Fig. 2.
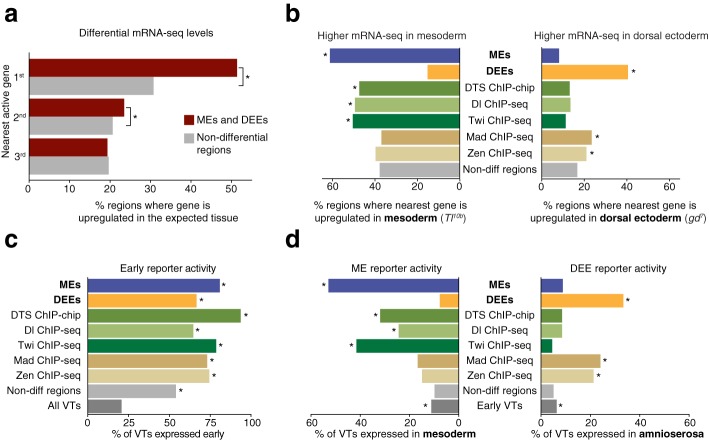
Genes near putative enhancers are differentially regulated across tissues. a The closest active genes near MEs and DEEs are significantly differentially expressed in the expected tissue (Tl 10b or gd 7) as compared to non-differential control regions (p < 10−30, one-sided chi-squared test). The second closest genes also show a small but significant enrichment (p < 0.034, one-sided chi-squared test), while the third nearest active genes are no longer enriched over the control. b Fraction of genes with higher mesodermal expression (left) is largest among MEs, and that with higher dorsal ectoderm expression (right) is largest among DEEs. As a comparison, the top 400 regions identified by transcription factor ChIP-seq data are shown: Dl and Twi ChIP-seq experiments from Tl 10b embryos, and Mad and Zen ChIP-seq experiments from gd 7 embryos. The star marks significance over non-differential regions (p < 0.01, one-sided chi-squared test). c Transgenic reporter activity of Vienna Tiles (VTs) that overlap MEs, DEEs, and transcription factor ChIP regions are preferentially expressed during early embryonic stages (with annotated expression in any tissue at stages 4–10). All groups show expression far above the average of all VTs (marked as star, p < 0.01, Fisher’s exact test). d VT reporter expression is more tissue-specific for MEs and DEEs compared to transcription factor ChIP regions. ME reporter activity (left) was defined as annotations by Kvon et al. [39] containing “mesoderm,” and DEE reporter activity (right) as annotations containing “amnioserosa.” The star marks significance over non-differential regions (p < 0.01, Fisher’s exact test). Number of regions: all VTs (7705), early expressed VTs (1595), VTs overlapping putative DV enhancers (148), MEs (68), DEEs (80)