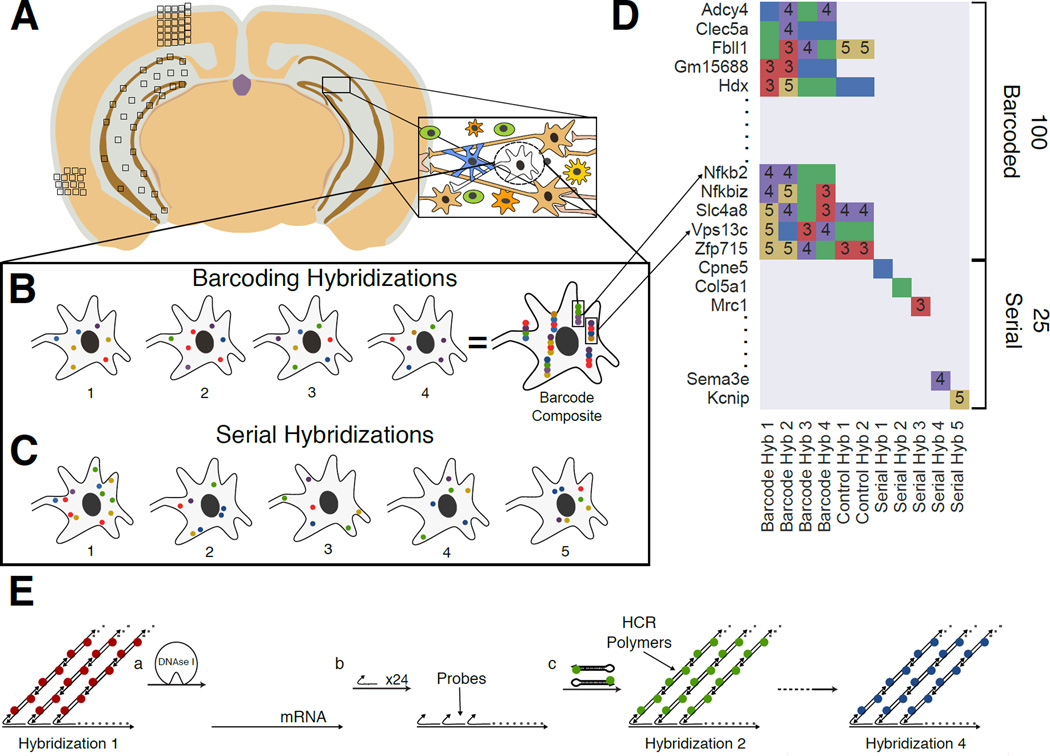
Fig. 1. Overview of the Sequential barcode FISH (seqFISH) in brain slices.
A. A coronal section from a mouse brain was mounted on a slide and imaged in all boxed areas. Each image was taken at 60x magnification. B. Example of barcoding hybridizations from one cell in field from A. The same points are re-probed through a sequence of 4 hybridizations (numbered). The sequence of colors at a given location provides a barcode readout for that mRNA (“barcode composite”). These barcodes are identified through referencing a lookup table abbreviated in D and quantified to obtain single cell expression. In principle, the maximum number of transcripts that can be identified with this approach scales to FN, where F is the number of fluorophores and N is the number of hybridizations. Error correction adds another round of hybridization. C. Serial smHCR is an alternative detection method where 5 genes are quantified in each hybridization and repeated N times. Serial hybridization scales as F*N. D. Schematic for multiplexing 125 genes in single cells. 100 genes are multiplexed in 4 hybridizations by seqFISH barcoding. This barcode scheme is tolerant to loss of any round of hybridization in the experiment. 25 genes are serially hybridized 5 genes at a time by 5 rounds of hybridization. Each number represents a color channel in single molecule HCR. As a control, 5 genes are measured both by double rounds of smHCR as well as barcoding in the same cell. E. SmHCR amplifies signal from individual mRNAs. After imaging, DNAse strips the smHCR probes from the mRNA, enabling rehybridization on the same mRNA (step a). The “color” of an mRNA can be modulated by hybridizing probes that trigger HCR polymers labeled with different dyes (step b). mRNA are amplified following hybridization by adding the complementary hairpin pair (step c). The DNAse smHCR cycle is repeated on the same mRNAs to construct a predefined barcode over time.