Abstract
Retrotransposons are mobile genetic elements that colonize eukaryotic genomes by replicating through an RNA intermediate. As retrotransposons can move within the host genome, defense mechanisms have evolved to repress their potential mutagenic activities. In the fission yeast Schizosaccharomyces pombe, the mRNA of Tf2 long terminal repeat retrotransposons is targeted for degradation by the 3′–5′ exonucleolytic activity of the exosome-associated protein Rrp6. Here, we show that the nuclear poly(A)-binding protein Pab2 functions with Rrp6 to negatively control Tf2 mRNA accumulation. Furthermore, we found that Pab2/Rrp6-dependent RNA elimination functions redundantly to the transcriptional silencing mediated by the CENP-B homolog, Abp1, in the suppression of antisense Tf2 RNA accumulation. Interestingly, the absence of Pab2 attenuated the derepression of Tf2 transcription and the increased frequency of Tf2 mobilization caused by the deletion of abp1. Our data also reveal that the expression of antisense Tf2 transcripts is developmentally regulated and correlates with decreased levels of Tf2 mRNA. Our findings suggest that transcriptional and post-transcriptional pathways cooperate to control sense and antisense RNAs expressed from Tf2 retroelements.
Keywords: Pab2, Abp1, Tf2, retrotransposon, S. pombe
TRANSPOSABLE elements (TEs) are a ubiquitous feature of eukaryotic genomes that are divided into two major groups: DNA transposons and retrotransposons. Notably, it is estimated that more than half of the human genome consists of TEs, of which only retrotransposons are thought to be still active today (Cordaux and Batzer 2009). Retrotransposons are divided into two main classes: long terminal repeat (LTR) retrotransposons and non-LTR retrotransposons. The genome of the fission yeast Schizosaccharomyces pombe is known to accommodate LTR-type retrotransposons (Tf1 and Tf2) of the gypsy family. Whereas full-length Tf1 elements can only be found in wild S. pombe isolates, 13 full-length Tf2 elements are still present in most laboratory strains of S. pombe (Esnault and Levin 2015), representing a powerful tool for the study of gene regulation by LTR retrotransposons. In addition to 13 full-length Tf2 elements, the S. pombe genome contains ∼250 LTR-derived fragments (solo LTRs), of which 35 are derived from Tf2 elements (Bowen et al. 2003). Interestingly, ∼70% of the Tf2 mobilization events are mediated by homologous recombination between a newly synthesized cDNA and a preexisting copy of Tf2, whereas the remaining 30% of transposition events occur via integration into a new chromosomal locus (Hoff et al. 1998).
Movement of TEs represents a source of threat to the integrity of the genome of a living organism, as it can result in insertions, deletions, and chromosomal rearrangements (Mita and Boeke 2016). Since retrotransposons are not infectious and only propagate within a host cell, their transposition activity is intimately linked to cell survival. Accordingly, a variety of mechanisms to repress the expression of retrotransposons and transposition activity have emerged through the course of evolution. In S. pombe, Tf2 expression is strongly repressed by a set of conserved centromeric-binding proteins of the CENP-B family. Specifically, it was shown that the CENP-B homolog, Abp1/Cbp1, promotes histone deacetylation and heterochromatin formation at Tf2 loci by recruiting the histone deacetylases Clr3 and Clr6 (Cam et al. 2008). Binding of Abp1 to LTRs also promotes the clustering of Tf2 elements into specialized higher-order structures (Cam et al. 2008), which appear to restrict homologous recombination between cDNAs and preexisting Tf2 (Johansen and Cam 2015; Murton et al. 2016).
RNA interference (RNAi) is an additional mechanism to repress the expression and propagation of TEs in a variety of organisms (Dumesic and Madhani 2014). The S. pombe RNAi machinery consists of the dsRNA ribonuclease Dcr1, the Argonaute family protein Ago1, and the RNA-dependent RNA polymerase Rdp1 (Goto and Nakayama 2012), which contribute to heterochromatin-mediated silencing at centromeric and telomeric regions as well as at mating-type loci (Volpe et al. 2002; Verdel et al. 2004; Cam et al. 2005). Surprisingly, whereas inactivation of the RNAi pathway in vertebrates, insects, and plants results in the derepression of TE expression (Girard and Hannon 2008), S. pombe RNAi mutants show only a slight increase in Tf2 mRNA expression (Hansen et al. 2005; Woolcock et al. 2011). Moreover, only trace amounts of Tf2-derived small interfering RNAs (siRNAs) are detected after deep sequencing of total small RNAs (Woolcock et al. 2011), consistent with the minor role of the RNAi pathway in Tf2 silencing under optimal S. pombe growth conditions. In contrast, cells deficient for Rrp6, a component of the RNA exosome complex of 3′–5′ exonucleases, produce substantial levels of Tf2-specific siRNAs and show a strong dependence on the RNAi machinery for Tf2 silencing (Yamanaka et al. 2013). These findings have led to a model in which the RNA exosome and the RNAi machinery function redundantly in the repression of Tf2 expression in S. pombe.
The exonucleolytic activity of Rrp6 also contributes to the post-transcriptional regulation of meiotic differentiation genes during the mitotic cell cycle of S. pombe. This exosome-dependent RNA degradation pathway requires polyadenylation of meiotic transcripts by the canonical poly(A) polymerase (Yamanaka et al. 2010) and poly(A) tail recognition by the nuclear poly(A)-binding protein Pab2 (St-Andre et al. 2010; Yamanaka et al. 2010; Chen et al. 2011), which physically associates with the RNA exosome (Lemay et al. 2010; Egan et al. 2014; Zhou et al. 2015). The Pab2/Rrp6-dependent RNA processing pathway also contributes to the maturation of polyadenylated snoRNA precursors (Lemay et al. 2010; Larochelle et al. 2012) and the degradation of inefficiently spliced transcripts (Lemieux et al. 2011). Interestingly, Pab2 is required for the accumulation of Tf2-specific siRNAs in an rrp6Δ mutant (Yamanaka et al. 2013); yet, how Pab2 contributes to siRNA production remains elusive. Furthermore, it is still unknown whether Pab2 contributes to the regulation of Tf2 expression.
