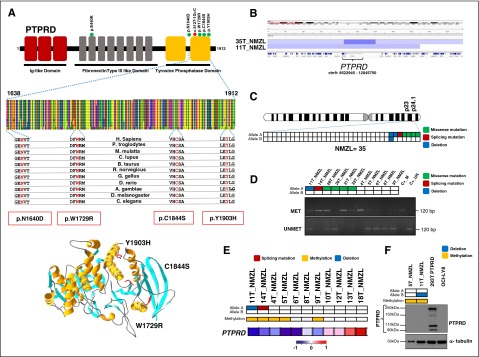
Figure 5.
PTPRD mutations in NMZL. (A) Schematic diagram of the PTPRD protein with its key functional domains (top). Color-coded symbols indicate the type and position of the mutations on the PTPRD protein (green, missense mutations; red, splicing site mutation). Multiple alignment of the PTPRD phosphatase domain amino acid sequences with the 11 orthologous PTPRD proteins. Conserved amino acids affected by mutations are color coded in red. Representation of the 3D structure of the PTPRD phosphatase domain mutations (bottom). The structural view of the PTPRD phosphatase domain was generated in DeepView-Swiss-PdbViewer (http://spdbv.vital-it.ch/) using the coordinates of the crystal structure of the PTPRS phosphatase domain (99% identity with PTPRD) (PDB 2fh7). Residues targeted by somatic mutations in NMZL are highlighted. (B) Graphic representation of segmentation data from 2 NMZL cases harboring PTPRD copy number losses visualized with Integrative Genomics Viewer (IGV) software. Each track represents one sample, where blue indicates region of a copy number loss. Individual genes in the region are aligned in the bottom panel. (C) Allelic (A or B) distribution of PTPRD genetic lesions in individual NMZL cases (green, missense mutation; red, splicing mutation; blue, deletion). (D) Methylation-specific PCR documenting aberrant methylation of the PTPRD-promoter CpG sites in NMZL cases (MET, methylated sample; UNMET, unmethylated sample; C+_M, positive control of methylated reaction; C+_UN, positive control of unmethylated reaction; bp, base pair). In the heatmap, rows correspond to the allelic status of PTPRD gene in NMZL (white, wild-type; green, mutated; blue, deleted). (E) Transcriptome analysis of PTPRD expression in primary NMZL. In the heatmaps, the first 2 rows correspond to the allelic status of the PTPRD gene in NMZL (white, wild-type; green, mutated; blue, deleted), the third row shows the methylation status of PTPRD (white, unmethylated; yellow, methylated), the last row shows the expression of PTPRD in NMZL (red, high expression; blue, low expression). (F) Western blot analysis of PTPRD protein expression in primary NMZL samples. The HEK 293T cells tranfected with a vector containing the wild-type PTPRD tagged with GFP at the C-terminal were used as positive control. The OCI-LY8 cells harboring a biallelic deletion of PTPRD were used as negative control. In the positive control, western blot analysis shows the GFP-tagged full-length PTPRD protein (250 kDa), as well as 2 N-terminal PTPRD protein products (150 kDa and 90k Da), and a GFP-tagged C-terminal PTPRD product (110 kDa) after cleavage process by a physiologic posttranslational modification called ectodomain shedding60 α-tubulin was used as a loading control. In the heatmaps, the first 2 rows correspond to the allelic status of PTPRD gene in NMZL (white, wild-type; blue, deleted), and the third row shows the methylation status of PTPRD (white, unmethylated; yellow, methylated).