Abstract
Purpose
New therapeutic approaches are needed for patients with thyroid cancer refractory to radioiodine treatment. An inhibitor of bromodomain and extraterminal domain (BET) proteins, JQ1, shows potent anti-tumor effects in hematological cancers and solid tumors. To evaluate whether JQ1 is effective against thyroid cancer, we examined anti-tumor efficacy of JQ1 using the ThrbPV/PVKrasG12D mouse, a model of anaplastic thyroid cancer.
Experimental Design
We treated ThrbPV/PVKrasG12D mice with vehicle or JQ1 at a dose of 50 mg/kg body weight/day starting at the age of 8 weeks for a 10-week period and monitored thyroid tumor progression.
Results
JQ1 markedly inhibited thyroid tumor growth and prolonged survival of these mice. Global differential gene expression analysis showed that JQ1 suppressed the cMyc (hereafter referred to as Myc) transcription program by inhibiting mRNA expression of Myc, ccnd1, and other related genes. JQ1-suppressed Myc expression was accompanied by chromatin remodeling as evidenced by increased expression of histones and hexamethylene bis-acetamide inducible 1, a suppressor of RNA polymerase II transcription elongation. Analyses showed that JQ1 decreased MYC abundance in thyroid tumors and attenuated the cyclin-CDK4-Rb-E2F3 signaling to decrease tumor growth. Further analysis indicated that JQ1 inhibited the recruitment of BDR4 to the promoter complex of the Myc and Ccnd1 genes in rat thyroid follicular PCCL3 cells, resulting in decreased MYC expression at the mRNA and protein levels to inhibit tumor cell proliferation.
Conclusions
These preclinical findings suggest that BET inhibitors may be an effective agent to reduce thyroid tumor burden for the treatment of refractory thyroid cancer.
Keywords: thyroid hormone receptors, tumorigenesis, genetically engineered mouse model, thyroid cancer, bromodomain and extraterminal protein inhibitor, JQ1
Introduction
Thyroid cancer is the most common malignancy of the endocrine organs (1). While well-differentiated thyroid cancer responds well to radioiodine therapy and usually has a favorable therapeutic outcome, the treatment of refractory thyroid carcinoma remains a major challenge. Intensive efforts have been made in the search for effective ways to treat refractory thyroid cancer (2).
Chromatin remodeling through histone acetylation is a key step in the regulation of the gene expression in both normal and tumor cells (3). Members of the bromodomain and extraterminal domain (BET) family of proteins, such as bromodomain-containing protein 4 (BRD4), interact with acetylated histones to assemble chromatin complexes and transcription activators at specific promoter sites (4). These BET proteins are potential therapeutic targets to treat cancers. Selective inhibitors of BET proteins, such as JQ1, inhibit the interaction of BET proteins with acetylated histones at promoter sites, resulting in an alteration of downstream signaling events (5, 6). The efficacy of BET inhibitors has been linked to their ability to suppress v-myc myelocytomatosis viral oncogene homolog (MYC) expression and downstream transcriptional targets (6, 7). Inhibitors of BET proteins have shown potent inhibitory activity against hematological malignant cancers and other types such as glioblastoma and lung and prostate cancers (8-11). Although it has been shown that JQ1 inhibits growth of established differentiated thyroid cancer cells in nude mice (12), the therapeutic efficacy of BET inhibitors in undifferentiated thyroid cancers has yet to be studied.
Genetic aberrations contribute to the development of thyroid cancer. RAS mutations, which are frequently detected in thyroid cancer, have high diagnostic negative predictive values for the disease (13, 14). We previously established a mouse model of follicular thyroid cancer, the ThrbPV/PV mouse (15, 16). After the mutant KrasG12D gene was targeted to the follicular thyroid cancer cells of ThrbPV/PV mice (ThrbPV/PVKrasG12D mice), the double mutant mice spontaneously developed metastatic undifferentiated follicular thyroid carcinoma resembling human anaplastic thyroid cancer with markedly shortened life expectancy (17). In the ThrbPV/PVKrasG12D mice, MYC was identified as a critical factor to promote the development of undifferentiated metastatic thyroid cancer (17). In the Kras-mutant non-small cell lung cancer mouse model, JQ1 treatment produces significant tumor regression via coordinate downregulation of the MYC-dependent program (18).
In this study, we investigated the therapeutic efficacy of JQ1 in the treatment of thyroid cancer in ThrbPV/PVKrasG12D mice and found that JQ1 inhibited growth and proliferation of thyroid tumors in them. JQ1 treatment suppressed the MYC functions and signaling that promote thyroid tumor growth via interference with BRD4 functions. Our findings suggest that BET inhibitors may be effective agents for the treatment of anaplastic thyroid cancer.
Materials and Methods
Animals and treatment of JQ1
The National Cancer Institute Animal Care and Use Committee approved the protocols for animal care and handling in the present study. Mice harboring the ThrbPV gene (ThrbPV/PV mice) and ThrbPV/PVKrasG12D mice were previously described (17, 18). JQ1was dissolved in DMSO solution to make a 100 mg/ml stock and administered by oral gavage daily at a dose of 50 mg/kg body weight/day starting at the age of 8 weeks for a 10-week period. The thyroids and lungs were dissected after mice were euthanized for weighing, histologic analysis, and biochemical studies.
Western blot analysis
The Western blot analysis was carried out as described by Zhu et al (17). Primary antibodies for CDK4 (#2906), p-Rb (#9307), and GAPDH (#2118) were purchased from Cell Signaling Technology (Danvers, MA). The E2F3 primary antibody (sc-878) and Rb (sc-50) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA). Primary antibody against Ki-67 (RB-9043-P0) was purchased from Neomarkers (Fremont, CA). The hexamethylene bis-acetamide inducible 1 (HEXIM1) primary antibody (A303-113A), and BRD4 (A301-985A50) were purchased from Bethyl Laboratories Inc (Montgomery, TX). Antibodies were used at the manufacturers’ recommended concentration. For control of protein loading, the blot was probed with the antibody against GAPDH.
Histological analysis and immunohistochemistry
Thyroid glands, heart, and lung were dissected and embedded in paraffin. Five-micrometer-thick sections were prepared and stained with hematoxylin and eosin (H&E). For each mouse, single random sections through the thyroid, lung, and heart were examined. Immunohistochemistry was performed with paraffin sections by standard methods.
Microarray analysis
Microarray analysis was carried out as described by Zhu et al (19). Briefly, biotinylated-aRNA samples from three individual mice of each group were used in hybridization of the GeneChip Mouse Exon 1.0 ST Array (affymetrix, Santa Clara, CA) and scanned on an Affymetrix GeneChip scanner 3000. Data were collected using Affymetrix GCOS software. Data processing and analysis were done by affy, limma, xps R/Bioconductor packages (http://www.bioconductor.org). Briefly, the robust multichip average (RMA) method was used for computing expression measures, and the Benjamini and Hochberg method was used for calculating the adjusted p values. Differentially expressed genes were selected by the adjusted p values with a minimum 2.0-fold change. The GEO array data submission is in progress.
RNA extraction and real time RT-PCR validation of microarray data
Total RNA from thyroids was isolated using TRIzol (Invitrogen, Carlsbad, CA) as indicated by the manufacturer’s protocol. Selected genes from microarray data were chosen for real time RT-PCR validation. A total 200 ng of RNA extracted from thyroids of ThrbPV/PVKrasG12D mice with vehicle or JQ1 treatment was used in the real-time RT-PCR. The reactions were performed with the QuantiTect SYBR RT-PCR kit (Qiagen, Germantown, MD) on an ABI 7900HT system. In each group, four to seven samples with triplicates were tested on the target genes. Data were analyzed using Prism V5 (GraphPad Software, Inc., La Jolla, CA). Primers were as follows.