Here we show that Pab2 negatively controls the expression of sense and antisense Tf2 transcripts. We also found that the absence of Pab2 and Rrp6 attenuated the derepression of Tf2 transcription caused by the deletion of abp1. Consistent with this, the increased frequency of Tf2 transposition detected in abp1Δ cells was reduced in the absence of Pab2. The attenuation of Tf2 derepression resulting from the deletion of pab2 in the abp1Δ mutant correlated with increased levels of antisense Tf2 transcripts in the abp1Δ/pab2Δ double mutant compared to single mutants. We also provide evidence supporting that the antisense Tf2 transcripts stabilized in the absence of Pab2-dependent RNA decay promote the repression of Tf2 expression via an RNAi-dependent pathway. Finally, meiosis synchronization experiments indicated that antisense Tf2 transcripts were expressed during specific stages of the meiotic differentiation program that correlated with decreased levels of Tf2 mRNA. Our study discloses a Pab2-dependent RNA elimination pathway that represses sense and antisense Tf2 RNA accumulation.
Materials and Methods
Strains, media, and genetics methods
All of the S. pombe strains used in this study are listed in Supplemental Material, Table S1 and File S1. Strains were grown to exponential phase at 30° in yeast extract medium containing appropriate amino acid supplements (YES). Meiotic entry synchronization experiments were performed as follows: diploid h+/h+ pat1-114/pat1-114 cells were grown to OD600 of 1.0 at 25° in Edinburgh minimum medium (EMM) containing 2% glucose and 0.5% NH4Cl. Cells were then washed and suspended in EMM without NH4Cl (EMM-N) at OD600 of 0.5 and incubated at 25° for 14 hr. Entry into the meiosis program was induced by shifting the cells to 34° to inactivate the Pat1 kinase. PCR-mediated gene disruptions were performed by a two-step approach using 200–300 nucleotides annealing to the target gene, modified from Bähler et al. (1998). Gene knockouts were confirmed by colony PCR and RT-PCR. Plasmid integrations at the ade6 locus were performed as previously described (Lemieux et al. 2011) using linearized DNA and selection for adenine prototrophy. Strains are available upon request.
DNA constructs
The expression of GFP under the control of adh1 promoter and trx1 terminator was performed by cloning PCR products corresponding to the adh1 promoter (1000 bp), the gfp cDNA, and the trx1 terminator (1000 bp) into pBPade6 (Beaudoin and Labbe 2006), generating pFB445.
The Tf2AS expression construct was generated by PCR amplification of a ∼2000-bp fragment (2045 to 102 of Tf2 elements) from genomic DNA corresponding to the Tf2 antisense transcripts region. To reduce transcription from the strong Tf2 LTR promoter, nucleotides 146–154 corresponding to the TATA box of the Tf2 element were deleted. The PCR product was cloned between the adh1 promoter and the trx1 terminator, generating pFB914. All constructs were confirmed by DNA sequencing.
RNA analyses
For Northern blot analysis, 20 μg of total RNA was resolved on a 1% agarose-formaldehyde gel and transferred onto nylon membrane by capillary diffusion. Membranes were cross-linked using a UV Stratalinker and incubated using 32P-labeled strand-specific RNA probes that were generated using T7/SP6 RNA polymerase. The RNA probes used to detect sense and antisense Tf2 RNAs were complementary to nucleotides 118–293 and 663–792, respectively, of the Tf2 ORF sequence. After extensive washing steps, membranes were exposed to Phosphor Screens and visualized using a Typhoon Trio instrument. For coordinate mapping analysis of Tf2 transcripts, 5′ rapid amplification of cDNA ends (RACE) was performed using the 5′ RACE System from Invitrogen, which relies on terminal deoxynucleotidyl transferase activity to add homopolymeric tails to the 3′ ends of cDNA. 3′ RACE was performed using a ligation-mediated PCR approach in which a 5′ preadenylated and 3′ predideoxycytosined oligonucleotide was ligated to the 3′ end of RNAs. Before PCR analysis, RNase H digestion was performed to remove complementary RNA to the cDNA.
Retrotransposition assay
For retrotransposition assays, strains carrying Tf2-12 neoAI constructs (Sehgal et al. 2007) were inoculated at 5 × 104 cells/ml in 40 independent 2.5-ml cultures and grown to saturation at 30°. Each saturated culture was quantified for its cell density and then 1 × 108 cells from each culture were plated on a G418 plate (150 µg/ml) and neo-resistant colonies were counted. Retrotransposition rates analysis was conducted using the Ma-Sandri-Sarkar Maximum Likelihood Estimator (MMS-LLE) methods on FALCOR web-tool (Hall et al. 2009).
Chromatin immunoprecipitation (ChIP) assays
ChIP assays were carried out as previously described (Larochelle et al. 2012). Briefly, cells were cross-linked with formaldehyde and lysed using a FastPrep instrument. Lysates were sonicated and chromatin extracts were incubated with an antibody specific for the C-terminal domain of RNA polymerase II (8WG16) that was previously coupled to protein-G sepharose. RNA polymerase II density was calculated as the enrichment of a specific Tf2 locus relative to a nontranscribed intergenic region (nucleotides 3,009,380–3,009,484 of chromosome I), as previously described (Larochelle et al. 2012). Quantification of the immunoprecipitated DNA was done by quantitative real-time PCR. ChIP assays using a control IgG antibody were also carried for all experiments, but were not shown since they were negative.
