Fig. 6.
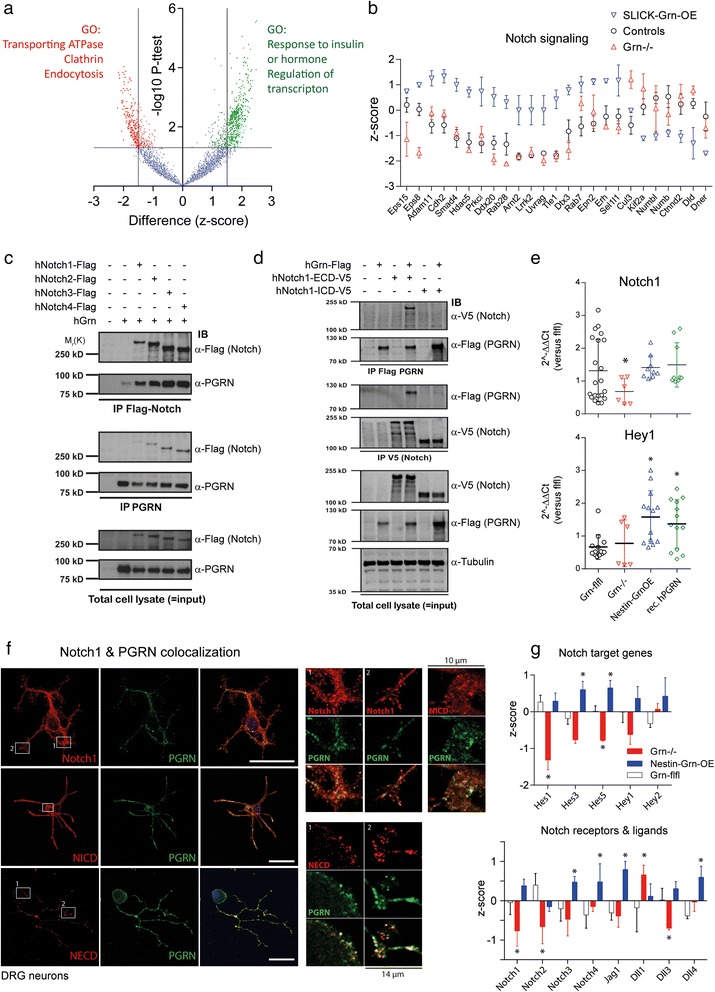
Proteome analysis and assessment of progranulin—Notch interactions. a Volcano plot showing differentially expressed proteins in the prefrontal cortex of SLICK-Grn-OE (n = 3) versus control mice (n = 6). LN-transformed LFQ (label free quantification) values were z-transformed and then submitted to 2-tailed independent t-tests, using FDR adjustments. The –log P values were plotted on the Y-axis versus the z-score difference on the x-axis. Upregulated proteins are shown in the upper right corner, downregulated proteins in the upper left corner. Significantly up- or downregulated proteins (P < 0.05; = − logP > 1.3) are highlighted in green or red color, respectively and were subsequently submitted to Gene Ontology (GO) enrichment analysis (genetrail); full results in Additional file 2: Table S1. b Scatter plots (mean and SD of the z-scores) of significantly regulated proteins, which are involved in Notch signaling or traffic. The list of regulated proteins was searched for proteins involved in Notch signaling or traffic by comparing them with Notch networks (biogrid, genemania) and with proteins that have GO annotations of “Notch signaling” (n = 3 per group, n = 6 for controls). c Co-immunoprecipitation analysis of interactions between Notch receptors and progranulin. HEK293 cells were transiently co-transfected with Flag-tagged human Notch receptors and full-length non-tagged human progranulin (hGrn). Pull-down was performed with anti-Flag (for Notch receptors) or anti-progranulin (N19 antibody) followed by immunoblot analysis with anti-Flag and anti-progranulin. Input blots with total protein lysates were done with 1/10th of the protein subjected to IP. Input blots of total protein lysates are shown in the bottom. d Co-immunoprecipitation analysis of interactions between the extracellular or intracellular domains (NECD and NICD) of human Notch1 with human progranulin. HEK293 cells were transiently co-transfected with hNotch1-ECD-V5 or hNotch1-ICD-V5 and Flag-tagged full-length human progranulin (hGrn-Flag, all tags were C-terminal). IPs were performed with anti-Flag and anti-V5 followed by immunoblot analysis with anti-V5 (for the NECD, NICD) and anti-Flag (for progranulin, PGRN). Input blots were done with 1/10th of the protein and beta tubulin served as loading control for input blots. e Scatter plots of Notch1 and Hey1 mRNA (QRT-PCR) in primary embryonic neuron cultures of Nestin-Grn-OE, Grn−/− and control mice, the latter without/with stimulation with recombinant human progranulin (rPGRN). Each symbol represents a culture, the line is the mean, the whiskers show the SD. Rps13 was used as housekeeping gene for calculation of ΔCt values. Asterisks show significant differences versus unstimulated control cultures. * P < 0.05, one-way ANOVA, posthoc t-tests according to Dunnett versus control. f Immunofluorescence wide-field laser scanning microscopic analysis of the co-expression of progranulin with Notch1, the NICD and the NECD of Notch1 in primary neurons of the dorsal root ganglia of adult STOP-Grnflfl mice. Scale bar 25 μm. The right panels show zoom-in images of the white rectangular regions (width 10 μm for Notch1 and NICD and 14 μm for NECD). Quantitative colocalization analysis revealed partial overlap (Additional file 2: Table S4). g Bar charts showing Z-cores (means + SD) of QRT-PCR mRNA levels of Notch target genes, Notch receptors and Notch ligands after nerve injury in DRGs. Results of ipsilateral and contralateral DRGs were summarized. Data are of n = 3 samples per genotype. Each sample consisted of the ipsilateral or contralateral L4 and L5 DRGs of each 3 mice. Hence, the analysis is based on tissue of 18 mice with 36 ipsilateral and 36 contralateral DRGs. Each sample was analyzed in 3 replicate RT-PCR runs for each gene. Two housekeeping genes, RNA Pol-III and Rps13, were used for calculation of ΔCt values. The ΔCt data of the respective genes were Z-transformed for each gene separately. Z-transformed data were submitted to 2-way ANOVA for “gene” by “genotype” and groups differed significantly with P = 0.0005. Subsequently, one-way ANOVAs were performed for each gene individually with subsequent post-hoc analyses according to Dunnett versus STOP-Grnflfl control mice. Asterisks show statistically significant results, * P < 0.05
