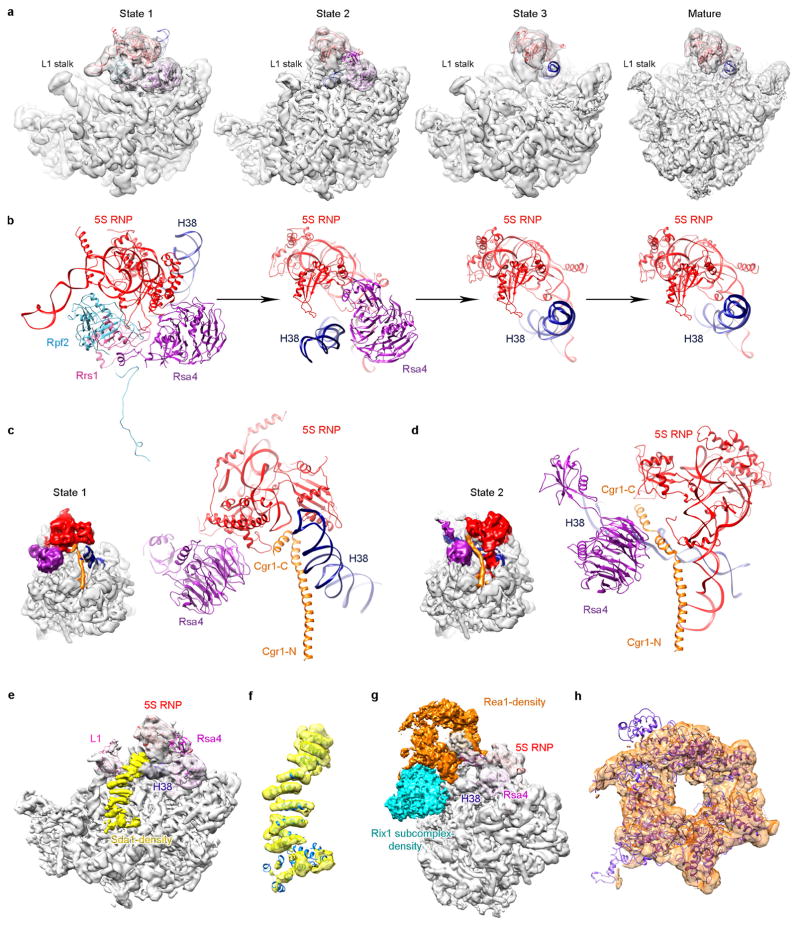
Extended Data Figure 9. Structures of different assembly states of the pre-60S ribosomal particles.
a, Cryo-EM density maps of three premature states (1–3) and the mature state are displayed in transparent surface representation, superimposed with models of the 5S RNA, H38 and associated central-protuberance-binding factors. b, Zoom-in views of the central protuberance regions in a. For clarification, only atomic models are shown. Comparison of these four states indicates that the 5S RNP rotates to a near-mature state (state 2) after Rpf2–Rrs1 leave, and further release of Rsa4 in state 3 results in a ‘mature-like’ conformation for the 5S RNP. H38 from these four states is in a series of continuous changes coupled with the 5S RNP conformational maturation. c, d, Spatial relationship of the 5S RNP, H38, Rsa4 and Cgr1 in state 1 (c) and state 2 (d). Note that repositioning of H38 from state 1 to state 2 is coupled with a dramatic conformational change on the C-terminal end of Cgr1. e–h, Additional assembly factors identified in the density map of state 2. One piece of additional density between H38 and L1 contains a characteristic HEAT repeat, which contacts the L1 stalk in an inward position (e). The atomic model of Sda1 (PDB accession number 5FL8)14 fits well with the segmented density (f). For clarification, densities immediately above Sda1 are not shown in e and f. A large piece of additional density in the map of state 2, composed of the Rix1 subcomplex and Rea1 (g, h). The density assignment was facilitated by the cryo-EM structure of Rix1–Rea1 particles14. Superimposition of the atomic model of Rea1 (PDB accession number 5FL8)14 with the segmented density map of Rea1 (h).