Antibodies
Rabbit polyclonal anti-Tf1 reverse transcriptase (RT) was a generous gift from Dr. H. L. Levin (National Institutes of Child Health and Human Development, National Institutes of Health, Bethesda, MD). Mouse monoclonal anti-α-tubulin B-5-1-2 was from Sigma. Alexa Fluor 680 goat anti-mouse was from Invitrogen. Donkey anti-rabbit IR Dye 800cw was from Odyssey. Mouse monoclonal anti-RNA polII CTD 8WG16 was from Covance.
Western blot
For Western blot analysis, 300 μg of total proteins were resolved by SDS-PAGE and transferred onto nitrocellulose membranes, which were incubated with antibodies against Tf1 RT (1:10,000) and tubulin α (1:5000). Membranes were then incubated with anti-rabbit (1:15,000) and anti-mouse (1:15,000) secondary antibodies analyzed using an Odyssey infrared imaging system.
Data availability
Strains and plasmids are available upon request. Table S1 contains all strains used in this study.
Results
Pab2 represses Tf2 mRNA expression
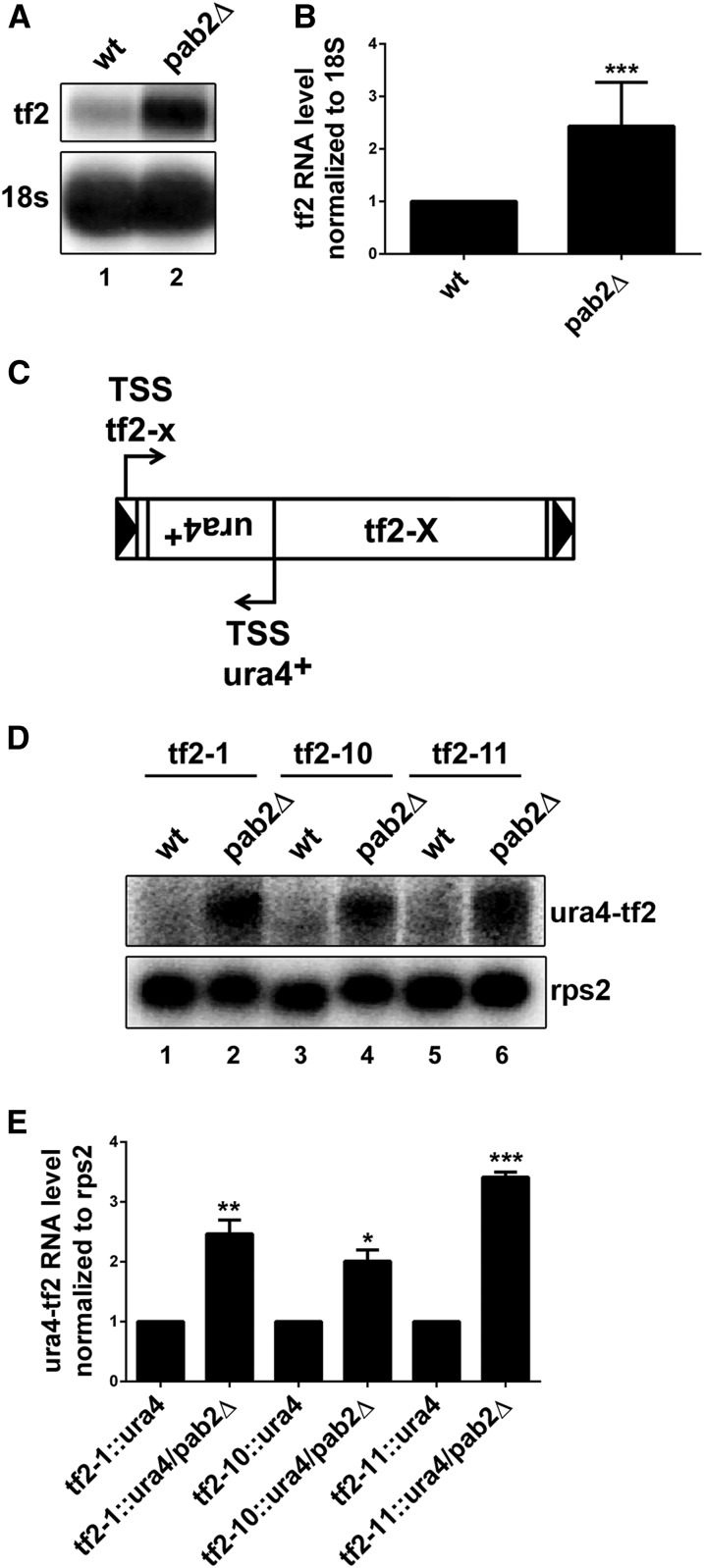
An analysis of gene expression changes in S. pombe pab2-null cells using DNA microarray revealed mRNA accumulation from Tf2 retrotransposons (Lemay et al. 2010). To independently validate this observation, we used Northern blotting to compare total RNA prepared from wild-type and pab2Δ cells, which confirmed the significant increase in Tf2 mRNA in the pab2Δ mutant (Figure 1, A and B). Analysis of Tf2 transcription start sites by 5′ RACE in the pab2 mutant generally mapped to 180-nt from the 5′ end of the LTR, which is consistent with previous observations (Rhind et al. 2011; Persson et al. 2016). As the S. pombe genome contains 13 copies of full-length Tf2 elements that are roughly 99% identical, discriminating between the mRNAs expressed from the different Tf2 elements using hybridization-based approaches is challenging. To investigate if the increased expression of Tf2 mRNA in the pab2Δ mutant originated from one specific locus or multiple Tf2 loci, we used a set of 13 strains in which each Tf2 element is independently tagged by the integration of a ura4 reporter gene (Anderson et al. 2009) (see Figure 1C). Accordingly, we deleted pab2 in three randomly selected strains from this set of 13 strains, where Tf2-1, Tf2-10, and Tf2-11 loci were individually tagged using ura4. Northern blot analysis using a ura4+-specific probe confirmed mRNA accumulation originating from all three Tf2 loci in pab2Δ cells (Figure 1, D and E). From these results, we conclude that Pab2 negatively controls Tf2 mRNA expression in S. pombe.