For the mouse endogenous control glyceraldehyde-3-phosphate dehydrogenase (Gapdh) gene: forward, 5’-cgtcccgtagacaaaatggt-3’; reverse, 5’-gaatttgccgtgagtggagt-3’. For the mouse cyclin-dependent kinase 4 (Cdk4) gene: forward, CGTTGGCTGTATCTTTGCAG; reverse, GGAAGGCAGAGATTCGCTTA. For the moue retinoblastoma 1 (Rb) gene: forward, TACTGGCCTGTGCTCTTGAA; reverse, TCTCCATCCTTGGACTGCTT. For the mouse E2F transcription factor 3 (E2f3) gene: forward, TACGGAGTCCCGATAGTCCA; reverse, CTTCTTTGGACAGGCCTTGA. For the mouse cyclin D1 (Ccnd1) gene: forward, GAGCCCAACCGAGACCAC; reverse, CTCTTCGCACTTCTGCTCCT. For the mouse hexamethylene bis-acetamide inducible 1 (Hexim1) gene: forward, GTGCTGCTGTTGTCCATGAA; reverse, TTCTGGACAGGCATTCGTCT. For the mouse lysine acetyltransferase 2A (Kat2a) gene: forward, ACGGAAATCGTCTTCTGTGC; reverse, CTTGGGCACCTTGATGTCTT. For the Myc gene: forward, TCCTGTACCTCGTCCGATTC; reverse, GGTTTGCCTCTTCTCCACAG.
Chromatin immunoprecipitation (ChIP) assay
ChIP assay with an established rat follicular thyroid cell line PCCL3 was performed as described previously (20) with slight modification. Briefly, PCCL3 cells (3 × 106 cells) transfected with KrasG12D and ThrbPV genes were fixed in 1% of formaldehyde for 10 min and quenched by addition of glycine with a 0.125 M final concentration for 5 min. Immunoprecipitation was carried out using Anti-BRD4 (A301-985A50) or IgG as a negative control. Subsequent steps in ChIP assays were carried out as described previously (20). The following primers were used for Chip assay. For the Myc gene: forward, CCTCGGAACCACTTTTTCCT; reverse, TACTGGCCACAGATCACAGC. For the Ccnd1 gene: forward, AGCCCTAAGTTCTCGTGCAG; reverse, CCAGCGAGGAGGAATAGATG.
Statistical analysis
All data are expressed as mean ± standard error. Statistical analysis was performed and p < 0.05 was considered significant. All statistical tests were two-sided. GraphPad Prism version 5.0 for Mac OS X was used to perform analyses of variances.
Results
JQ1 inhibits thyroid tumor growth in ThrbPV/PVKrasG12D mice
We treated ThrbPV/PVKrasG12D mice with vehicle or JQ1 for 10 weeks and evaluated the effect on thyroid carcinogenesis. Analysis of Kaplan-Meier cumulative survival curves showed that no ThrbPV/PVKrasG12D mice treated with JQ1 died during that period. By contrast, only 50% of ThrbPV/PVKrasG12D mice survived after 10 weeks of vehicle treatment (Figure 1A). Among the vehicle-treated mice, 50% of mice could not survive before reaching the experimental end-point of 10-week due to massive enlarged thyroid tumors. In contrast, all JQ1-treated mice survived to the end of 10-week treatment period. The differences between the survival rates of the ThrbPV/PVKrasG12D mice treated with vehicle only or with JQ1 were statistically significant (p=0.01). These results indicate that JQ1 treatment led to improved survival of ThrbPV/PVKrasG12D mice.
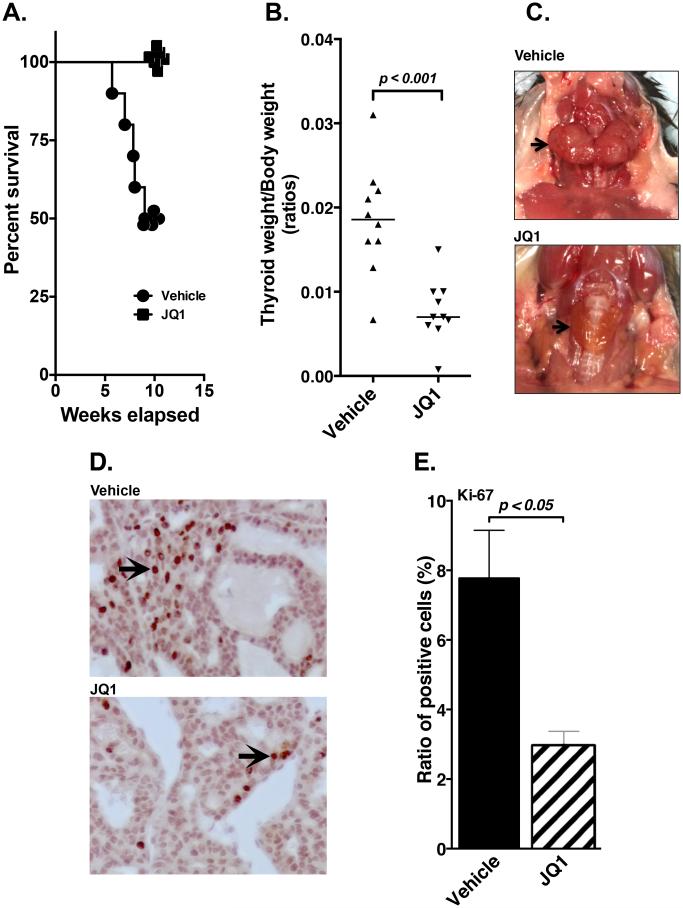
Figure 1.
JQ1treatment improves survival and decreases proliferation of thyroid tumor cells in JQ1-treated ThrbPV/PVKrasG12D mice. (A) The Kaplan–Meier survival curves for ThrbPV/PVKrasG12D mice treated with vehicle (n=10) or JQ1 (n=10) for 10 weeks. The Kaplan–Meier cumulative survival analysis was performed using GraphPad Prism version 5.0 for Mac OS X. Survival rates of ThrbPV/PVKrasG12D mice with vehicle treatment and mice with JQ1 treatment were significantly different (p<0.05) (B) Thyroid glands of the mice treated with vehicle (n=10) or JQ1 (n=10) were dissected and compared. The difference between the thyroid weights of ThrbPV/PVKrasG12D mice treated with vehicle or JQ1 was significant (p=0.0002). (C) Representative thyroid tumors from vehicle-treated or JQ1-treated ThrbPV/PVKrasG12D mice. (D) Thyrocyte proliferation in ThrbPV/PVKrasG12D mice was determined by immunohistochemical assessment of the Ki-67 proliferation marker, as described in Materials and Methods. Representative images of Ki-67-positive thyrocytes (indicated by arrows) in the mice treated with vehicle or JQ1 is shown at × 400 magnification. (E) Quantification of Ki-67-positive cells in thyroid sections of ThrbPV/PVKrasG12D mice treated with vehicle or JQ1. The percentage of Ki-67-positive cells and total cells was determined from 10–12 bright fields.
Enlarged thyroid tumor mass is one of the major causes for the death of mice. We examined thyroid weight of the mice treated with vehicle or JQ1. In vehicle-treated ThrbPV/PVKrasG12D mice the weight of thyroid was 374.9 ± 46.39 mg (n=10) whereas in JQ1-treated mice it was 158.4 ± 27.40 mg (n=10), a significant difference (Figure 1B). The representative thyroid tumors after vehicle or JQ1 treatment are shown in Figure 1C. It is clear that with JQ1 treatment there was decreased tumor growth with less vascularity (lower panel, Figure 1C). These results indicate that JQ1 decreased thyroid tumor growth of ThrbPV/PVKrasG12D mice, resulting in their improved survival. However, it is of interest to note that JQ1 did not affect thyroid malignant progression of ThrbPV/PVKrasG12D mice. The frequency of occurrence of capsular invasion, vascular invasion, anaplasia and lung metastases was similar between vehicle-treated and JQ1-treated ThrbPV/PVKrasG12D mice (Supplemental Table I).
