found in each treatment group (Control is the combination of both Col-0 and cerk1 plants treated with water) (Fig. 3). We identified eight proteins specifically localized to the nucleus of Col-0 plants after chitosan treatment (listed in Table 1). Although most of these are not obvious MTI components, a clear trend toward ribosomal proteins and translation is obvious. Proteins 1 (S19E family ribosomal protein), 4 (ribosomal protein l6), 5 (S19E family ribosomal protein) and 7 (RNAse Z activity involved in tRNA processing) are all involved in translation. Protein 8 (DNA-binding transcriptional regulator) is engaged in transcription regulation while protein 6 (small nuclear ribonucleoprotein G) binds RNA and could be involved in either transcription or translation. Most of these proteins have been reported to be modulated at the transcription level after biotic or abiotic stress, but have not previously been linked with the MAMP response.18-21
Figure 3.
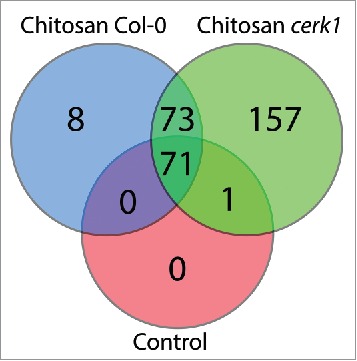
Venn diagram displaying the number of proteins identified in the nucleus for each condition.
Table 1.
Nuclear localized proteins identified by LC-MS-MS in Col-0 plants following chitosan treatment.
| Protein description | Uniprot ID | AGI |
|---|---|---|
| Ribosomal protein S19e family protein | D7KGE2 | AT5G61170 |
| HAD superfamily, subfamily IIIB acid phosphatase | Q9ZWC4 | AT1G04040 |
| Galactose mutarotase-like superfamily protein | Q8LFH1 | AT3G47800 |
| Ribosomal protein L6 family protein | Q8L9N4 | AT1G18540 |
| Ribosomal protein S19e family protein | D7MUI1 | AT5G61170 |
| Probable small nuclear ribonucleoprotein G | O82221 | AT2G23930 |
| Encodes a protein with RNase Z activity suggesting a role in tRNA processing | Q8L633 | AT2G04530 |
| DNA-binding storekeeper protein-related transcriptional regulator | O23063 | AT4G00390 |
157 proteins were only detected in the nucleus of cerk1 plants after chitosan treatment (reported in Table S2). It is striking that so many protein are unique to cerk1 as it has an impaired sensing of chitin8 and as we observed only a weak transcriptional reprogramming in our RT-qPCR results (Fig. 1). On the other hand, it is known that while chitin and chitosan responses largely overlap, 33% of chitosan elicited genes are not elicited by chitin.14 Table 2 groups the proteins possessing the molecular functions most likely to affect early MTI responses (transcription factor and DNA/RNA-binding protein) and excludes those from metabolisms. Many of those may regulate gene expression or mRNA metabolism, as several additional proteins are RNA helicases that may influence transcription or translation. Interestingly, one resistance protein of the Toll/Interleukin receptor (TIR) family (At4g16990) was found: it is known as RLM3 and is required for resistance to Leptosphaeria maculans and other necrophytic pathogens.22
Table 2.
Subset of nuclear localized proteins identified by LC-MS-MS in cerk1 plants following chitosan treatment.
