Fig. 4.
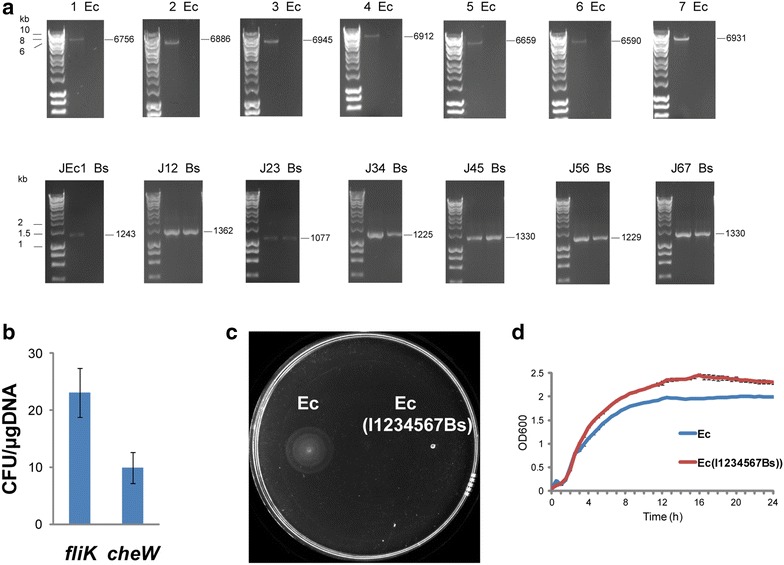
Chromosomal integration of 50 kb DNA into E. coli. a PCR verification of the integration of the seven DNA fragments (50 kb DNA length in total) into the E. coli K12 MG1655 chromosome. Ec E. coli K12 MG1655 wild type; Bs B. subtilis 168 wild type; 1,2,3,4,5,6,7: DNA fragments 1,2,3,4,5,6,7 in Ec(I1234567Bs); JEc1, J12, J23, J34, J45, J56, J67: junctions between the E. coli chromosome and DNA fragment 1, DNA fragments 1 and 2, DNA fragments 2 and 3, DNA fragments 3 and 4, DNA fragments 4 and 5, DNA fragments 5 and 6, DNA fragments 6 and 7 in Ec(I1234567Bs), respectively. Expected amplicon sizes are shown on the right. b Efficiency of integration of the 50 kb B. subtilis DNA into the fliK and cheW loci in the E. coli K12 MG1655 chromosome. Integration efficiencies were calculated from the number of colony forming units per µg of electroporated DNA. The figure shows means and standard deviations calculated from integration of seven DNA fragments. c Motility of the E. coli K12 MG1655 wild type (Ec) and strain Ec(I1234567Bs) harboring integration of the 50 kb B. subtilis DNA in the fliK locus of the flagellar region 3b. Overnight E. coli cultures (2 μl, OD600 of 1.0) were inoculated on the motility agar plates and analyzed after 5 h of growth at 37 °C. d Growth rate of the E. coli K12 MG1655 wild type (Ec) and strain Ec(I1234567Bs) harboring integration of the 50 kb B. subtilis DNA in the fliK locus of the flagellar region 3b measured with the microplate reader (Fluostar Omega). The means and standard errors were calculated from three independent experiments