JQ1 decreases proliferation of thyroid tumor cells
To determine whether the decreased thyroid growth was due to decreased proliferation of tumor cells in ThrbPV/PVKrasG12D mice, we examined Ki-67-positive cells in the thyroids of mice (Figure 1D). Ki-67-positive thyroid epithelial cells were clearly detected in thyroid sections of ThrbPV/PVKrasG12D mice treated with vehicle (Figure 1D). In contrast, only a few were detected in the thyroid sections of ThrbPV/PVKrasG12D mice after JQ1 treatment (Figure 1D). To quantify the percentage of cells undergoing cell proliferation, we calculated the average ratio of Ki-67-positive cells to total cells from 10–12 bright fields at high magnification (× 400) of each section. The quantitative data are shown in Figure 1E. In the mice treated with vehicle, 7.8% of thyroid epithelial cells were actively proliferating (Figure 1E). In ThrbPV/PVKrasG12D mice treated with JQ1, the ratio was only 3.0% (Figure 1E), indicating a 60% reduction in the proliferation of thyroid epithelial cells of ThrbPV/PVKrasG12D mice. However, it is important to point out that the Ki-67 immunohistochemical sections did not show any correlation of positive nuclei with histologic type and, with JQ1 treatment, this lack of correlation with histology persisted. Nonetheless, these findings suggested that decreased proliferation contributed to the significant reduced thyroid weight of ThrbPV/PVKrasG12D mice after JQ1 treatment.
Differential gene expression profiles in the thyroid tumors of ThrbPV/PVKrasG12D mice treated with vehicle or JQ1
To identify the factors contributing to JQ1-induced suppression of thyroid tumors in ThrbPV/PVKrasG12D mice, we carried out microarray analysis using thyroid tumor samples of mice treated with vehicle or JQ1 (n=3 for each treatment). A hierarchical clustering analysis shows that the gene expression profiles in the mice treated with JQ1 were distinct from those in the mice treated with vehicle (Figure 2A). Analysis of the array data between the two groups of mice showed that 82 genes were differentially expressed (>2.0-fold change, adjusted p < 0.05). Of those genes, 38 were upregulated and 44 were downregulated in thyroid tissues of the JQ1-treated mice.
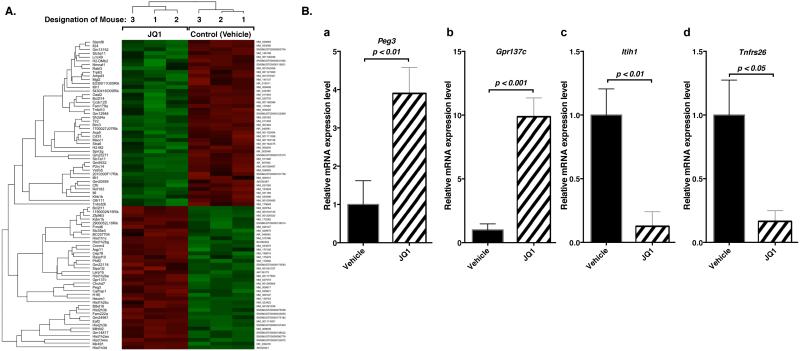
Figure 2.
Global differential gene expression analysis of thyroid tumors in JQ1- or vehicle-treated ThrbPV/PVKrasG12D mice. (A) A heat-map presentation of hierarchical clustering analysis (average of Euclidean distance) of the most highly expressed genes selected by fold change (>2) and adjusted p values (<0.05) with BH method (50) from the mice treated with vehicle or with JQ1. (B) The mRNA expression levels of the Gpr137c, Peg3, Itih1, and Tnfrsf26 genes were measured by real-time RT–PCR in thyroid tumors. Reactions were performed in 200 ng of total RNA. The Gapdh gene was used as a reference.
Table 1 lists the top 13 genes whose expression was increased by JQ1. Among these transcripts, 8 of 13 genes were in the category of histones and histone modification (Hist1h3d, Hist1h2bg, Hist1h2ao, Hist2h3b, Hist1h2bc, Hist1h4m, Hist1h1c, and Hexim1). HEXIM1 is critical for histone modification. Our gene expression profiling suggested that JQ1 treatment induced histone remodeling, leading to the expression changes of proteins critical for thyroid tumor growth. Therefore, we chose the genes essential for cell proliferation for the validation of our gene expression data. Beside histone proteins, two transcripts (Peg3 and Gpr137c) are related to suppression of tumor cell growth (21, 22). Among the top downregulated genes after JQ1 treatment, two transcripts (Itih1 and Tnfrsf26) are associated with tumor growth-promoted cell signaling (23). Consistently with their reported functions, we observed an increased mRNA expression of the Peg3 and Gpr137c genes in ThrbPV/PVKrasG12D mice by JQ1 treatment as compared with vehicle treatment (Figure 2B-a & b). In contrast, we observed that mRNA expression of the Itih1 and Tnfrsf26 genes in ThrbPV/PVKrasG12D mice with JQ1 treatment was lower than with vehicle treatment (Figures 2B-c and d). These data validated the differentially expressed genes identified in our gene expression analysis.
Table 1.
The expression of top 13 genes differentially increased by JQ1
| ID | Gene Symbol | Chromosome | Fold Change | Adjusted P Value |
|---|---|---|---|---|
| 1 | Hist1h3d | 13 | 15.5 | 0.003 |
| 2 | Hist1h2bg | 13 | 7.5 | 0.003 |
| 3 | Hist1h2ao | 13 | 7.2 | 0.011 |
| 4 | Gpr137c | 14 | 5.1 | 0.011 |
| 5 | Hist2h3b | 3 | 4.3 | 0.003 |
| 6 | Hist1h2bc | 13 | 4.2 | 0.033 |
| 7 | Hist1h4m | 13 | 4.1 | 0.011 |
| 8 | Larp1b | 3 | 3.9 | 0.030 |
| 9 | Peg3 | 7 | 3.5 | 0.010 |
| 10 | Hexim1 | 11 | 3.3 | 0.005 |
| 11 | Cnnm4 | 1 | 3.1 | 0.010 |
| 12 | Aqp11 | 7 | 2.8 | 0.044 |
| 13 | Hist1h1c | 13 | 2.8 | 0.017 |
JQ1 decreases MYC expression in thyroid tumors of ThrbPV/PVKrasG12D mice
MYC is an oncoprotein critical in cell proliferation and has been identified as one of the major targets of JQ1 (6). We showed previously that the expression of the Myc gene is highly elevated at both the mRNA and the protein levels in thyroid tumors of ThrbPV/PVKrasG12D mice (17, 19). Elevated MYC expression is associated with aggressive thyroid tumor growth in ThrbPV/PVKrasG12D mice (17, 19). We therefore first evaluated the effect of JQ1 on the expression of Myc mRNA. Figure 3A shows that JQ1 suppressed the expression of Myc mRNA by 40%. However, intriguingly, our microarray analysis found no significant differential expression of the Myc gene between the vehicle- and the JQ1-treated groups (Figure 2A). To examine the discrepancy between the gene expression findings shown in Figure 3A and the microarrays analysis, we identified the sequences of the 24 mouse Myc probes used in the microarray. We used the cDNA sequence of the mouse Myc gene as a reference for DNA sequence alignments and found that 14 of 24 probes aligned with the Myc sequence, but 10 sequences did not. The validation of the JQ1 effect on the Myc expression shown by the independent Q/PCR analysis (Figure 3A) was further confirmed by the evaluation of the canonical transcriptional signatures of MYC-dependent genes. We also used Gene Set Enrichment Analysis (24) and found Myc-dependent gene sets were enriched in the JQ1-down-regulated genes (Supplemental Table II), among which the YU_MYC_TARGET_YU (25) gene set shows strong correlation (FDR q-val:0.006) with the JQ1 down-regulated genes (Figure 3B). Taken together, these results suggest that the expression of the Myc gene was suppressed by JQ1.