| Protein description |
Uniprot ID |
AGI |
|---|---|---|
| Transcription factor or transcriptional regulator | ||
| MED16, Mediator of RNA polymerase II transcription subunit 16, positive regulation of SAR | F4JGZ1 | AT4G04920 |
| Small RNA degrading nuclease 3, regulation of transcription | F4K3N3 | AT5G67240 |
| ACT domain-containing small subunit of acetolactate synthase protein | Q93YZ7 | AT2G31810 |
| Trihelix transcription factor ASIL2, sequence-specific DNA binding transcription factors | Q9LJG8 | AT3G14180 |
| VERNALIZATION INDEPENDENCE 5, regulation of transcription, DNA binding | D7KW58 | AT1G61040 |
| Sequence-specific DNA binding transcription factors | Q8LF33 | AT3G11100 |
| Short life 1, PHD finger and BAH motif containing putative transcription factor | F4JV93 | AT4G39100 |
| Mediator of RNA polymerase II transcription subunit 32 | Q84VW5 | AT1G11760 |
| CALMODULIN-BINDING TRANSCRIPTION ACTIVATOR 2, CAMTA2 | Q6NPP4 | AT5G64220 |
| EARLY BOLTING IN SHORT DAYS, chromatin assembly or disassembly | O65462 | AT4G22140 |
| RNA-binding protein | ||
| NUCLEOSTEMIN-LIKE 1, nucleolar GTP- binding protein involved in RNA methylation | Q93Y17 | AT3G07050 |
| RPT2a encodes the 26S proteasome subunit, regulate gene silencing via DNA methylation | Q9SZD4 | AT4G29040 |
| EMBRYO DEFECTIVE 2770, RNA-directed DNA methylation, mRNA splicing | Q9ZT71 | AT4G03430 |
| Serine/arginine-rich SC35-like splicing factor | Q8L3X8 | AT3G55460 |
| RZ1B, Putative RNA-binding involved in cold tolerance | O22703 | AT1G60650 |
| WD-40 protein involved in histone deacetylation in response to abiotic stress | Q9FN19 | AT5G67320 |
| TOUGH, Interacts with TATA-box binding protein 2. RNA binding | Q8GXN9 | AT5G23080 |
| THO complex subunit 7B, component THO/TREX complex | Q9M8T6 | AT3G02950 |
| Small RNA degrading nuclease 3, regulation of transcription | F4K3N3 | AT5G67240 |
| RNA binding (RRM/RBD/RNP motifs), RNA processing | F4J9U9 | AT3G12640 |
| mRNA splicing factor, Cwf18 | Q9MAB2 | AT3G05070 |
| SWI/SNF complex subunit SWI3C, ATP-dependent chromatin-remodeling complex | Q9XI07 | AT1G21700 |
| Splicing factor U2af large subunit B, Necessary for the splicing of pre-mRNA | Q8L716 | AT1G60900 |
| Small nuclear ribonucleoprotein | Q9SUM2 | AT4G30220 |
| Small nuclear ribonucleoprotein family protein, mRNA splicing | Q9C6K5 | AT1G76860 |
| nuclear cap-binding protein, mRNA metabolism | Q9XFD1 | AT5G44200 |
| RNA-binding protein-related | F4JM55 | AT4G28990 |
We also analyzed the proteins common between Col-0 and cerk1 nuclei after chitosan treatment (presented at the intersection in Fig. 3). A total of 73 proteins were identified and most of these were either DNA/RNA-binding proteins or ribosomal proteins. Table 3 shortlists the proteins sorted by molecular function, uncovering several DNA/RNA-binding proteins linked with chromatin remodeling and RNA maturation (see full list in Table S3). Receptor for activated C kinase 1 A (RACK1A) was one of the few proteins in Table 3 that was neither ribosomal nor DNA/RNA-binding. This protein was recently shown to act as a scaffold protein in a new immune signaling pathway.23 The subset common between the two genotypes (Col-0 and cerk1) and the two treatments (water and chitosan) – those at the intersection of the three circles (listed in Table S4) – mostly contained proteins from the chloroplasts and mitochondria as well as many enzymes from primary metabolism that likely contaminated the nuclear preparations, which explains that they were found in all genotypes and treatments. This set of proteins also contained some constitutive nuclear components, such as nucleoporins, spliceosome assembly proteins and polymerases, but few RNA- or DNA-binding proteins and ribosomal components, strengthening the results obtained in chitosan-treated plants in which we observed some specificity among RNA- or DNA-binding proteins and ribosomal components.
Table 3.
Subset of nuclear localized proteins identified by LC-MS-MS in both cerk1 AND Col-0 plants following chitosan treatment.
