Fig. 1.
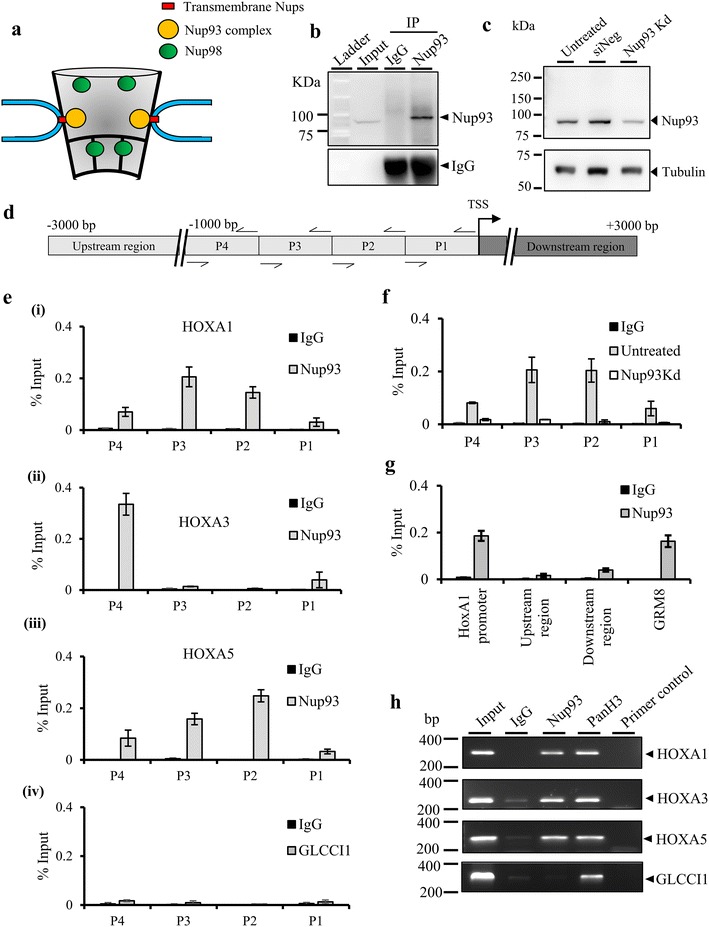
Nup93 associates with HOXA1, HOXA3 and HOXA5 promoters. a Pictorial representation of the nuclear pore complex (NPC) showing the relative location of the Nup93 sub-complex, and Nup98 within the nuclear pore complex [103]. b Immunoprecipitation of Nup93 using anti-Nup93 antibody on whole-cell extracts of DLD1 cells (a representative full blot from three independent biological replicates, N = 3). Anti-rabbit heavy chain IgG shows equal precipitation efficiency in IgG and Nup93 lanes. c A representative full Western blot showing siRNA-mediated depletion of Nup93 in DLD1 cells (lane Nup93 Kd), Untreated Untreated cells, siNeg ON-TARGETplus non-targeting siRNA control (a representative blot from three independent biological replicates, N = 3). d Pictorial representation of primer pair positions (P1–P4) on the promoter of HOXA1, HOXA3 and HOXA5 genes, respectively, upstream region and downstream region indicates primer position outside the HOXA1 promoter, arrow indicates transcription start site (TSS). e ChIP experiments were performed using antibodies specific to Nup93 and IgG control. Nup93 ChIP-qPCR on (i) HOXA1 (ii) HOXA3 (iii) HOXA5 and (iv) GLCCI1 promoters, respectively. Y-axis: immunoprecipitated DNA relative to 1% input, corrected for ChIP using non-specific IgG (N = 2, data from two independent biological replicates that include a total of six technical replicates), error bar standard error of mean (SEM). f Nup93 ChIP-qPCR was performed in untreated and Nup93 knockdown cells for HOXA1 promoter using primer pairs P1–P4. g Nup93 ChIP-qPCR using primer pairs outside HOXA1 promoter regions (upstream region and downstream region) and primers for a Nup93-associated gene (GRM8), used as a positive control. h ChIP-PCR amplification of HOXA1, HOXA3 and HOXA5. GLCCI1 used as negative control. Nup93 binds to ~300–600 bp on each of these HOXA promoters, (N = 2, representative data from two independent biological replicates)
