FIG. 2.
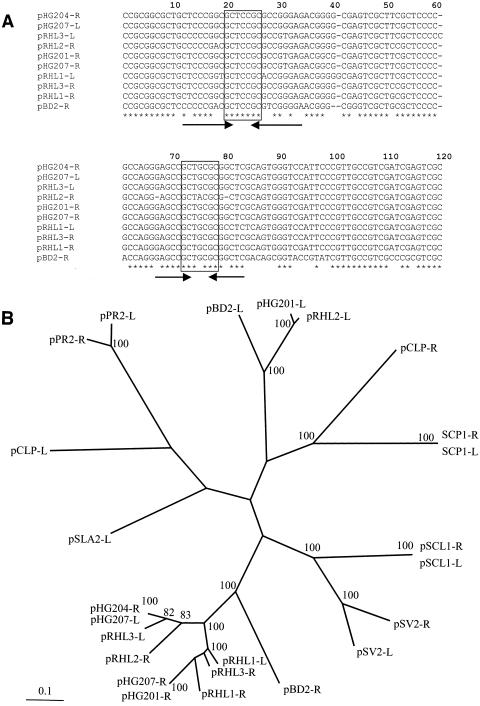
Sequence analysis of actinomycete invertron telomeres. (A) Alignment of rhodococcal invertron telomere nucleotide sequences. The nucleotide sequences are derived from each of the three RHA1 invertrons (except for pRHL2-L), as well as pHG201 of R. opacus MR11, pHG204 of R. opacus MR22 (31), pBD2 of R. erythropolis (60), and pHG207 of R. sp. strain MR2253 (30). Strictly conserved nucleotides are indicated with hodococcus asterisks. The two sets of inverted repeats are indicated with arrows. The GCTXCGC central motif is boxed. (B) Radial view of best maximum-parsimony tree obtained by PHYLIP analyses of actinomycete telomeres. The first 800 nucleotides of each telomere were aligned. Sequences were taken from each of the plasmids in 2a, as well as the following invertrons: S. clavuligerus pSCL1 (65), S. coelicolor A3 SCP1, S. violaceoruber pSV2 (59), S. rochei 7434AN4 pSLA2-L (28), Planobispora rosea pPR1 and pPR2 (47), and M. celatum pCLP (46).