| Protein description |
Uniprot ID |
AGI |
|---|---|---|
| Miscellanous | ||
| Homologous to the co-chaperon DNAJ protein | Q94AW8 | AT3G44110 |
| EPITHIOSPECIFIER MODIFIER 1, defense response to bacterium | Q9LJG3 | AT3G14210 |
| RECEPTOR FOR ACTIVATED C KINASE 1 A, MAP-kinase scaffold activity | O24456 | AT1G18080 |
| DNA-RNA binding proteins | ||
| Nuclear RNA binding protein A-like protein | Q8LDQ7 | AT5G47210 |
| GLYCINE-RICH RNA-BINDING PROTEIN 7, DNA binding, RNA binding | C0Z2N6 | AT2G21660 |
| mRNA splicing factor | B3H6J5 | AT3G49601 |
| RNA BINDING PROTEIN, RNA modification, RNA processing, RNA stabilization | Q04836 | AT4G24770 |
| RNA polymerase I-associated factor PAF67 | F4JY76 | AT5G25754 |
| ATWTF1, RNA recognition domain | A0MFS5 | AT4G01037 |
| GENERAL REGULATORY FACTOR 3, 14-3-3 gene | P42644 | AT5G38480 |
| COPPER RESPONSE DEFECT 1, putative ZIP protein, DNA binding | Q9M591 | AT3G56940 |
| Histone deacetylase HDT2 | Q56WH4 | AT5G22650 |
| MAR-binding filament-like protein 1, DNA-binding protein | Q9LW85 | AT3G16000 |
| Nucleosome assembly protein 1-like 1 | B3H684 | AT4G26110 |
| Emsy N Terminus and plant Tudor-like domain, defense response to fungus | Q9C7C4 | AT3G12140 |
| Histone deacetylase HD2A | F4J378 | AT3G44750 |
| Serine/arginine-rich SC35-like splicing factor | Q9LHP2 | AT3G13570 |
| U2 SMALL NUCLEAR RIBONUCLEOPROTEIN B, splicing | O22922 | AT2G30260 |
| DEK domain-containing chromatin associated protein | Q84JB7 | AT5G63550 |
| ATGRP8, glycine-rich protein with RNA binding domain at the N-terminus. | B9DFJ8 | AT4G39260 |
| MLP-LIKE PROTEIN 423, defense response, mRNA modification | Q93VR4 | AT1G24020 |
| Involved in translation | ||
| LOS1, translation elongation factor 2 | Q9ASR1 | AT1G56070 |
| Ribosomal protein L4/L1 family | F4KDU5 | AT5G02870 |
| EMBRYO DEFECTIVE 2184, structural constituent of ribosome | Q9FWS4 | AT1G75350 |
| Eukaryotic translation initiation factor 3 subunit E | Q9C5Z3 | AT3G57290 |
| Eukaryotic translation initiation factor 3 subunit B | F4K4D5 | AT5G27640 |
| 40S ribosomal protein S3a-1 | Q9CAV0 | AT3G04840 |
| 40S ribosomal protein S16-3 | A8MRX2 | AT5G1838 |
| Ribosomal protein L19 | Q8W101 | AT1G02780 |
| 40S ribosomal protein S20-1 | P49200 | AT3G45030 |
| Translation elongation factor EF1B/ribosomal protein S6 | D7KNE3 | AT5G19510 |
| Elongation factor 1-β 2 | Q9SCX3 | AT5G19510 |
| Ribosomal protein S10p/S20e family protein | Q9LK61 | AT3G13120 |
| Ribosomal protein L10 family protein | B5X0P0 | AT5G13510 |
| 50S ribosomal protein L19-2 | Q8RXX5 | AT5G47190 |
| TRANSLATION INITIATION FACTOR 3 SUBUNIT H1 | Q9C5Z2 | AT1G10840 |
| RIBOSOMAL PROTEIN S10E B | Q9FFS8 | AT5G41520 |
| 60S ribosomal protein L36-2 | Q9M352 | AT3G53740 |
| 60S ribosomal protein L17-1 | Q93VI3 | AT1G27400 |
| 40S ribosomal protein S24e | Q9SS17 | AT3G04920 |
| Elongation factor 1B β | A8MRC4 | AT1G30230 |
The MTI response depends on the recognition of conserved molecular pathogen patterns at the cell surface by pathogen recognition receptors.1 Genetic screening has largely contributed to our understanding of plant defense24 and to the molecular dissection of the defense signaling pathways.25 We used HPLC-ESI-tandem MS, a phenotype-independent approach to discover components participating in the establishment of defense responses resulting from MAMP recognition.
