Table 2. Parameter estimates and likelihood scores for the PR gene under different branch-site models.
| Codon model | Hypothesis | Estimates of ω parameters | Positive selection | 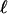 |
|---|---|---|---|---|
| M3 k = 2 | H0: BG = G-FG = B-FG | p0 = 0.81, ω0 = 0.02 | None | -6176.81 |
| (p1 = 0.18), ω1 = 0.33 | ||||
| Model B, ω2 = 1 | H1: BG = B-FG ≠ G-FG | p0 = 0.50, ω0 = 0.02 | G-FG: not allowed | -6167.16 |
| p1 = 0.12, ω1 = 0.31 | ||||
| (p2+3 = 0.38), (ω2 = 1) | BG+B-FG: none | |||
| H2: BG = G-FG ≠ B-FG | p0 = 0.78, ω0 = 0.03 | B-FG: not allowed | -6164.15 | |
| p1 = 0.13, ω1 = 0.38 | ||||
| (p2+3 = 0.08), (ω2 = 1) | BG+G-FG: none | |||
| H3: BG ≠ G-FG = B-FG | p0 = 0.75, ω0 = 0.03 | FG1 + FG2: not allowed | -6153.40 | |
| p1 = 0.13, ω1 = 0.36 | ||||
| (p2+3 = 0.12), (ω2 = 1) | BG: none | |||
| Model B | H1: BG = B-FG ≠ G-FG | p0 = 0.78, ω0 = 0.02 | G-FG: 5 sites | -6159.24 |
| p1 = 0.18, ω1 = 0.31 | ||||
| (p2+3 = 0.03), ω2 = 76 | BG+B-FG: none | |||
| H2: BG = G-FG ≠ B-FG | p0 = 0.82, ω0 = 0.03 | B-FG: 10 sites | -6156.29 | |
| p1 = 0.13, ω1 = 0.39 | ||||
| (p2+3 = 0.05), ω2 = 99 | BG+G-FG: none | |||
| H3: BG ≠ G-FG = B-FG | p0 = 0.80, ω0 = 0.03 | FG1+FG2: 16 sites | -6142.67 | |
| p1 = 0.13, ω1 = 0.37 | ||||
| (p2+3 = 0.06), ω2 = 14 | BG: none |
Tree topology, BG, G-FG, and B-FG are presented in Fig 1. 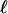 , log likelihood score. Boldface indicates an estimate value >1. Parentheses indicate a parameter value obtained by subtraction.
, log likelihood score. Boldface indicates an estimate value >1. Parentheses indicate a parameter value obtained by subtraction.
