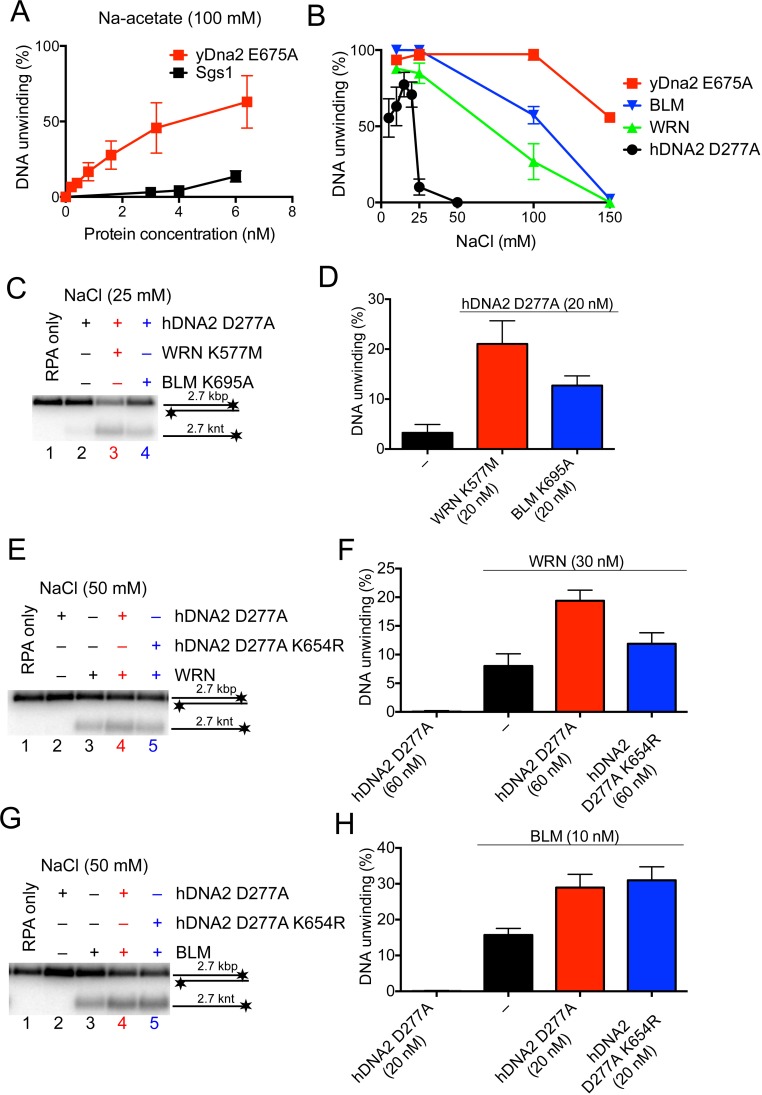
Figure 7. The helicase activity of hDNA2 functionally integrates with BLM or WRN helicases.
(A) Quantitation of 2.7 kbp-long dsDNA unwinding by yDna2 E675A or Sgs1 with 400 nM of yeast RPA. Reactions were supplemented with 100 mM sodium acetate and 5 mM magnesium acetate and incubated at 30°C. Averages shown, n = 2–3; error bars, SEM (B) Quantitation of DNA unwinding by yDna2 E675A (1 nM), BLM (10 nM), WRN (30 nM), hDNA2 D277A (30 nM) and its dependence on NaCl concentration. Reactions were supplemented with indicated NaCl concentrations and 2 mM magnesium acetate and incubated at 37°C. Averages shown, n = 2–3; error bars, SEM. (C) Representative 1% agarose gel showing DNA unwinding by hDNA2 D277A (20 nM) and its stimulation by helicase-deficient WRN K577M (20 nM) and BLM K695A (20 nM) variants. The reactions were supplemented with 25 mM NaCl and contained 215 nM hRPA. (D) Quantitation of experiments such as shown in Figure 7C. Averages shown, n = 5–7; error bars, SEM. (E) Representative 1% agarose gel showing the interplay of wild type WRN (30 nM) and nuclease-deficient hDNA2 D277A (60 nM) or nuclease- and helicase-deficient hDNA2 D277A K654R (60 nM) mutants. The reactions were supplemented with 50 mM NaCl and 215 nM hRPA. (F) Quantitation of experiments such as shown in Figure 7E. Averages shown, n = 3–4; error bars, SEM. (G) Representative 1% agarose gel showing the interplay of wild type BLM (10 nM) and nuclease-deficient hDNA2 D277A (20 nM) or nuclease- and helicase-deficient hDNA2 D277A K654R (20 nM) mutants. The reactions were supplemented with 50 mM NaCl and 215 nM hRPA. (H) Quantitation of experiments such as shown in Figure 7G. Averages shown, n = 2–4; error bars, SEM.