Interestingly, proteins that were either part of the ribosome or actively participated in translation were over-represented following chitosan treatment in both genotypes (Table 1, 2, 3). Since ribosomes are assembled in the nucleus, it is not surprising to observe many ribosomal proteins in our nuclear proteomes, but it is interesting that their identity differed in different genotypes and whether the plants had been exposed to chitosan or not. It is well-known that ribosome composition is highly heterogeneous and varies during plant development to ensure translational regulation.26 Hence, we could speculate that ribosome subunits, which are highly heterogeneous,27 may disassemble and reassemble after elicitor detection and triggering of MTI. As is observed in development, such reassembly could promote MTI oriented translational regulation. Recently, JIP60, a barley protein that mediates a translational switch toward stress and defense protein synthesis in the presence of jasmonate and at senescence, was discovered.28 More recently the ribosomal coding genes RPL12 and RPL19 were shown to be involved in nonhost disease resistance in Nicotiana and Arabidopsis and also play a minor role in basal resistance against virulent pathogens.29
Another type of proteins abundantly observed in our study were DNA-modifying enzymes that have the capacity to affect chromatin remodeling and in doing so to further impact transcription. The role of chromatin remodelling proteins in regulating Arabidopsis defense responses has been reviewed by Berr et al.30 Mutation of chromatin-remodeling enzymes results in pleiotropic phenotypes not specifically associated with MTI or ETI but in which prominent players in transcriptional repression and activation at the onset of these processes are affected.
Various families of RNA-binding proteins, including proteins linked to mRNA splicing, export and maturation, were also identified after elicitation by chitosan. RNA export defects have previously been shown to suppress NB-LRR-mediated immunity,31,32 basal responses32 and response to abiotic stress,33 suggesting that even more proteins involved in RNA metabolism may participate in defense responses.
As reviewed by Boller and Felix (2009), many molecular events unfold during the first 15 min of MAMP recognition and they set a point of no return upon which cells commit to the massive transcriptional reprogramming required for the establishment of the basal response. Consequently, we chose to concentrate our analysis on early nuclear recruitment of molecular components following MAMP detection. While the MTI response is clearly dependant on MAPK pathways, our data indicate that ribosome reorganization, DNA modification and RNA maturation could play major roles during the early MAMP response. Specific proteins affecting translation or switching it to defense mode need to be investigated further. Similarly, the participation of chromatin-remodeling and RNA-modifying enzymes should be studied. Our results demonstrate that nuclear proteomic is a valid, phenotype-independent approach to uncover factors involved in various cellular processes.
Supplementary Material
Disclosure of potential conflicts of interest
No potential conflicts of interest were disclosed.
References
- 1.Boller T, Felix G. A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Ann Rev Plant Biol 2009; 60:379-406; PMID:19400727; http://dx.doi.org/15548740 10.1146/annurev.arplant.57.032905.105346 [DOI] [PubMed] [Google Scholar]
- 2.Kunze G, Zipfel C, Robatzek S, Niehaus K, Boller T, Felix G. The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 2004; 16:3496-507; PMID:15548740; http://dx.doi.org/ 10.1105/tpc.104.026765 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nurnberger T, Jones JD, Felix G, Boller T. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 2007; 448:497-500; PMID:17625569; http://dx.doi.org/ 10.1038/nature05999 [DOI] [PubMed] [Google Scholar]
- 4.Asai ST, Tena G, Plotnikova J, Willmann MR, Chiu WL, Gomez-Gomez L, Boller T, Ausubel FM, Sheen J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 2002; 415:977-83; PMID:11875555; http://dx.doi.org/ 10.1038/415977a [DOI] [PubMed] [Google Scholar]
