Fig. 10.
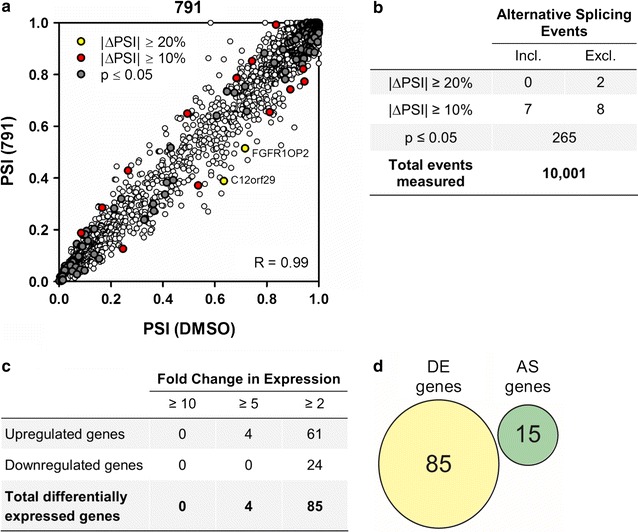
791 does not appreciably alter cellular gene expression or alternative splicing events. HeLa HIVrtTA∆Mls cells were incubated with 791 (30 µM) for 24 h. Cells were subsequently harvested for total RNA isolation and samples used for cDNA library preparation and Illumina sequencing. a Mean alternative splicing changes (PSI or percent spliced in) were plotted comparing DMSO and compound treatment (N = 2, RNA-seq). |ΔPSI| ≥ 10% and 20% are represented as red and yellow dots, respectively. AS genes with exon inclusion/exclusion ≥20% are labelled. Statistically significant alternative splicing changes with |ΔPSI| ≤ 10% are indicated by the gray dots (Student’s t test, two-tailed). Error bars not shown. b Summary of altered exon inclusion or exclusion (Incl. or Excl.) with compound treatment (RNAseq, N = 2). c Differentially expressed (DE) genes described as cRPKM fold change ≥2 or ≤0.5 with compound treatment relative to DMSO treatment (p ≤ 0.05, 11,406 genes, N = 2). Fold change distribution of differentially expressed genes based on compound treatment relative to DMSO treatment within the RNAseq dataset (p ≤ 0.05, 1020 genes, N = 2). d Venn diagrams comparing DE and AS (|ΔPSI| ≥ 10%) events with 791 (N = 2). See Additional files 10, 11: Tables S5 and S6 for a complete description of the data obtained