Figure 1.
Pab2 represses expression of Tf2 mRNA. (A) Northern blot analysis of total RNA prepared from wild-type (lane 1) and pab2Δ (lane 2) cells. The blot was hybridized using probes complementary to Tf2 open reading frame and 18s rRNA. (B) Quantification of Northern blots data for Tf2 mRNA levels. The data and error bars represent the average and SD from at least three independent experiments. *** P < 0.001; Student’s t-test. (C) Schematic representation of an individual ura4+-tagged Tf2 element. (D) Northern blot analysis of total RNA prepared from wild-type (lanes 1, 3, and 5) and pab2Δ (lanes 2, 4, and 6) cells expressing Tf2-1 (lanes 1 and 2), Tf2-10 (lanes 3 and 4), and Tf2-11 (lanes 5 and 6) ura4+-tagged constructs. The blot was hybridized using probes complementary to ura4+ antisense sequence and the rps2 mRNA. (E) Quantification of Northern blots data for ura4+ antisense RNA levels. The data and errors bars represent the average and SEM from at least three independent experiments. * P < 0.05, ** P < 0.01, *** P < 0.001; two-way ANOVA including Sidak’s multiple comparison test.
The absence of Pab2 and Rrp6 attenuates the Tf2 derepression caused by the deletion of abp1
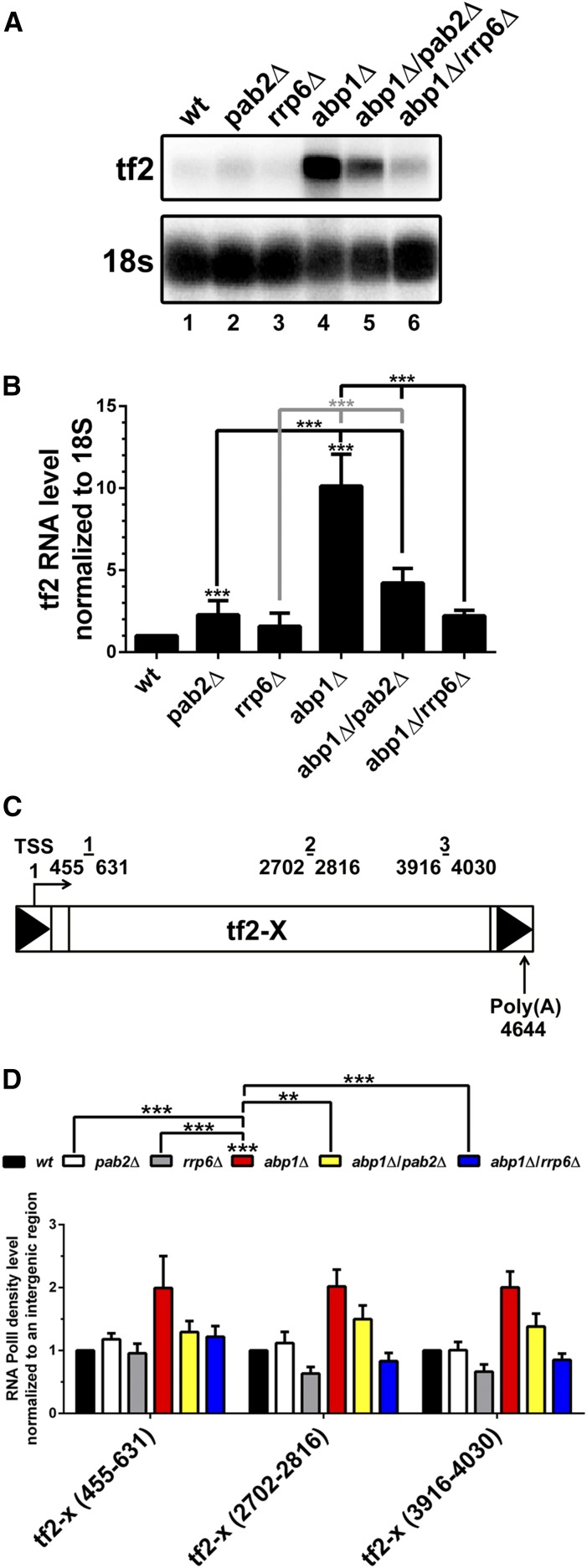
Abp1, a fission yeast CENP-B homolog, is a key factor involved in the transcriptional repression of Tf2 elements (Cam et al. 2008). Interestingly, high-throughput screening of genetic interactions in fission yeast reported a negative genetic interaction between pab2 and abp1 (Roguev et al. 2008), an observation that we have independently validated (Figure S1). To further characterize this genetic interaction at the molecular level, we compared the expression of Tf2 mRNA in the abp1Δ/pab2Δ double mutant to abp1Δ and pab2Δ single mutants by Northern analysis. As shown in Figure 2A (lane 4; quantifications in Figure 2B), high levels of Tf2 mRNA were detected in the absence of Abp1, consistent with previous observations (Cam et al. 2008). Unexpectedly, although Abp1 and Pab2 both negatively regulate Tf2 expression, the abp1Δ/pab2Δ double mutant showed reduced Tf2 mRNA levels relative to the abp1Δ single mutant (Figure 2A, compare lanes 4–5; Figure 2B). Because Rrp6, a 3′–5′ exonuclease that can be found associated with the nuclear exosome (Allmang et al. 1999), functions with Pab2 in the negative regulation of meiotic differentiation genes (St-Andre et al. 2010; Yamanaka et al. 2010) and specific intron-containing genes (Lemieux et al. 2011), we sought to examine the effect of deleting rrp6 on Abp1-mediated Tf2 repression. Notably, as seen for Pab2, we found that the abp1Δ/rrp6Δ double mutant showed a marked decrease in Tf2 mRNA accumulation as compared to the single abp1Δ mutant (Figure 2A, compare lanes 4 and 6; Figure 2B).
