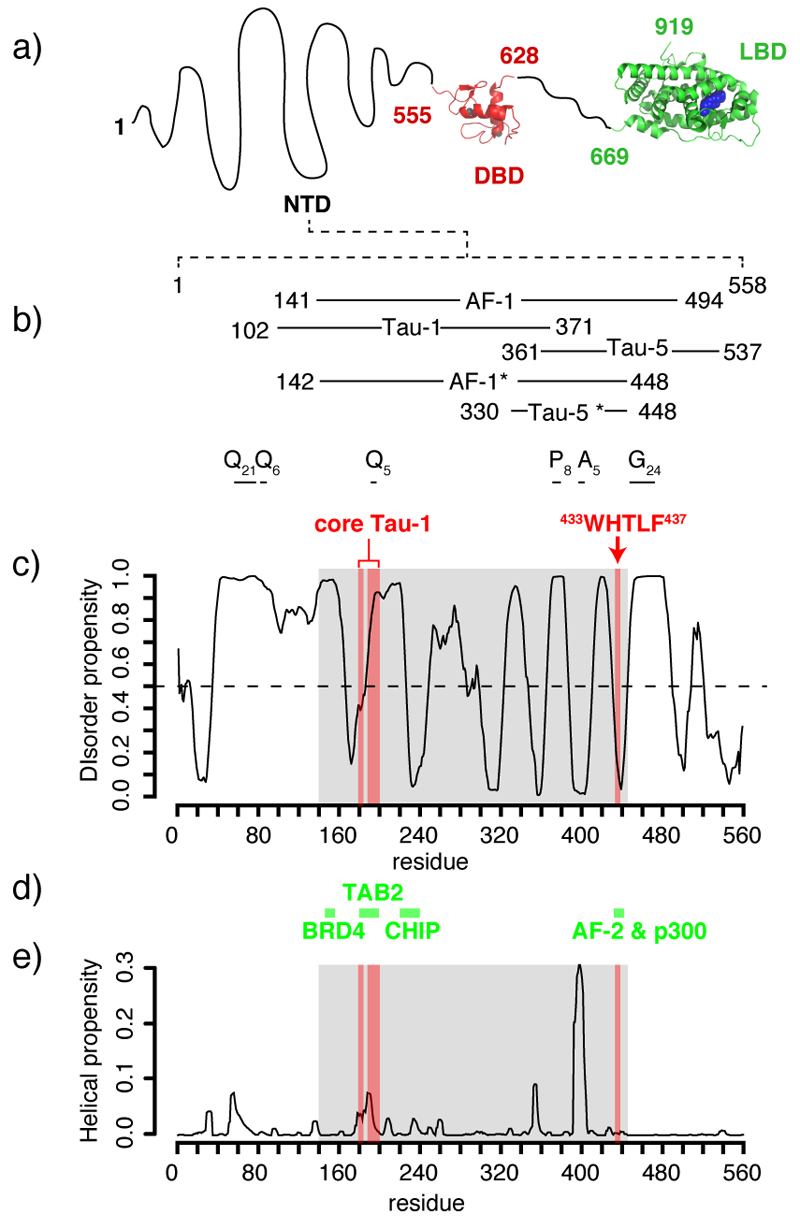
Figure 1.
Predicted properties of the sequence of the transactivation domain of AR a) Domain organization of AR 34,39 with an indication of the position of Zn atoms in the DBD (grey) and of dihydrotestosterone (DHT) in the LBD (blue). b) Definitions of activation function 1 (AF-1), transcription acctivation units 1 and 5 (Tau-1 and Tau-5), AF-1* and Tau-5* (the regions of sequence studied in this work) with an indication of the regions of low sequence complexity such as polyGln (Qn), polyPro (Pn), polyAla (An) and polyGly (Gn) tracts. c) Propensity to disorder of the NTD predicted by PONDR VL-XT26 with an indication of the functional motifs defining the core of Tau-1 and Tau-5, shaded in red and of the region of sequence studied in this work, shaded in grey. d) Positions of the motifs of the NTD of AR involved in protein protein interactions and acronyms of the binding partners (see main text for details). e) Propensity to adopt α-helical conformations predicted by Agadir 36, as a function of residue number, with an indication of the core of Tau-1 and Tau-5 (shaded in red) and AF-1* (shaded in grey).