Fig. 5.
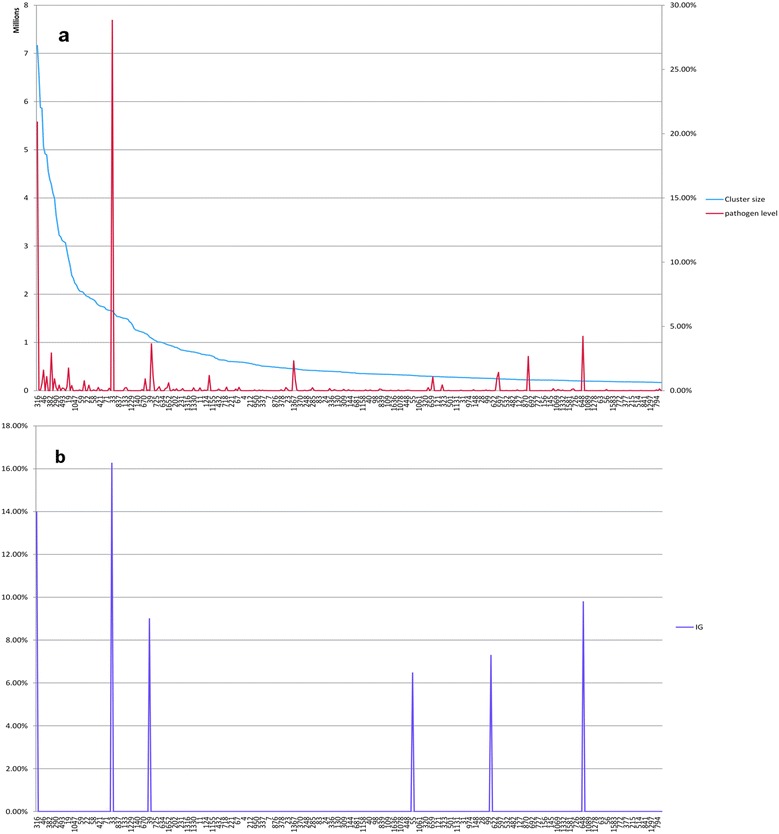
Evaluation of final third-level (3L) clusters. a Third-level clusters’ content in target (pathogen) genome sequences. 3L-clusters (labelled with their identifier on the x axis) are sorted according to their size (measured on the left axis, blue line). Red peaks show the fraction of total pathogen reads embedded in each 3L-cluster (measured on the right axis). b Relevance of final third-level clusters to disease status. 3L-clusters coordinates on the x axis are the same as in Fig. 5a; purple peaks represent information gain (IG) for each 3L-cluster with respect to sick versus healthy class assignments (see Methods). Note that only six 3L-clusters have IG values above background: the three highest red peaks (representing 3L-clusters embodying the bulk of the pathogen genome) correspond to the three highest IG purple peaks (first, second and last peaks); the remaining high IG peaks are correlated to the opposite (i.e., healthy) label, and are devoid of sequences from the pathogen strain