Figure 2.
Deletion of pab2 and rrp6 attenuates the Tf2 derepression observed in the absence of Abp1. (A) Northern blot analysis of total RNA prepared from wild-type (lane 1), pab2Δ (lane 2), rrp6Δ (lane 3), abp1Δ (lane 4), abp1Δ/pab2Δ (lane 5), and abp1Δ/rrp6Δ (lane 6) cells. The blot was hybridized using probes complementary to the Tf2 open reading frame and 18s rRNA. (B) Quantification of Northern blots data for Tf2 mRNA levels. The data and error bars represent the average and SD from at least three independent experiments. *** P < 0.001; Student’s t-test. (C) Schematic representation of Tf2 genes. Bars above the Tf2 retrotransposon show the positions of PCR products used for the ChIP analysis. Arrows indicates transcriptional start site (TSS) and polyadenylation site (Poly(A)). (D) ChIP assays measuring RNAPII density along Tf2 loci. The input and coprecipitated DNA were quantified by qPCR along the Tf2 loci using specific primer as shown in (C). ChIP data are presented as the mean of the relative level of RNAPII density normalized to a control intergenic region. Error bars represent the SEM for at least three independent experiments. ** P < 0.01, *** P < 0.001; two-way ANOVA including Tukey’s multiple comparison test.
We next examined whether the suppressive effect of Pab2 and Rrp6 deficiencies on Tf2 derepression in abp1Δ cells was mediated at the transcriptional level. For this, we measured RNA polymerase II (RNAPII) occupancy along Tf2 elements by chromatin immunoprecipitation (ChIP) assays (Figure 2C). As shown in Figure 2D, the pab2Δ single mutant demonstrated levels of RNAPII similar to wild-type cells, indicating that the increase in Tf2 mRNA levels detected in the absence of Pab2 is the result of defects in a post-transcriptional mechanism. In contrast, we found increased levels of RNAPII density at Tf2 loci in abp1Δ cells (Figure 2D), consistent with the role of Abp1 in Tf2 silencing via transcriptional repression (Cam et al. 2008). Notably, RNAPII occupancy at Tf2 was significantly reduced in abp1Δ/rrp6Δ and abp1Δ/pab2Δ double mutants compared to the abp1Δ single mutant (Figure 2D). Our results indicate that the attenuation of Tf2 derepression resulting from the absence of Pab2 and Rrp6 in abp1Δ cells is mediated by the restoration of transcriptional repression.
The increased frequency of Tf2 transposition in abp1Δ cells is reduced in the absence of Pab2
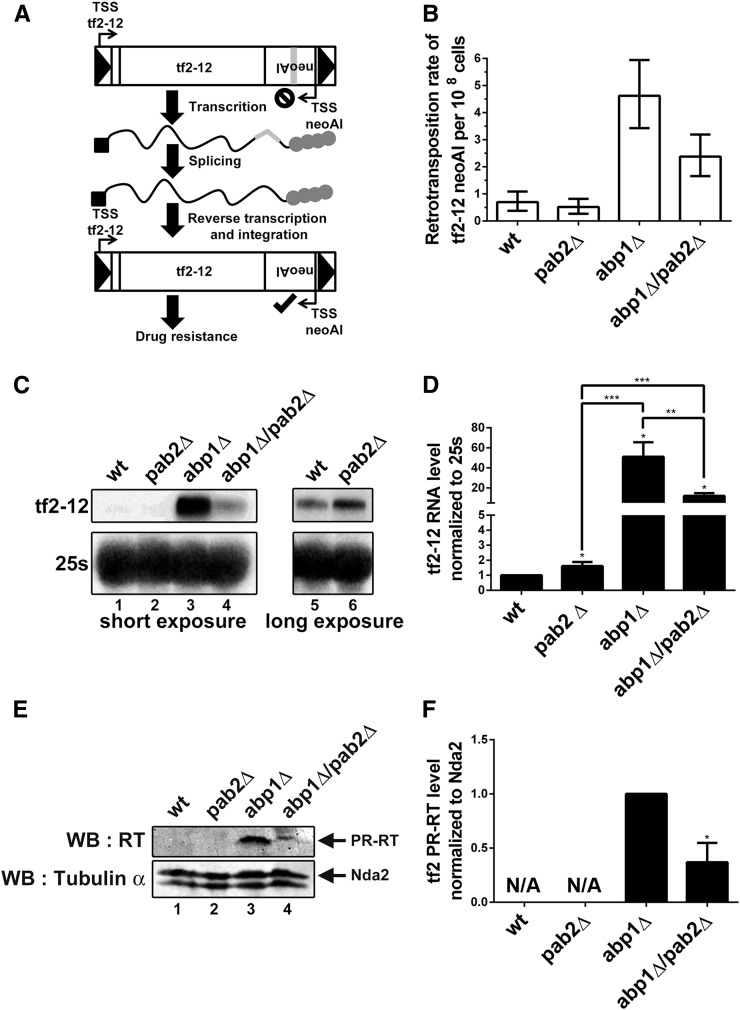
Given that the deletion of pab2 reduced the Tf2 derepression detected in abp1Δ cells, we examined the functional relevance of this attenuation on Tf2 transposition frequency. To measure Tf2 mobilization, we used the Tf2-12 neoAI strain (Sehgal et al. 2007) in which an intron-containing neomycin-resistant gene was inserted into the 3′ UTR of Tf2-12 in the opposite orientation to the Tf2-coding sequence (see Figure 3A). According to the organization of this construct, only cells in which splicing of the artificial intron was completed in the Tf2-12 mRNA will allow G418 resistance after a new Tf2-12 cDNA mobilization. The absence of Abp1 clearly increased the transposition frequency of Tf2-12 (Figure 3B), consistent with recent results analyzing Tf2 transposition frequency (Johansen and Cam 2015; Murton et al. 2016). In contrast, the ∼twofold increase in Tf2 mRNA level detected in pab2Δ cells did not result in a noticeable increase in the rate of Tf2 transposition (Figure 3B). However, as shown for Tf2 mRNA expression, the Tf2 transposition frequency in abp1Δ/pab2Δ cells was significantly lower than in the abp1Δ single mutant (Figure 3B).