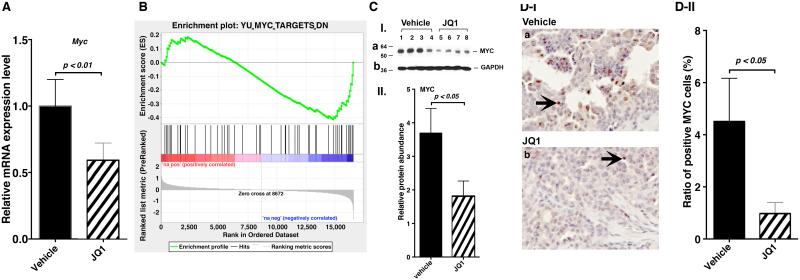
Figure 3.
JQ1 treatment suppresses the expression of the Myc gene in thyroid tumors of ThrbPV/PVKrasG12D mice. (A) The expression of the Myc mRNA in thyroid tumors of JQ1-treated ThrbPV/PVKrasG12D mice was suppressed. Total RNAs prepared from thyroid tumors of the ThrbPV/PVKrasG12D mice treated with vehicle or JQ1 were used for the analysis of RNA expression by Q/PCR analysis. (B). A representative enrichment plot: YU_MYC_TARGETS_UP Profile (25). One of the gene sets is significantly enriched in the downregulated genes by JQ1 treatment. (C-I-a) Western blot analyses for MYC in the thyroid tumors of ThrbPV/PVKrasG12D mice treated with vehicle or JQ1. Analysis with GAPDG (C-I-b) as loading control was carried out as described in Materials and Methods. (C-II) The band intensities were quantified by image analysis, and relative MYC abundance was determined using GAPDH as loading control. (D-I) Proliferation of thyroid follicular cells in ThrbPV/PVKrasG12D mice was shown by immunohistochemical analysis using anti-MYC antibody staining, as described in Materials and Methods. A representative image of MYC-positive thyrocytes (indicated by arrows) in the mice with vehicle or JQ1 treatment is shown at × 400 magnification. (D-II) Quantification of MYC-positive cells in thyroid sections of ThrbPV/PVKrasG12D mice treated with vehicle or JQ1. The ratio of MYC-positive cells and total cells was determined from 10–12 bright fields.
Having validated that the expression of the Myc gene was suppressed by JQ1 at the mRNA levels, we moved on to examine whether MYC was associated with thyroid tumor proliferation. We first evaluated the MYC protein abundance in thyroid tumors of ThrbPV/PVKrasG12D mice. MYC expression was first assessed at the protein level by Western blot analysis (Figure 3C-I-a). GAPDH was used as the loading control (Figure 3C-I-b). There was a relatively high level of the MYC protein in thyroid tumors of ThrbPV/PVKrasG12D mice treated with vehicle (n=4). After JQ1 treatment, the MYC protein abundance was significantly reduced (n=4). Quantitative analysis of MYC band intensities indicated that MYC abundance was reduced to 52.6% of that of ThrbPV/PVKrasG12D mice treated with vehicle (Figure 3C-II), suggesting the contribution of MYC in suppressing thyroid tumor growth. To further evaluate whether the decreased thyroid growth was associated with decreased MYC protein abundance, we examined MYC-positive cells in the thyroids of mice (Figure 3D). A high number of thyroid epithelial cells that were MYC positive were detected in thyroid sections of ThrbPV/PVKrasG12D mice with vehicle treatment (Figure 3D-I-a). In contrast, only a few were detected in the thyroid sections of ThrbPV/PVKrasG12D mice after JQ1 treatment (Figure 3D-I-b). To quantify the percentage of cells with MYC-positive cells, we calculated the average ratios of MYC-positive cells to total cells from 10-12 bright fields at high magnification (X 400) for each section. The quantitative data are shown in Figure 3D-II. In the mice treated with vehicle, 4.5% of thyroid epithelial cells were MYC-positive (Figure 3D-II). In ThrbPV/PVKrasG12D mice treated with JQ1, the ratio was decreased to 1.0% (Figure 3D-II), indicating a 71.4% reduction in the MYC-positive thyroid tumor cells of ThrbPV/PVKrasG12D mice. These findings indicate that decreased MYC abundance was associated with the significantly reduced thyroid weight of ThrbPV/PVKrasG12D mice after JQ1 treatment.
JQ1 reduces cyclin D-CDK4-Rb-E2F3-signaling in thyroid tumors of ThrbPV/PVKrasG12D mice
To dissect the downstream molecular events responsible for JQ1-induced inhibition of thyroid tumor growth in ThrbPV/PVKrasG12D mice, we examined altered cell signaling pathways involved in tumor growth and progression. It is known that MYC regulates, cyclin D1, cyclin-dependent protein kinase 4 (CDK4), retinoblastoma tumor suppressor protein (Rb), and E2F3 to affect cell proliferation (26). We therefore evaluated whether these regulators were affected in thyroid tumors of ThrbPV/PVKrasG12D mice treated with JQ1. The E2F3 transcription factor is a master regulator of proliferation (26). MYC and E2F3 collaborate together to drive potent cell proliferation (26). Rb binds directly to E2F3 proteins as a molecular scaffold and suppresses E2F3 functions. Phosphorylation of Rb by CDK4 relieves E2F3 to drive cell cycle progression (27). As expected, p-Rb (S780) protein levels were decreased in JQ1-treated ThrbPV/PVKrasG12D mice (Figure 4A-I-a) without apparent changes in the total Rb protein level (Figure 4A-I-b). The ratio of p-Rb (S780) to total Rb was decreased (Figure 4A-I-a and 4A-I-b). Quantitative analysis indicated that the ratio of p-Rb to total Rb after JQ1 treatment was reduced to 42.6% of that of ThrbPV/PVKrasG12D mice treated with vehicle (Figure 4A-II-a). Accordingly, JQ1 treatment decreased E2F3 protein abundance (Figure 4A-I-c). Quantitative analysis indicated that E2F3 abundance after JQ1 treatment was reduced to 54.7% of that of ThrbPV/PVKrasG12D mice with vehicle treatment (Figure 4A-II-b). These results indicate that reduced MYC abundance and decreased E2F3 functions are responsible for reduced thyroid tumor growth.
Figure 4.
JQ1 alters cyclin D1-CDK4-Rb-E2F3 signaling and AT2A/HEXIM1 abundance in thyroid tumors of ThrbPV/PVKrasG12D mice. (A-I) Total protein extracts were prepared from thyroid tumors. Western blot analysis was carried out for p-Rb(S780) (a), total Rb (b), E3F3 (c), CDK4 (d), cyclin D1 (e), KAT2A (f), HEXIM1 (g), and GAPDH (h), as described in Materials and Methods. (A-II) Quantitative analysis of relative protein expression levels of the ratio of p-Rb(S780) to total Rb (a), E3F3 (b), CDK4 (c), cyclin D1 (d), KAT2A (e), HEXIM1 (f) in mice treated with vehicle or JQ1. (B). Decreased Kat2a (panel a) and increased Hexim1 (panel b) mRNA expression in thyroid tumors of ThrbPV/PVKrasG12D mice treated with JQ1. Total RNAs prepared from thyroid tumors of the ThrbPV/PVKrasG12D mice treated with vehicle or JQ1 were used for the analysis of RNA expression.
Cyclin D1 is one of the major cyclins that interact with cyclin-dependent protein kinases (Cdks). In proliferating cells, cyclin D-Cdk complex accumulation is crucial for cell cycle progression. Specifically, the cyclin D-Cdk4/6 complex partially phosphorylates Rb, resulting in the release of the E2F3 important for cell cycle progression. The observation that p-Rb in thyroid tumors was reduced after JQ1 treatment prompted us to examine whether cyclin D1/Cdk4/6 was altered. The CDK4 protein level was lower in JQ1-treated ThrbPV/PVKrasG12D mice than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4A-I-d). Quantitative analysis indicated that CDK4 protein abundance after JQ1 treatment was reduced to 42.6% of that of ThrbPV/PVKrasG12D mice with vehicle treatment (Figure 4A-II-c). Similarly, cyclin D1 protein abundance was lower in JQ1-treated ThrbPV/PVKrasG12D mice than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4A-I-e). Quantitative analysis showed that cyclin D1 protein abundance after JQ1 treatment was reduced to 46.9% of that of ThrbPV/PVKrasG12D mice with vehicle treatment (Figure 4A-II-d). These data support the idea that JQ1 reduced MYC abundance and other regulators to suppress thyroid tumor growth.
