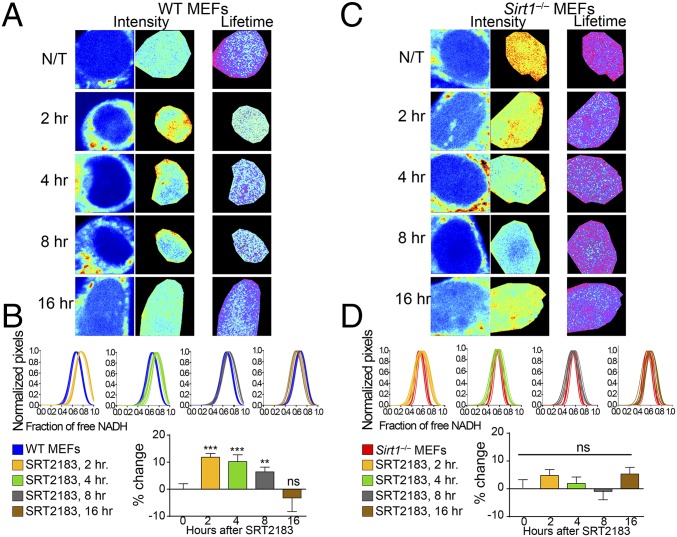
Fig. 2.
Metabolic transitions in the cell nucleus depend on SIRT1 activity. (A) Intensity and FLIM (lifetime) images from untreated (N/T) WT MEFs or those treated with SRT2183 (10 μM) for 2, 4, 8, and 16 h as indicated. (B) Histograms in Upper show comparative analyses by overlapping the fractional free NADH distributions from nontreated single WT MEFs (blue lines) and single WT MEFs treated with SRT2183 (10 μM). The fraction of free NADH is represented in the x axis, where 1 = 100% free NADH; n = 10–15 single cells per experiment, and each line represents the distribution of the fraction of free NADH for each single cell. Graphs in Lower depict the percentage change on the fraction of free NADH. Data were normalized for nontreated WT MEFs = 0. Means ± SEM of three independent experiments are presented. **P < 0.01 by two-tailed MWU; ***P < 0.001 by two-tailed MWU. (C) Intensity and FLIM (lifetime) images from N/T Sirt1−/− MEFs or those treated with SRT2183 (10 μM) for 2, 4, 8, and 16 h as indicated. (D) Histograms in Upper show comparative analyses by overlapping the fractional free NADH distributions from nontreated single Sirt1−/− MEFs (red lines) and single Sirt1−/− MEFs treated with SRT2183 (10 μM). The fraction of free NADH is represented in the x axis, where 1 = 100% free NADH; n = 10–15 single cells per experiment, and each line represents the distribution of the fraction of free NADH for each single cell. Graphs in Lower depict the percentage change on the fraction of free NADH. Data were normalized for nontreated Sirt1−/− MEFs = 0. Means ± SEM of three independent experiments are presented. ns, Not significant by two-tailed MWU.