Figure 3.
The absence of Pab2 reduces the mobilization frequency of Tf2 elements in the abp1Δ mutant. (A) Diagram of the system used for calculating the Tf2 retrotransposition rates. The endogenous Tf2-12 element was marked with a neoAI cassette, which is interrupted by an artificial intron. Mobilization of Tf2-12 mRNA with spliced neoAI cassette will allow expression of the drug resistance gene, thereby allowing growth on media containing G418. (B) Quantification of Tf2-12 retrotransposition rate by fluctuation analysis. The data and error bars represent the average and 95% confidence interval from 40 independent experiments. (C) Northern blot analysis of total RNA prepared from wild-type (lanes 1 and 5), pab2Δ (lanes 2 and 6), abp1Δ (lane 3), and abp1Δ/pab2Δ (lane 4) cells expressing Tf2-12 neoAI. The blots were hybridized using probes complementary to neoAI antisense RNA and the 25S rRNA. (D) Quantification of Northern blots data for Tf2-12 mRNA levels. The data and error bars represent the average and SD from at least three independent experiments.(* P < 0.05, ** P < 0.01, *** P < 0.001; Student’s t-test. (E) Western blot analysis of total proteins prepared from wild-type (lane 1), pab2Δ (lane 2), abp1Δ (lane 3), and abp1Δ/pab2Δ (lane 4) cells expressing Tf2-12 neoAI. The blot was analyzed using antibodies specific for the reverse transcriptase (RT) and tubulin. (F) Quantification of Western blots data for Tf2 protease-reverse transcriptase (PR-RT) levels. The data and errors bars represent the average and SD from at least three independent experiments. * P < 0.05; Student’s t-test.
To validate that Tf2-12 neoAI mRNA expression behaves similarly to endogenous Tf2, we analyzed Tf2-12 neoAI mRNA levels by Northern blotting using a probe specific to the neomycin resistance gene. As shown in Figure 3C, the expression profile of Tf2-12 neoAI was similar to endogenous Tf2 elements: an increase in the level of Tf2-12 neoAI mRNA in the abp1Δ single mutant, which was reduced by the deletion of pab2 (compare lanes 3–4; quantification in Figure 3D). In addition, the single deletion of pab2 resulted in increased levels of Tf2-12 neoAI mRNA (Figure 3C, lanes 5–6; Figure 3D), as seen for endogenous Tf2 (Figure 1). Western blot analysis also revealed the accumulation of Tf2 protease-reverse transcriptase (PR-RT) fusion in abp1Δ relative to wild-type cells (Figure 3E, compare lanes 1 and 3). Lower PR-RT levels were detected in the abp1Δ/pab2Δ double mutant as compared to the single abp1Δ mutant (Figure 3E, lanes 3–4; Figure 3F). PR-RT was not detected in extracts from wild-type and pab2Δ cells (Figure 3E, lanes 1–2), probably due to trace levels of this protein in these strains. These results show that in the absence of Tf2 silencing by Abp1, the attenuation of Tf2 derepression caused by the deletion of pab2 correlates with a reduction in the Tf2 mobilization rate.
