Figure 1.
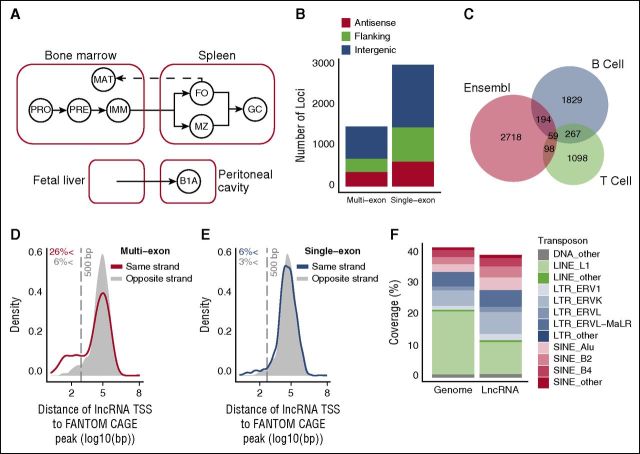
Identification of lncRNAs expressed in B cells. (A) Schematic representation of the ontogenetic relationships between B-cell populations used to generate the lncRNA catalog. Solid arrows indicate developmental progression through B-cell stages or activation (to GC B cells). Dashed line indicates recirculation of follicular B cells back to the bone marrow. B1A, B1a B cells; FO, follicular B cells; GC, germinal center B cells; IMM, immature B cells; MAT, mature B cells; MZ, marginal zone B cells; PRE, pre-B cells ; PRO, pro-B cells. (B) Genomic distribution of the 1491 multiexon and 3025 single-exon lncRNAs identified by this study. Positions are described relative to Ensembl v72 protein-coding gene annotations as antisense (overlapping a coding gene on antisense strand), flanking (<5 kb from coding gene), and intergenic (>5 kb from coding gene). (C) Overlap between the 2349 intergenic lncRNAs identified by this study (B cell), with those identified in T lineage cells18 (T cell), and those annotated in Ensembl v78. Kernel density plots representing the distribution of distance between each multiexon (D) and single-exon (E) intergenic lncRNA TSS and the nearest annotated TSS on the same strand that appeared in 2 or more of the 128 mouse cell lines considered by the FANTOM5 consortium. CAGE, cap analysis of gene expression. Shaded regions indicate a null distribution as measured by distance to the nearest FANTOM5 annotated TSS on the opposing strand. Vertical gray dashed line indicates a distance of 500 bp. (F) Coverage of the genome and of lncRNA exons by the indicated transposon elements.