We also examined whether the protein expression of other cellular effectors was changed after JQ1 treatment. MYC recruits K (lysine) acetyltransferase 2A (KAT2A) for transcription activation (28, 29). KAT2A protein abundance was lower in JQ1-treated ThrbPV/PVKrasG12D mice than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4A-I-f). Quantitative analysis indicated that KAT2A abundance after JQ1 treatment was reduced to 36.5% of that of ThrbPV/PVKrasG12D mice with vehicle treatment (Figure 4A-II-e). HEXIM1 is a crucial regulator of the transcription elongation factor b (P-TEFb) by inhibiting RNA polymerase II transcriptional elongation (26). It was reported that a BET inhibitor OTX015 induced the decreases of MYC accompanied by an HEXIM1 increase in acute leukemia cells (30). We found that HEXIM1 protein was higher in JQ1-treated ThrbPV/PVKrasG12D mice than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4A-I-g). Quantitative analysis indicated that HEXIM1 abundance in JQ1 treated ThrbPV/PVKrasG12D mice was increased by 1.8 fold compared to vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4A-II-f).
We further examined the mRNA expression of Kat2a and Hexim1 genes. Similar to the JQ1-suppressed Myc mRNA expression (see Figure 3A), we found lower Kat2a mRNA expression in JQ1-treated ThrbPV/PVKrasG12D mice than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4B-a). In contrast, we observed that Hexim1 mRNA expression in JQ1-treated ThrbPV/PVKrasG12D mice was higher than in vehicle-treated ThrbPV/PVKrasG12D mice (Figure 4B-b). These data suggest that JQ1 affected the transcription of the MYC-regulated genes to suppress thyroid tumor growth.
JQ1 decreases the MYC abundance and inhibits proliferation in PCCL3 cells
The PCCL3 cell line is an established non-transformed rat thyroid cell line that has been used frequently to study the biology and gene regulation of the thyroid (31). This cell line has also been used as a model to study the carcinogenesis of the thyroid. For example, transfection of the paired box 8 (PAX8)-peroxisome proliferator-activated receptor γ (PPARγ) fusion protein (PAX8-PPARγ) into PCCL3 cells confers PCCL3 cells with the ability to invade and to form anchorage independent colonies (32). In addition, activation of the RAS oncogene induces the cancer phenotype of genomic instability in thyroid PCCL3 cells (33). Previously, we showed that co-expression of KRASG12D and TRβPV in PCCL3 cells led to a significant increase in Myc mRNA expression while neither KRASG12D nor TRβPV alone in PCCL3 cells significantly increased expression (17). Consistently, Western blot analysis has shown that neither KRASG12D nor TRβPV alone increased the abundance of MYC proteins whereas their co-expression significantly increased that protein abundance (17). We examined whether JQ1 treatment reduced MYC in the PCCL3 cells stably expressing both KRASG12D and TRβPV (PCCL3-PV KRASG12D cells). As shown in Figure 5A, MYC protein abundance was significantly reduced after JQ1 treatment. Quantitative analysis indicated that the MYC protein abundance in PCCL3 cells with JQ1 treatment was reduced to 39.2% of that in PCCL3-PV KRASG12D cells with vehicle treatment (Figure 5B). Moreover, QPCR analysis showed that the Myc mRNA was also decreased in JQ1-treated PCCL3-PV KRASG12D cells (Figure 5C). The results showed that PCCL3-PV KRASG12D cells had a similar response to JQ1 treatment as seen in thyroid tumors of ThrbPV/PVKrasG12D mice.
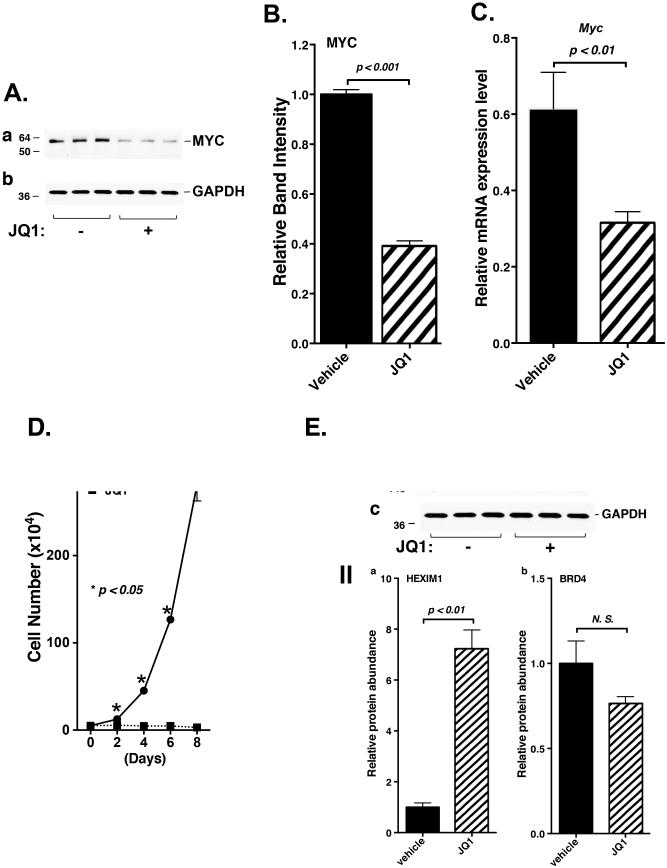
Figure 5.
JQ1 treatment reduced cell proliferation and decreases the protein abundance critical for cell proliferation in rat thyroid PCCL3 cells. (A) Protein level of MYC in the PCCL3 cells stably transfected with vector or vector expressing both KRASG12D and TRβPV. Total protein extracts were prepared from the PCCL3 cells 24 hr after the cells were treated with vehicle or JQ1. (B). Quantitative analysis of relative protein expression level of MYC in the mice treated with vehicle or JQ1. (C) The mRNA level of the Myc gene was measured by real-time RT–PCR in the PCCL3 cells stably expressing both KRASG12D and TRβPV or Neo-control only. Reactions were performed in 200 ng of total RNA. Gapdh gene was used as a reference. (D) Cell proliferation in the PCCL3 cells expressing both KRASG12D and TRβPV treated with vehicle or JQ1. Cell growth was analyzed as described in Materials and Methods. Data are expressed as mean ± standard error (SE) (n=3). Cell lines with treatment are as marked. (E). Protein level of BRD4 and HEXIM1 in the PCCL3 cells stably expressing both KRASG12D and TRβPV. Total protein extracts were prepared from the PCCL3 cells 24 hr after the cells were treated with vehicle or JQ1. (F-a) JQ1 treatment reduced the recruitment of BRD4 on the promoter of the Myc gene. ChIP assay was carried out using normal rabbit IgG or BRD4 antibody as described in Materials and Methods. (F-b) JQ1 treatment reduced the recruitment of BRD4 on the promoter of the Ccnd1 gene. ChIP assay was carried out using normal rabbit IgG or BRD4 antibody as described in Materials and Methods.
We further evaluated whether the JQ1 treatment could affect the proliferation of the PCCL3-PV KRASG12D cells. Figure 5D shows that the proliferation rate of PCCL3-PV KRASG12D cells treated with JQ1 was markedly lower than that of the control PCCL3-PV KRASG12D cells treated with vehicle. These results indicate that JQ1 treatment led to proliferation inhibition of PCCL3-PV KRASG12D cells as it did in thyroid tumor cells of ThrbPV/PVKrasG12D mice.