Pab2 and Rrp6 repress the accumulation of Tf2 antisense RNAs
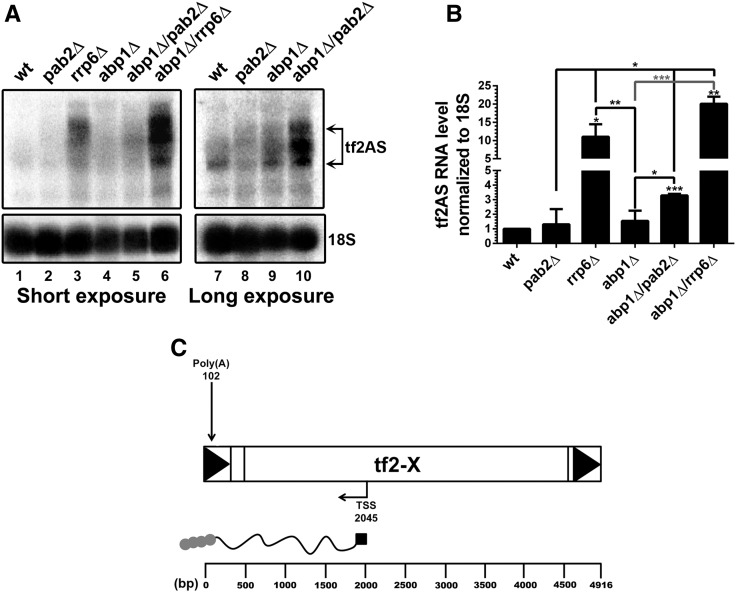
We next explored possible mechanisms that may explain how the absence of Pab2 or Rrp6 attenuated the Tf2 derepression observed in the abp1Δ mutant. One clue came from results showing that Tf2 elements are transcribed bidirectionally, generating sense and antisense transcripts (Lorenz et al. 2012). Given the role of Pab2 in Rrp6-mediated RNA turnover (Lemay et al. 2010; St-Andre et al. 2010; Yamanaka et al. 2010; Lemieux et al. 2011), we reasoned that the attenuating effects of pab2 and rrp6 deletions on Tf2 derepression in abp1Δ cells could be the consequence of Tf2 antisense RNA mis-regulation. To address whether Tf2 antisense (TF2AS) transcripts could be involved in the reduction of Tf2 mRNA levels in the double abp1Δ/pab2Δ mutant, we performed Northern blot analysis using RNA probes complementary to potential antisense Tf2 RNAs. As shown in Figure 4A, Tf2 antisense transcripts were clearly detected in rrp6Δ cells (lane 3), whereas no significant changes in Tf2AS RNA levels were observed in the abp1Δ and pab2Δ single mutants relative to the wild-type strain (Figure 4A, compare lanes 8–9 to lane 7; quantifications in Figure 4B). The major Tf2 antisense transcript was roughly 2000-nt-long and mapped to the 5′ half of Tf2 elements (Figure 4C). 5′ and 3′ RACE assays revealed that Tf2AS RNA had a major transcriptional start site (TSS) and a polyadenylation site located at nucleotides 2045 and 102 of Tf2 elements, respectively (Figure 4C and Table S2). Notably, we found that the levels of Tf2AS transcripts were significantly increased in both abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants relative to rrp6Δ, pab2Δ, or abp1Δ single mutants (Figure 4A, see lanes 6 and 10; Figure 4B). These results suggest that Pab2 and Rrp6 function in a parallel pathway to Abp1 in the repression of antisense Tf2 RNAs.
Figure 4.
Pab2 and Rrp6 function in a pathway redundant to Abp1 to suppress the accumulation of antisense Tf2 RNAs. (A) Northern blot analysis of total RNA prepared from wild-type (lanes 1 and 7), pab2Δ (lanes 2 and 8), rrp6Δ (lane 3), abp1Δ (lanes 4 and 9), abp1Δ/pab2Δ (lanes 5 and 10), and abp1Δ/rrp6Δ (lane 6) cells. The blots were hybridized using probes complementary to Tf2 antisense RNA and 18s rRNA. (B) Quantification of Northern blots data for Tf2 antisense RNA levels. The data and error bars represent the average and SD from at least three independent experiments. * P < 0.05, ** P < 0.01, *** P < 0.001; Student’s t-test. (C) Schematization of antisense Tf2 RNA. Arrows indicate transcriptional start site (TSS) and polyadenylation site (Poly(A)), as determined by 5′ and 3′ RACE, respectively.
Tf2 antisense transcripts stabilized in the absence of Pab2-mediated RNA decay trigger RNAi-dependent repression of Tf2 expression
It was previously shown that the absence of Rrp6 in fission yeast promotes the production of Tf2-specific siRNAs, resulting in RNAi-dependent heterochromatin formation and transcriptional repression at Tf2 (Yamanaka et al. 2013). We therefore suspected that the accumulation of endogenous Tf2AS transcripts caused by the deletion of rrp6 and pab2 in the abp1Δ mutant (Figure 4) could be responsible for the attenuation of Tf2 transcription in cis. As deletion of genes encoding RNAi components (dcr1, ago1, or rdp1) in abp1Δ/pab2Δ or abp1Δ/rrp6Δ genetic backgrounds resulted in synthetic lethality, it was impossible to test whether the restoration of transcriptional repression in abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants was due to RNAi-dependent gene silencing. To determine whether Tf2AS transcripts could promote Tf2 repression in trans, we exogenously expressed Tf2AS transcripts under the control of the adh1 promoter using a chromosomally integrated construct. Exogenous Tf2AS was unable to silence Tf2 expression in trans in the context of the strong Tf2 derepression observed in either the abp1Δ single mutant or the abp1Δ/pab2Δ double mutant (Figure S2 and data not shown). Interestingly, however, we found that exogenous Tf2AS suppressed the accumulation of Tf2 mRNA detected in the pab2Δ single mutant (Figure 5A, lanes 3–4; quantification in Figure 5B). Notably, the levels of exogenous Tf2AS transcripts were increased by ∼fourfold in the pab2Δ mutant compared to wild-type cells (Figure 5A, compare lanes 2 and 4; Figure 5C, lanes 2 and 5; Figure 5D). The increase in the levels of exogenous Tf2AS RNA in the pab2Δ mutant was specific to Tf2AS, as the expression of gfp using the same expression construct did not result in increased levels of gfp mRNA in the pab2Δ mutant relative to wild-type cells (Figure 5C, compare lanes 3 and 6; quantification Figure 5D).
Figure 5.
