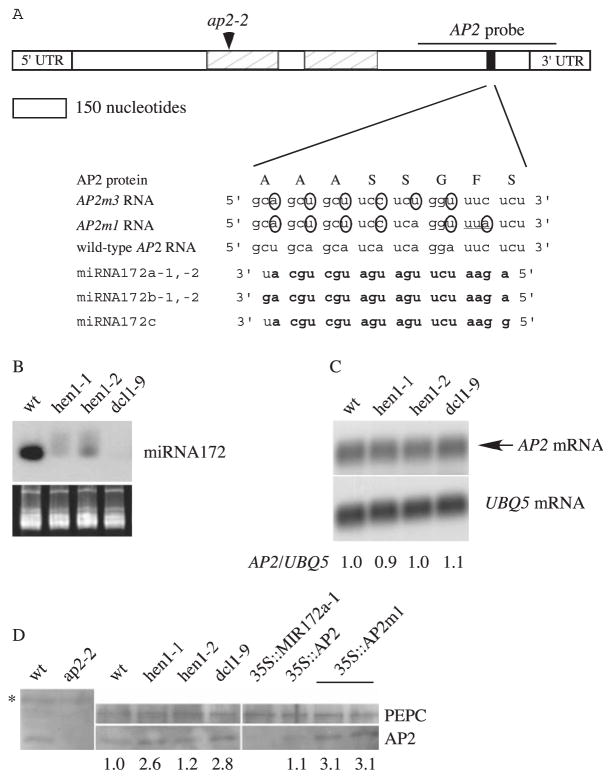
Fig. 1.
Sequence complementarity between AP2 RNA and miRNA172 and AP2 RNA and protein accumulation in various genotypes. (A) A diagram of the AP2 mRNA showing the AP2 domains (hatched rectangles) and the miRNA172 binding site (black rectangle). The mutant nucleotides in AP2m1 and AP2m3 are circled. Although AP2m1 causes a Phe-to-Leu amino acid substitution (underlined codon), AP2m3 does not change the amino acid sequence. Nucleotides in miRNA172 that can base-pair with AP2 RNA (with G:U pairing allowed) are in bold. The ap2-2 allele is a splice acceptor site mutation that would generate a stop codon at the indicated position (arrowhead). 3′UTR, 3′ untranslated region; 5′UTR, 5′ untranslated region. (B) Accumulation of miRNA172 in inflorescences. The stained gel below indicates the amount of RNA in each lane. wt, wild type. (C) AP2 RNA abundance in inflorescences. The numbers indicate the relative abundance of AP2 RNA in various genotypes. (D) AP2 protein abundance in inflorescences. Although the AP2 antisera reacted with a number of proteins (one indicated by the asterisk), the identity of the AP2 signal was revealed by its absence in the ap2-2 mutant. The numbers indicate the abundance of AP2, as calculated by using phosphoenolpyruvate carboxylase (PEPC) as the loading control.