JQ1 inhibits binding of BDR4 to the promoter of the Myc gene
Besides JQ1-induced inhibition of cell proliferation, we also examined whether JQ1-induced inhibition of the PCCL3 proliferation was associated with the changes of protein abundance important for histone remodeling as observed in thyroid tumors. We examined BRD4 and HEXIM1 proteins for which these two regulators had opposite effects on RNA polymerase II transcriptional elongation. HEXIM1 sequesters P-TEFb to inhibit RNA polymerase II transcriptional elongation, resulting in transcription repression. Western blotting analysis showed that the band intensities of HEXIM1 proteins after JQ1 treatment in PCCL3-PV KRASG12D cells were significantly elevated, as we observed in thyroid tumors (Figure 5E-I-a and Figure 5E-II-a). BRD4 plays a crucial role in chromatin remodeling in the Myc promoter (6). To assess whether the BRD4 protein abundance was affected by JQ1, we also carried out Western blot analysis to determine BRD4 protein levels in the PCCL3-PV KRASG12D cells. Figure 5E-I-b and Figure 5E-II-b show that the band intensities of BRD4 proteins after JQ1 treatment were not significantly altered. These data indicated that reduced MYC abundance after JQ1 treatment was not due to decreased BRD4 protein abundance. The finding that JQ1 treatment did not alter abundance of BRD4 protein prompted us to consider whether JQ1 could suppress recruitment of BRD4 to the Myc promoter. Using PCCL3-PV KRASG12D cells, we carried out ChIP assays to evaluate whether the recruitment of BRD4 to the Myc promoter was decreased. After JQ1 treatment, the recruitment of BRD4 to the Myc promoter was reduced to 50.5% of that seen in PCCL3-PV KRASG12D cells treated with vehicle (Figure 5F-a). These results indicate that, after JQ1 treatment, a reduced level of BRD4 was recruited to the Myc promoter, resulting in reduced Myc expression. We also examined whether JQ1 could suppress recruitment of BRD4 to the promoter of other genes that were affected by JQ1 treatment. By ChIP analysis we found that, after JQ1 treatment, the recruitment of BRD4 to the Ccnd1 promoter was significantly reduced (Figure 5F-b) in PCCL3-PV KRASG12D cells. These findings supported the notion that decreased MYC abundance and other key regulators are responsible for JQ1-induced suppression of thyroid tumor growth in ThrbPV/PVKrasG12D mice.
Discussion
Using a mouse model of undifferentiated thyroid cancer, we showed that JQ1 was effective in prolonging survival and decreasing thyroid tumor growth. Molecular analyses demonstrated that the expression of the Myc gene was inhibited by JQ1 at both the mRNA and protein levels in the thyroid tumors of ThrbPV/PVKrasG12D mice. Previously, we have shown that elevated Myc expression contributes to the initiation of aggressive undifferentiated thyroid cancer in ThrbPV/PVKrasG12D mice (17, 19). The present studies showed that JQ1 inhibited tumor cell proliferation accompanied by Myc transcription repression in thyroid tumors of ThrbPV/PVKrasG12D mice. MYC has been associated with the regulation of growth factors and signaling pathways critical for tumor growth. In this study, we found that JQ1 treatment attenuated the cyclin D1-CDK4-Rb-CDK4-E2F3 signaling to decrease tumor cell proliferation in the thyroid of ThrbPV/PVKrasG12D mice. Therefore, the present study has elucidated that one mechanism by which JQ1 acts to inhibit tumor growth of ThrbPV/PVKrasG12D mice is through the inhibition of the Myc transcription program. This conclusion is supported by the gene expression array analysis in which we found that the MYC-regulated signature genes were enriched in JQ1-affected differentially regulated genes (Figure 2C and Supplemental Table 2). Moreover, using ChIP analysis, we have shown that JQ1 decreased the recruitment of BRD4 to the Myc transcription complex (Figure 5F-a). We also found that JQ1 increased the Heximi1 gene expression. BET proteins and HEXIM1 are two opposing regulators of the positive transcription elongation factor P-TEFb, which is the master modulator of RNA polymerase II during transcriptional elongation (34). Overexpressed HEXIM1 limits the availability of active positive transcription elongation factor P-TEFb. As a result of BET inhibition and increased HEXIM1, JQ1 altered the Myc transcription program. Thus, JQ1 could act to affect the Myc transcription program as others have shown for multiple myeloma and in acute lymphoblastic leukemia (6, 35, 36), but it could also act in thyroid tumors as demonstrated in the present studies.
Our differential gene expression data also suggest that JQ1 may change other genes important for thyroid tumor growth. We observed that seven histone 1 members are increased after JQ1 treatment. The histone H1 family comprises eleven members, each one transcribed by its own gene (37). Recent studies led to the recognition that epigenetic alterations of histones can have a profound impact on transcription programs in the absence of changes in DNA sequences and thus affect cellular behaviors such as differentiation and proliferation. Aberrant epigenetic alterations of histones are known to be involved in carcinogenesis (36, 38). Whether the elevated histone 1 members observed in our gene expression arrays had altered epigenetic changes in methylation, acetylation or the phosphorylation of the histone tails are currently not known. However, it is reasonable to expect that the increased expression of the seven members of histone H1 could lead to chromatin remodeling to impact genes involved in cell proliferation. Indeed, using ChIP assays, we found that less BRD4 was recruited to the Ccnd1 promoter complex in the thyroid tumors of JQ1-treated ThrbPV/PVKrasG12D mice, resulting in the suppression of the Ccnd1 expression. Cyclin D1 is a critical regulator in cell cycle progression and its deregulation is frequent in cancer (39). Moreover, activation of the cyclin-dependent kinases is a well-documented mechanism for their oncogenic actions (39). Taken together, our data support a mechanism in which the BET protein inhibitor JQ1 induces chromatin remodeling, resulting in transcription alteration of multiple genes to inhibit thyroid tumor growth in ThrbPV/PVKrasG12D mice (Figure 6).
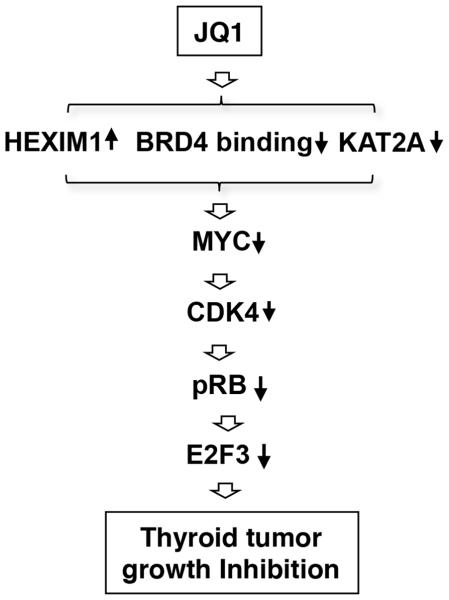
Figure 6. The proposed molecular pathway and affected regulators by which JQ1 acts to inhibit tumor growth of ThrbPV/PVKrasG12D mice.
JQ1 increases HEXIM1 and reduces KAT2A expression. JQ1 further inhibits the binding of BRD4 to the promoter of the Myc gene. These combined effects lead to reduced MYC expression and its downstream effectors to inhibit thyroid tumor growth.
