Fig. 1.
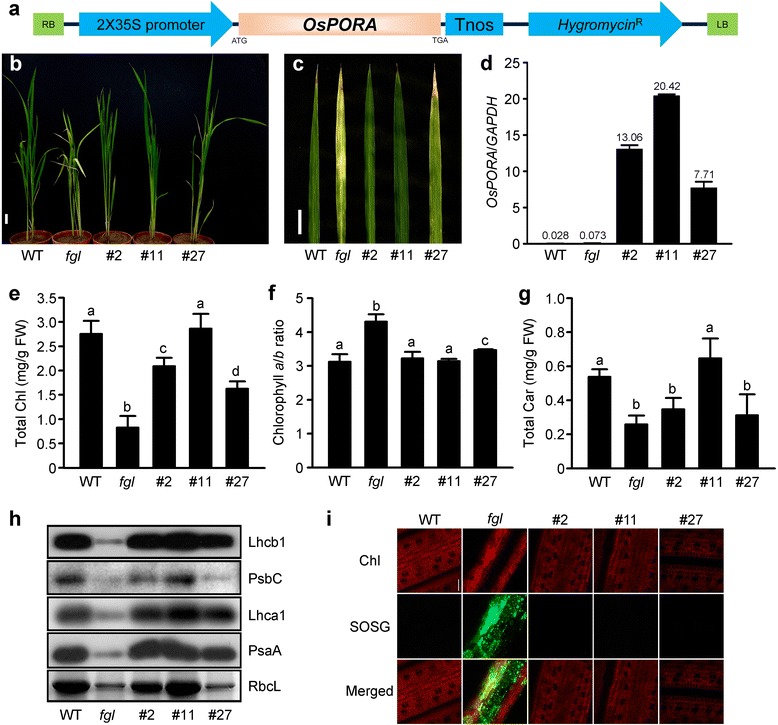
Characterization of 35S:OsPORA/fgl (OPAO) plants grown under natural sunlight in the greenhouse. a Structure of the construct used to generate the OPAO transgenic plants. b-c Phenotypes of WT, fgl, and three independent OPAO T0 plants. Plants (b) and leaf tip regions of the fourth leaf blades of plants (c) grown in the greenhouse. Scale bars = 5 cm (b) and 2 cm (c). d Transcript levels of OsPORA in WT, fgl, and OPAO plant lines #2, #11, and #27. Means and standard deviations were obtained from three independent leaf parts. The numbers on the bars represent relative values from qRT-PCR. e-g Measurement of photosynthetic pigments in WT, fgl, and three OPAO lines. Total chlorophyll (Chl) contents (e), Chl a/b ratio (f), total carotenoid (Car) contents (g). The data were obtained from five independent leaf regions (e-g). h Immunoblot analysis of photosystem proteins. Lhcb1, PsbC, Lhca1, and PsaA were detected using specific antibodies, and Rubisco large subunit (RbcL) was detected by Coomassie Blue staining. Similar results were obtained in three independent experiments (h). i Detection of singlet oxygen (1O2) accumulation. Red indicates Chl autofluorescence (upper panels), green indicates SOSG fluorescence (middle panels), and merged Chl and SOSG fluorescent signals are shown (lower panels). Similar results were obtained from at least five independent leaf disks. Scale bar = 50 μm (i). Leaves were sampled from 2-month-old plants grown in the greenhouse. Letters a, b, and c on the bars indicate significantly different values at the 1% level according to Duncan’s multiple range test (e-g)
