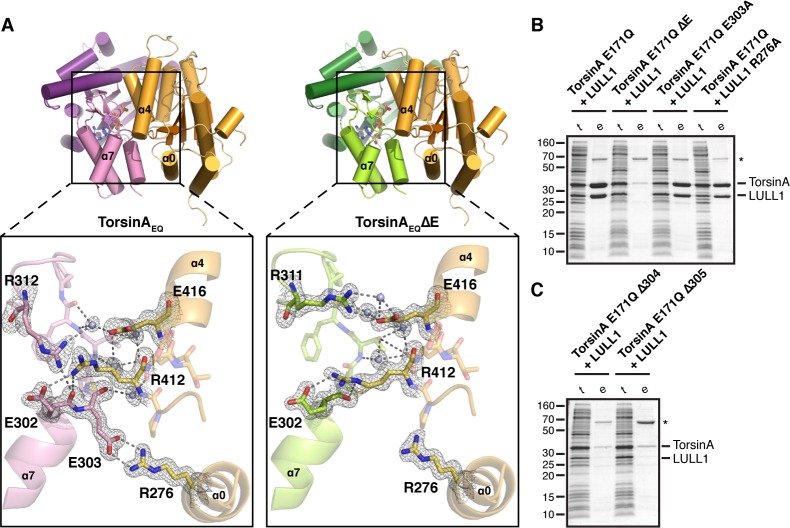
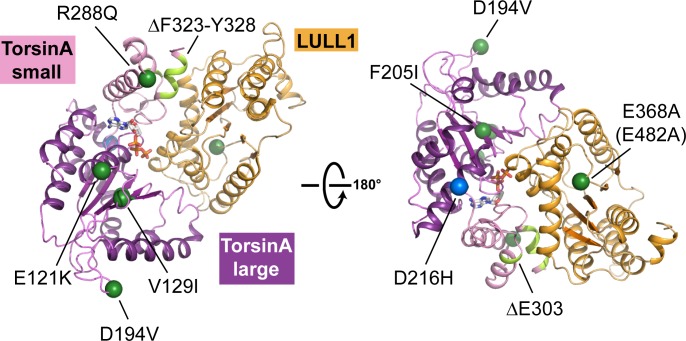
Figure 2. Analysis of the TorsinA-LULL1 interface.
(A) Side-by-side comparison of TorsinA-ATP-LULL1 (left) and TorsinAΔE-ATP-LULL1 (right). Zoomed insets show the atomic details of the interactions between TorsinA/TorsinAΔE and LULL1, with a focus on the ΔE303 area. (B and C) Mutational analysis of the TorsinA-LULL1 interface. Substitution or deletion of residues involved in TorsinA-LULL1 binding were probed using a Ni-affinity co-purification assay with recombinant, bacterial-expressed protein. Only TorsinA is His-tagged. SDS-PAGE analysis is shown. Lack of binding is observed by the absence of complex (uncomplexed His-tagged TorsinA is insoluble). t, total lysate, e, Ni eluate. Asterisk denotes an unrelated contaminant.