The present studies showed the remarkable effect of JQ1 on the inhibition of tumor growth in ThrbPV/PVKrasG12D mice. However, under our experimental conditions, we did not detect changes in the frequency of occurrence of capsular invasion, vascular invasion, anaplasia, and lung metastasis (see Supplemental Table I). These observations raised the possibility that JQ1 could preferentially target the transcription program of a subset of genes selectively involved in growth regulation. This notion is supported by the observations that BRD4, a target for JQ, is generally involved in the transcription of active genes and is associated with most active enhancers and promoters (termed super-enhancers) in tumor cells (40). JQ1 caused a preferential loss of BRD4 at super-enhancers, resulting in preferential loss of transcription at super-enhancer-associated genes such as the MYC oncogene (40). Since MYC is a master regulator of cell proliferation (41), the cell proliferation, but not other tumor phenotypes (e.g., invasiveness), in thyroid tumors of ThrbPV/PVKrasG12D mice was selectively and preferentially inhibited. These observations further supported the idea that the proliferation and invasiveness of thyroid tumor cells most likely is regulated by different discrete set of genes during carcinogenesis. On the basis of these considerations, we showed that JQ1 clearly was effective in the inhibition of tumor growth to reduce tumor burden as a first line treatment. However, its combination with other agents that minimize tumor cell invasiveness would be needed to prevent metastasis and reduce mortality. At present, newly developed BET inhibitors are being tested in various phases of clinical trials for solid tumors, hematological cancers, and other clinical conditions (42-44). Our findings from the present pre-clinical studies would be of value in the consideration of using second-generation JQ1 analogs in combination with other agents in clinical trials for the treatment of patients with refractory and undifferentiated thyroid cancer. This notion is of importance in view of the recent findings from clinical trials of target therapy using tyroinse kinase inhibitors for differentiated and poorly differentiated thyroid cancer (45). Evidence is accumulating to suggest that one potential limitation in the target therapy is the eventual emergence of resistance to treatment (45). The reasons for the development of resistance from target therapy are not entirely clear. However, the possibility exists that BET inhibitors in combination with target therapy agents could overcome the resistance in thyroid cancer as shown for metastatic breast cancer (46, 47), for metastatic melanoma (48) and lung cancer (49). Thus, the present studies have raised new possibility for the effective treatment modalities for poorly differentiated and undifferentiated thyroid cancer.
Supplementary Material
Statement of translational relevance.
New therapeutic approaches are needed for patients with thyroid cancers refractory to radioiodine treatment. Inhibitors interfering with the interaction of bromodomain and extra terminal domain (BET) proteins with acetylated histones have potent anti-tumor effects in hematological malignancy and a few solid tumors. However, it is not known whether a BET inhibitor such as JQ1 is effective against thyroid cancer. Using a preclinical mouse model mimicking anaplastic thyroid cancer (ThrbPV/PVKrasG12D mice), we evaluated the effect of JQ1 on thyroid carcinogenesis. We found JQ1 treatment significantly prolonged survival, inhibited tumor growth, and attenuated transcriptional programs critical for tumor cell proliferation. Our preclinical findings indicated that use of BET inhibitors could be a promising therapeutic strategy for anaplastic thyroid cancer for which treatment options are very limited.
Acknowledgments
This research was supported by the Intramural Research Program of the Center for Cancer Research, National Cancer Institute, National Institutes of Health.
Footnotes
Disclosure of Potential Conflicts of Interest
The authors declare no conflicts of interest.
References
- 1.Sherman SI. Thyroid carcinoma. Lancet. 2003;361:501–11. doi: 10.1016/s0140-6736(03)12488-9. [DOI] [PubMed] [Google Scholar]
- 2.Lee HJ, Yun HJ, Kim S. Lenvatinib in radioiodine-refractory thyroid cancer. The New England journal of medicine. 2015;372:1868. doi: 10.1056/NEJMc1503150. [DOI] [PubMed] [Google Scholar]
- 3.Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. doi: 10.1016/j.cell.2007.02.005. [DOI] [PubMed] [Google Scholar]
- 4.Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012;149:214–31. doi: 10.1016/j.cell.2012.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–73. doi: 10.1038/nature09504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–17. doi: 10.1016/j.cell.2011.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mertz JA, Conery AR, Bryant BM, Sandy P, Balasubramanian S, Mele DA, et al. Targeting MYC dependence in cancer by inhibiting BET bromodomains. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:16669–74. doi: 10.1073/pnas.1108190108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lockwood WW, Zejnullahu K, Bradner JE, Varmus H. Sensitivity of human lung adenocarcinoma cell lines to targeted inhibition of BET epigenetic signaling proteins. Proceedings of the National Academy of Sciences of the United States of America. 2012;109:19408–13. doi: 10.1073/pnas.1216363109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cheng Z, Gong Y, Ma Y, Lu K, Lu X, Pierce LA, et al. Inhibition of BET bromodomain targets genetically diverse glioblastoma. Clinical cancer research : an official journal of the American Association for Cancer Research. 2013;19:1748–59. doi: 10.1158/1078-0432.CCR-12-3066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Shimamura T, Chen Z, Soucheray M, Carretero J, Kikuchi E, Tchaicha JH, et al. Efficacy of BET bromodomain inhibition in Kras-mutant non-small cell lung cancer. Clinical cancer research : an official journal of the American Association for Cancer Research. 2013;19:6183–92. doi: 10.1158/1078-0432.CCR-12-3904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Asangani IA, Dommeti VL, Wang X, Malik R, Cieslik M, Yang R, et al. Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer. Nature. 2014;510:278–82. doi: 10.1038/nature13229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gao X, Wu X, Zhang X, Hua W, Zhang Y, Maimaiti Y, et al. Inhibition of BRD4 suppresses tumor growth and enhances iodine uptake in thyroid cancer. Biochemical and biophysical research communications. 2016;469:679–85. doi: 10.1016/j.bbrc.2015.12.008. [DOI] [PubMed] [Google Scholar]
- 13.Xing M. Clinical utility of RAS mutations in thyroid cancer: a blurred picture now emerging clearer. BMC medicine. 2016;14:12. doi: 10.1186/s12916-016-0559-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Nikiforov YE, Nikiforova MN. Molecular genetics and diagnosis of thyroid cancer. Nature reviews Endocrinology. 2011;7:569–80. doi: 10.1038/nrendo.2011.142. [DOI] [PubMed] [Google Scholar]
- 15.Kaneshige M, Suzuki H, Kaneshige K, Cheng J, Wimbrow H, Barlow C, et al. A targeted dominant negative mutation of the thyroid hormone alpha 1 receptor causes increased mortality, infertility, and dwarfism in mice. Proceedings of the National Academy of Sciences of the United States of America. 2001;98:15095–100. doi: 10.1073/pnas.261565798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Suzuki H, Willingham MC, Cheng SY. Mice with a mutation in the thyroid hormone receptor beta gene spontaneously develop thyroid carcinoma: a mouse model of thyroid carcinogenesis. Thyroid : official journal of the American Thyroid Association. 2002;12:963–9. doi: 10.1089/105072502320908295. [DOI] [PubMed] [Google Scholar]