- 5.Zipfel C, Robatzek S, Navarro L, Oakeley EJ, Jones JD, Felix G, Boller T. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 2004; 428:764-7; PMID:15085136; http://dx.doi.org/ 10.1038/nature02485 [DOI] [PubMed] [Google Scholar]
- 6.Zhou J, Wu S, Chen X, Liu C, Sheen J, Shan L, He P. The Pseudomonas syringae effector HopF2 suppresses Arabidopsis immunity by targeting BAK1. Plant J 2014; 77:235-45; PMID:24237140; http://dx.doi.org/ 10.1111/tpj.12381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wan J, Zhang XC, Stacey G. Chitin signaling and plant disease resistance. Plant Signal Behavior 2008; 3:831-3; PMID:19704513; http://dx.doi.org/18042724 10.4161/psb.3.10.5916 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Miya A, Albert P, Shinya T, Desaki Y, Ichimura K, Shirasu K, Narusaka Y, Kawakami N, Kaku H, Shibuya N. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc Natl Acad Sci U S A 2007; 104:19613-8; PMID:18042724; http://dx.doi.org/ 10.1073/pnas.0705147104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Petutschnig EK, Jones AM, Serazetdinova L, Lipka U, Lipka V. The lysin motif receptor-like kinase (LysM-RLK) CERK1 is a major chitin-binding protein in Arabidopsis thaliana and subject to chitin-induced phosphorylation. J Biol Chem 2010; 285:28902-11; PMID:20610395; http://dx.doi.org/ 10.1074/jbc.M110.116657 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cao Y, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, Joachimiak A, Stacey S. The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. Elife 2014; 3, 1–19; PMID:25340959; http://dx.doi.org/18263776 10.7554/eLife.03766 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wan J, Zhang XC, Neece D, Ramonell KM, Clough S, Kim SY, Stacey MG, Stacey G. A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 2008; 20:471-81; PMID:18263776; http://dx.doi.org/ 10.1105/tpc.107.056754 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gao M, Liu J, Bi D, Zhang Z, Cheng F, Chen S, Zhang Y. MEKK1, MKK1/MKK2 and MPK4 function together in a mitogen-activated protein kinase cascade to regulate innate immunity in plants. Cell Res 2008; 18:1190-8; PMID:18982020; http://dx.doi.org/ 10.1038/cr.2008.300 [DOI] [PubMed] [Google Scholar]
- 13.Nitta Y, Ding P, Zhang Y. Identification of additional MAP kinases activated upon PAMP treatment. Plant Signal Behavior 2014; 9:e976155; PMID:25482788; http://dx.doi.org/21240536 10.4161/15592324.2014.976155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Povero G, Loreti E, Pucciariello C, Santaniello A, Di Tommaso D, Di Tommaso G, Kapetis D, Zolezzi F, Piaggesi A, Perata P. Transcript profiling of chitosan-treated Arabidopsis seedlings. J Plant Res 2011; 124:619-29; PMID:21240536; http://dx.doi.org/ 10.1007/s10265-010-0399-1 [DOI] [PubMed] [Google Scholar]
- 15.Libault M, Wan J, Czechowski T, Udvardi M, Stacey G. Identification of 118 Arabidopsis transcription factor and 30 ubiquitin-ligase genes responding to chitin, a plant-defense elicitor. Mol Plant-Microbe Interactions 2007; 20:900-11; PMID:17722694; http://dx.doi.org/14617066 10.1094/MPMI-20-8-0900 [DOI] [PubMed] [Google Scholar]
- 16.Bae MS, Cho EJ, Choi EY, Park OK. Analysis of the Arabidopsis nuclear proteome and its response to cold stress. Plant J 2003; 36:652-63; PMID:14617066; http://dx.doi.org/ 10.1046/j.1365-313X.2003.01907.x [DOI] [PubMed] [Google Scholar]
- 17.Ito J, Batth TS, Petzold CJ, Redding-Johanson AM, Mukhopadhyay A, Verboom R, Meyer EH, Millar AH, Heazlewood JL. Analysis of the Arabidopsis cytosolic proteome highlights subcellular partitioning of central plant metabolism. J Proteome Res 2011; 10:1571-82; PMID:21166475; http://dx.doi.org/ 10.1021/pr1009433 [DOI] [PubMed] [Google Scholar]
- 18.Ascencio-Ibáñez JT, Sozzani R, Lee T-J, Chu T-M, Wolfinger RD, Cella R, Hanley-Bowdoin L. Global analysis of arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection. Plant Physiol 2008; 148:436-54; PMID:18650403; http://dx.doi.org/16776300 10.1104/pp.108.121038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ditt RF, Kerr KF, de Figueiredo P, Delrow J, Comai L, Nester EW. The Arabidopsis thaliana transcriptome in response to agrobacterium tumefaciens. Mol Plant-Microbe Interact 2006; 19:665-81; PMID:16776300; http://dx.doi.org/ 10.1094/MPMI-19-0665 [DOI] [PubMed] [Google Scholar]
