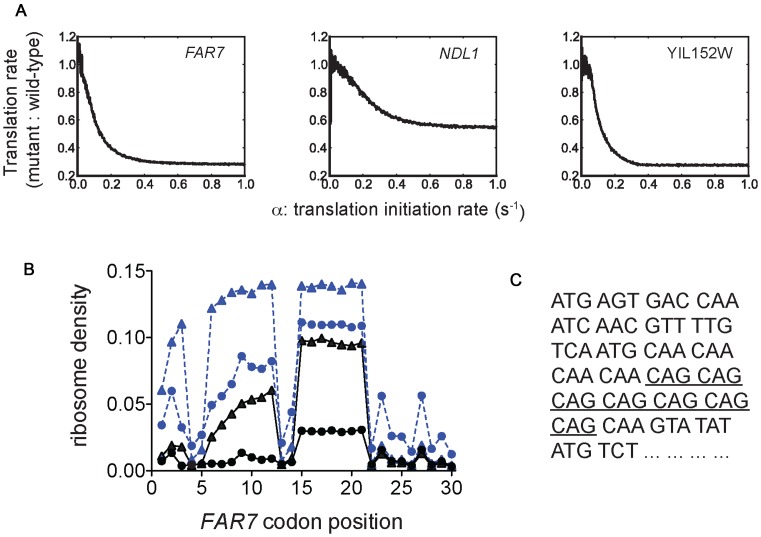
Figure 5.
The rate of translation elongation at CAG codons, relative to the rate of translation initiation, governs ribosome queue formation and thus 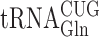 sensitivity. (A) TASEP simulation of translation of three
sensitivity. (A) TASEP simulation of translation of three 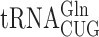 -sensitive mRNAs was conducted across a range of values of α, the translation initiation rate (Figure 2). Simulations were conducted in either a wild-type tRNA background, or a sup70-65 tRNA background, and a ratio of these translational efficiencies plotted against each value of the translation initiation rate. (B) These same TASEP simulations were used to record the codon-specific ribosomal density across the FAR7 ORF to indicate the positions of ribosome queuing. The ribosomal density across the FAR7 was recorded in a wild-type strain (filled circle symbols) and the sup70-65 mutant condition (open triangle symbols), at the physiological initiation rate of 0.3 events/s (dashed lines, blue symbols), and again at a 6-fold slower rate of 0.05 events/s (solid lines). The ribosomal density across codons 1–30 is presented, showing that the ribosomal density at the mRNA 5′ end in the sup70-65 mutant is greater than that in the wild-type at the high initiation rate, but that ribosome queues dissipate at the lower initiation rate, eliminating this density differential at the 5′-most codons. (C) Codons 1–24 of the FAR7 open reading frame, with the positions of the CAG codons indicated (underlined).
-sensitive mRNAs was conducted across a range of values of α, the translation initiation rate (Figure 2). Simulations were conducted in either a wild-type tRNA background, or a sup70-65 tRNA background, and a ratio of these translational efficiencies plotted against each value of the translation initiation rate. (B) These same TASEP simulations were used to record the codon-specific ribosomal density across the FAR7 ORF to indicate the positions of ribosome queuing. The ribosomal density across the FAR7 was recorded in a wild-type strain (filled circle symbols) and the sup70-65 mutant condition (open triangle symbols), at the physiological initiation rate of 0.3 events/s (dashed lines, blue symbols), and again at a 6-fold slower rate of 0.05 events/s (solid lines). The ribosomal density across codons 1–30 is presented, showing that the ribosomal density at the mRNA 5′ end in the sup70-65 mutant is greater than that in the wild-type at the high initiation rate, but that ribosome queues dissipate at the lower initiation rate, eliminating this density differential at the 5′-most codons. (C) Codons 1–24 of the FAR7 open reading frame, with the positions of the CAG codons indicated (underlined).