- 17.Zhu X, Zhao L, Park JW, Willingham MC, Cheng SY. Synergistic signaling of KRAS and thyroid hormone receptor beta mutants promotes undifferentiated thyroid cancer through MYC up-regulation. Neoplasia. 2014;16:757–69. doi: 10.1016/j.neo.2014.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kaneshige M, Kaneshige K, Zhu X, Dace A, Garrett L, Carter TA, et al. Mice with a targeted mutation in the thyroid hormone beta receptor gene exhibit impaired growth and resistance to thyroid hormone. Proceedings of the National Academy of Sciences of the United States of America. 2000;97:13209–14. doi: 10.1073/pnas.230285997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhu X, Zhu YJ, Kim DW, Meltzer P, Cheng SY. Activation of integrin-ERBB2 signaling in undifferentiated thyroid cancer. American journal of cancer research. 2014;4:776–88. [PMC free article] [PubMed] [Google Scholar]
- 20.Kim DW, Park JW, Willingham MC, Cheng SY. A histone deacetylase inhibitor improves hypothyroidism caused by a TRalpha1 mutant. Human molecular genetics. 2014;23:2651–64. doi: 10.1093/hmg/ddt660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zong G, Wang H, Li J, Xie Y, Bian E, Zhao B. Inhibition of GPR137 expression reduces the proliferation and colony formation of malignant glioma cells. Neurol Sci. 2014;35:1707–14. doi: 10.1007/s10072-014-1817-2. [DOI] [PubMed] [Google Scholar]
- 22.Jiang X, Yu Y, Yang HW, Agar NY, Frado L, Johnson MD. The imprinted gene PEG3 inhibits Wnt signaling and regulates glioma growth. J Biol Chem. 2010;285:8472–80. doi: 10.1074/jbc.M109.069450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Paris S, Sesboue R, Delpech B, Chauzy C, Thiberville L, Martin JP, et al. Inhibition of tumor growth and metastatic spreading by overexpression of inter-alpha-trypsin inhibitor family chains. Int J Cancer. 2002;97:615–20. doi: 10.1002/ijc.10120. [DOI] [PubMed] [Google Scholar]
- 24.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America. 2005;102:15545–50. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yu D, Cozma D, Park A, Thomas-Tikhonenko A. Functional validation of genes implicated in lymphomagenesis: an in vivo selection assay using a Myc-induced B-cell tumor. Ann N Y Acad Sci. 2005;1059:145–59. doi: 10.1196/annals.1339.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Humbert PO, Verona R, Trimarchi JM, Rogers C, Dandapani S, Lees JA. E2f3 is critical for normal cellular proliferation. Genes Dev. 2000;14:690–703. [PMC free article] [PubMed] [Google Scholar]
- 27.Miles WO, Dyson NJ. RB-loss puts focus on Myc. Nature cell biology. 2015;17:968–9. doi: 10.1038/ncb3213. [DOI] [PubMed] [Google Scholar]
- 28.Kenneth NS, Ramsbottom BA, Gomez-Roman N, Marshall L, Cole PA, White RJ. TRRAP and GCN5 are used by c-Myc to activate RNA polymerase III transcription. Proceedings of the National Academy of Sciences of the United States of America. 2007;104:14917–22. doi: 10.1073/pnas.0702909104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu X, Tesfai J, Evrard YA, Dent SY, Martinez E. c-Myc transformation domain recruits the human STAGA complex and requires TRRAP and GCN5 acetylase activity for transcription activation. J Biol Chem. 2003;278:20405–12. doi: 10.1074/jbc.M211795200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Coude MM, Braun T, Berrou J, Dupont M, Bertrand S, Masse A, et al. BET inhibitor OTX015 targets BRD2 and BRD4 and decreases c-MYC in acute leukemia cells. Oncotarget. 2015;6:17698–712. doi: 10.18632/oncotarget.4131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fusco A, Berlingieri MT, Di Fiore PP, Portella G, Grieco M, Vecchio G. One- and two-step transformations of rat thyroid epithelial cells by retroviral oncogenes. Mol Cell Biol. 1987;7:3365–70. doi: 10.1128/mcb.7.9.3365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vu-Phan D, Grachtchouk V, Yu J, Colby LA, Wicha MS, Koenig RJ. The thyroid cancer PAX8-PPARG fusion protein activates Wnt/TCF-responsive cells that have a transformed phenotype. Endocrine-related cancer. 2013;20:725–39. doi: 10.1530/ERC-13-0058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Saavedra HI, Knauf JA, Shirokawa JM, Wang J, Ouyang B, Elisei R, et al. The RAS oncogene induces genomic instability in thyroid PCCL3 cells via the MAPK pathway. Oncogene. 2000;19:3948–54. doi: 10.1038/sj.onc.1203723. [DOI] [PubMed] [Google Scholar]
- 34.Chen R, Yik JH, Lew QJ, Chao SH. Brd4 and HEXIM1: multiple roles in P-TEFb regulation and cancer. BioMed research international. 2014;2014:232870. doi: 10.1155/2014/232870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ott CJ, Kopp N, Bird L, Paranal RM, Qi J, Bowman T, et al. BET bromodomain inhibition targets both c-Myc and IL7R in high-risk acute lymphoblastic leukemia. Blood. 2012;120:2843–52. doi: 10.1182/blood-2012-02-413021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chaidos A, Caputo V, Gouvedenou K, Liu B, Marigo I, Chaudhry MS, et al. Potent antimyeloma activity of the novel bromodomain inhibitors I-BET151 and I-BET762. Blood. 2014;123:697–705. doi: 10.1182/blood-2013-01-478420. [DOI] [PubMed] [Google Scholar]
- 37.Scaffidi P. Histone H1 alterations in cancer. Biochimica et biophysica acta. 2015 doi: 10.1016/j.bbagrm.2015.09.008. [DOI] [PubMed] [Google Scholar]
- 38.Paluch BE, Naqash AR, Brumberger Z, Nemeth MJ, Griffiths EA. Epigenetics: A primer for clinicians. Blood Rev. 2016 doi: 10.1016/j.blre.2016.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Musgrove EA, Caldon CE, Barraclough J, Stone A, Sutherland RL. Cyclin D as a therapeutic target in cancer. Nat Rev Cancer. 2011;11:558–72. doi: 10.1038/nrc3090. [DOI] [PubMed] [Google Scholar]
- 40.Loven J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell. 2013;153:320–34. doi: 10.1016/j.cell.2013.03.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Dang CV. MYC, microRNAs and glutamine addiction in cancers. Cell Cycle. 2009;8:3243–5. doi: 10.4161/cc.8.20.9522. [DOI] [PubMed] [Google Scholar]
- 42.Ferri E, Petosa C, McKenna CE. Bromodomains: Structure, function and pharmacology of inhibition. Biochem Pharmacol. 2015 doi: 10.1016/j.bcp.2015.12.005. [DOI] [PubMed] [Google Scholar]
- 43.Moros A, Rodriguez V, Saborit-Villarroya I, Montraveta A, Balsas P, Sandy P, et al. Synergistic antitumor activity of lenalidomide with the BET bromodomain inhibitor CPI203 in bortezomib-resistant mantle cell lymphoma. Leukemia. 2014;28:2049–59. doi: 10.1038/leu.2014.106. [DOI] [PubMed] [Google Scholar]
- 44.Odore E, Lokiec F, Cvitkovic E, Bekradda M, Herait P, Bourdel F, et al. Phase I Population Pharmacokinetic Assessment of the Oral Bromodomain Inhibitor OTX015 in Patients with Haematologic Malignancies. Clin Pharmacokinet. 2016;55:397–405. doi: 10.1007/s40262-015-0327-6. [DOI] [PubMed] [Google Scholar]
- 45.Viola D, Valerio L, Molinaro E, Agate L, Bottici V, Biagini A, et al. Treatment of advanced thyroid cancer with targeted therapies: ten years of experience. Endocrine-related cancer. 2016;23:R185–205. doi: 10.1530/ERC-15-0555. [DOI] [PubMed] [Google Scholar]
- 46.Stratikopoulos EE, Dendy M, Szabolcs M, Khaykin AJ, Lefebvre C, Zhou MM, et al. Kinase and BET Inhibitors Together Clamp Inhibition of PI3K Signaling and Overcome Resistance to Therapy. Cancer Cell. 2015;27:837–51. doi: 10.1016/j.ccell.2015.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Stuhlmiller TJ, Miller SM, Zawistowski JS, Nakamura K, Beltran AS, Duncan JS, et al. Inhibition of Lapatinib-Induced Kinome Reprogramming in ERBB2-Positive Breast Cancer by Targeting BET Family Bromodomains. Cell Rep. 2015;11:390–404. doi: 10.1016/j.celrep.2015.03.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Paoluzzi L, Hanniford D, Sokolova E, Osman I, Darvishian F, Wang J, et al. BET and BRAF inhibitors act synergistically against BRAF-mutant melanoma. Cancer Med. 2016 doi: 10.1002/cam4.667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Xu C, Buczkowski KA, Zhang Y, Asahina H, Beauchamp EM, Terai H, et al. NSCLC Driven by DDR2 Mutation Is Sensitive to Dasatinib and JQ1 Combination Therapy. Mol Cancer Ther. 2015;14:2382–9. doi: 10.1158/1535-7163.MCT-15-0077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practicaland powerful approach to multipletesting. J R Stat Soc Ser B. 1995;57:289–300. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.