- 20.Sharma N, Cram D, Huebert T, Zhou N, Parkin IAP. Exploiting the wild crucifer Thlaspi arvense to identify conserved and novel genes expressed during a plant's response to cold stress. Plant Mol Biol 2006; 63:171-84; PMID:16972165; http://dx.doi.org/ 10.1007/s11103-006-9080-4 [DOI] [PubMed] [Google Scholar]
- 21.Charron J-BF, Ouellet F, Houde M, Sarhan F. The plant Apolipoprotein D ortholog protects Arabidopsis against oxidative stress. BMC Plant Biol 2008; 8:86; PMID:18671872; http://dx.doi.org/ 10.1186/1471-2229-8-86 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Staal J, Kaliff M, Dewaele E, Persson M, Dixelius C. RLM3, a TIR domain encoding gene involved in broad-range immunity of Arabidopsis to necrotrophic fungal pathogens. Plant J 2008; 55:188-200; PMID:18397376; http://dx.doi.org/ 10.1111/j.1365-313X.2008.03503.x [DOI] [PubMed] [Google Scholar]
- 23.Cheng Z, Li JF, Niu Y, Zhang XC, Woody OZ, Xiong Y, Djonović S, Millet Y, Bush J, McConkey BJ, et al.. Pathogen-secreted proteases activate a novel plant immune pathway. Nature 2015; 521:213-6; PMID:25731164; http://dx.doi.org/ 10.1038/nature14243 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Glazebrook J, Rogers EE, Ausubel FM. Use of arabidopsis for genetic dissection of plant defense responses. Ann Rev Genetics 1997; 31:547-69; PMID:9442907; http://dx.doi.org/22682562 10.1146/annurev.genet.31.1.547 [DOI] [PubMed] [Google Scholar]
- 25.Monaghan J, Germain H, Weihmann T, Li X. Dissecting plant defense signal transduction: Modifier of snc1 in Arabidopsis. Canadian J Plant Pathol 2010; 32:35-42; http://dx.doi.org/ 10.1080/07060661003621001 [DOI] [Google Scholar]
- 26.Horiguchi G, Van Lijsebettens M, Candela H, Micol JL, Tsukaya H. Ribosomes and translation in plant developmental control. Plant Sci 2012; 191-192:24-34; PMID:22682562; http://dx.doi.org/ 10.1016/j.plantsci.2012.04.008 [DOI] [PubMed] [Google Scholar]
- 27.Giavalisco P, Wilson D, Kreitler T, Lehrach H, Klose J, Gobom J, Fucini P. High heterogeneity within the ribosomal proteins of the Arabidopsis thaliana 80S ribosome. Plant Mol Biol 2005; 57:577-91; PMID:15821981; http://dx.doi.org/ 10.1007/s11103-005-0699-3 [DOI] [PubMed] [Google Scholar]
- 28.Rustgi S, Pollmann S, Buhr F, Springer A, Reinbothe C, von Wettstein D, Reinbothe S. JIP60-mediated, jasmonate- and senescence-induced molecular switch in translation toward stress and defense protein synthesis. Proc Natl Acad Sci U S A 2014; 111:14181-6; PMID:25225401; http://dx.doi.org/ 10.1073/pnas.1415690111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nagaraj S, Senthil-Kumar M, Ramu VS, Wang K, Mysore KS. Plant ribosomal proteins, RPL12 and RPL19, play a role in nonhost disease resistance against bacterial pathogens. Front Plant Sci 2015; 6:1192; PMID:26779226; http://dx.doi.org/0.3389/fpls.2015.01192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Berr A, Menard R, Heitz T, Shen WH. Chromatin modification and remodelling: a regulatory landscape for the control of Arabidopsis defence responses upon pathogen attack. Cell Microbiol 2012; 14:829-39; PMID:22405188; http://dx.doi.org/ 10.1111/j.1462-5822.2012.01785.x [DOI] [PubMed] [Google Scholar]
- 31.Germain H, Qu N, Cheng YT, Lee E, Huang Y, Dong OX, Gannon P, Huang S, Ding P, Li Y, et al.. MOS11: a new component in the mRNA export pathway. PLoS Genetics 2010; 6:e1001250; PMID:21203492; http://dx.doi.org/ 10.1371/journal.pgen.1001250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang Y, Li X. A putative nucleoporin 96 Is required for both basal defense and constitutive resistance responses mediated by suppressor of npr1-1,constitutive 1. Plant Cell 2005; 17:1306-16; PMID:15772285; http://dx.doi.org/ 10.1105/tpc.104.029926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Dong CH, Hu X, Tang W, Zheng X, Kim YS, Lee BH, Zhu JK. A putative Arabidopsis nucleoporin, AtNUP160, is critical for RNA export and required for plant tolerance to cold stress. Mol Cell Biol 2006; 26:9533-43; PMID:17030626; http://dx.doi.org/ 10.1128/MCB.01063-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