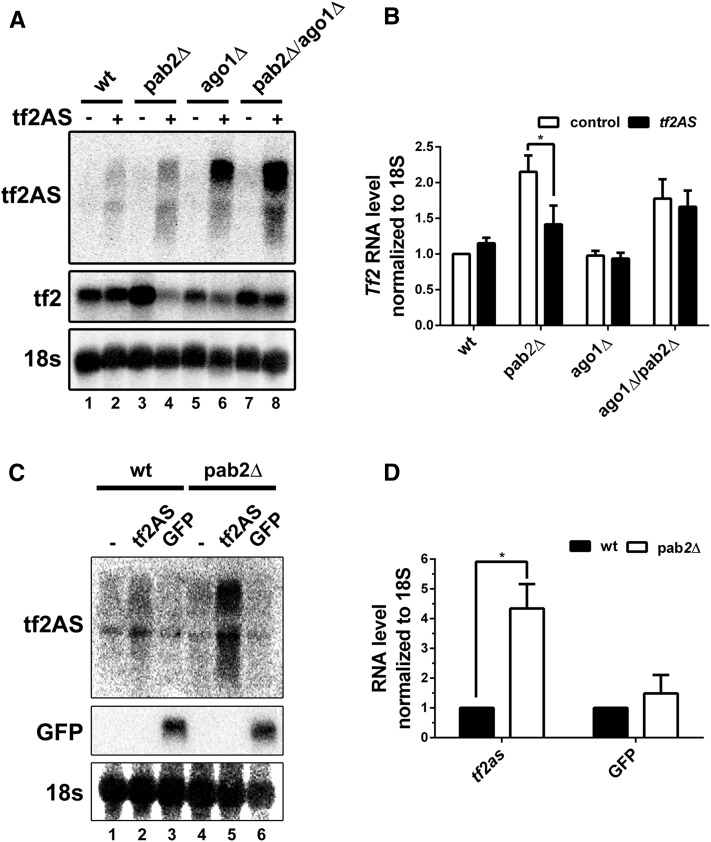
Ago1-dependent repression of Tf2 mRNA up-regulation in the pab2Δ mutant by exogenous expression of antisense Tf2 RNAs. (A) Northern blot analysis of total RNA prepared from wild-type (lanes 1–2), pab2Δ (lanes 3–4), ago1Δ (lanes 5–6), and pab2Δ/ago1Δ (lanes 7–8) cells expressing exogenous Tf2AS in trans (lanes 2, 4, 6, and 8) or transformed with the empty vector (lanes 1, 3, 5, and 7). The blot was hybridized using probes complementary to Tf2 antisense RNA, Tf2 mRNA, and 18s rRNA. (B) Quantification of Northern blots data for Tf2 mRNA levels. The data and error bars represent the average and SEM from at least three independent experiments. * P < 0.05; two-way ANOVA including Sidak’s multiple comparison test. (C) Northern blot analysis of total RNA prepared from wild-type (lanes 1–3) and pab2Δ (lanes 4–6) cells expressing exogenous Tf2AS RNA (lanes 2 and 5), gfp mRNA (lanes 3 and 6), or transformed with the empty vector (lanes 1 and 4). The blot was hybridized using probes complementary to Tf2 antisense RNA, gfp mRNA, and 18s rRNA. (D) Quantification of Northern blots data for Tf2 antisense RNA and gfp mRNA levels. The data and error bars represent the average and SEM from at least three independent experiments. * P < 0.05; two-way ANOVA including Sidak’s multiple comparison test.
The aforementioned data suggested that the absence of Pab2 results in Tf2AS stabilization, which could induce RNAi-dependent Tf2 repression in trans. To test whether the repressive effect of exogenous Tf2AS on Tf2 mRNA levels in the pab2Δ mutant required RNA interference, we deleted ago1, which encodes an essential RNAi factor. Notably, we found that the overall accumulation of exogenous Tf2AS in cells deleted for ago1 was greater than in Ago1-containing cells (Figure 5A, compare lanes 6 and 8 to 2 and 4). Yet, despite greater levels of Tf2AS in the absence of Ago1, the repressive effect of exogenous Tf2AS expression in the pab2Δ mutant was lost in the double pab2Δ/ago1Δ mutant (Figure 5A, lanes 7–8; Figure 5B). Similarly, the high levels of Tf2AS transcripts in the single ago1Δ mutant did not affect Tf2 mRNA levels (Figure 5A, lanes 5–6; quantifications in Figure 5B), suggesting that RNAi is required for Tf2 repression by exogenous Tf2AS RNA. Taken together, these results support a model in which the accumulation of Tf2AS transcripts in the absence of Pab2-dependent RNA decay activates an RNAi-mediated mechanism of Tf2 repression.
Tf2 antisense transcripts are induced during specific stages of meiotic differentiation
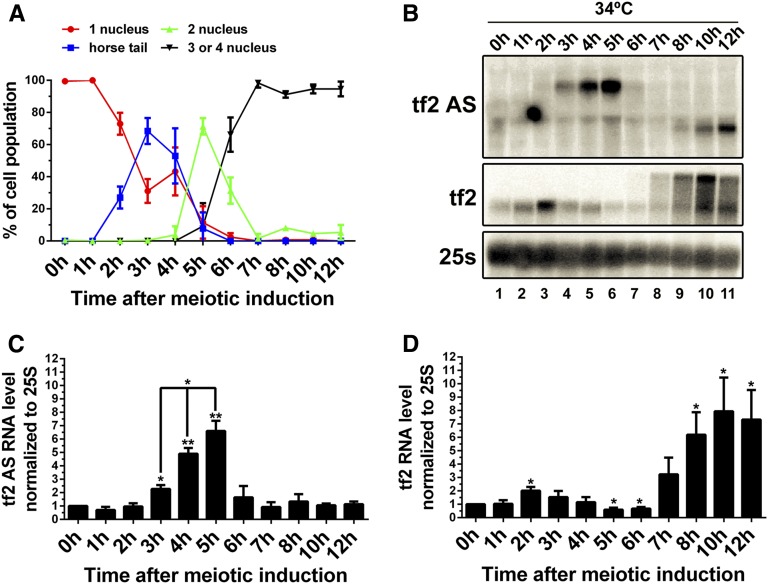
In S. pombe, waves of sense and antisense transcription accompany the meiotic differentiation program (Mata et al. 2002; Bitton et al. 2011). We thus examined the profile of Tf2 antisense transcript accumulation during the meiotic differentiation program by using diploid h+/h+ pat1-114/pat1-114 temperature-sensitive cells (Bähler et al. 1991), which allow the synchronization of sexual differentiation via inactivation of the Pat1 kinase (Figure 6A). Whereas the expression of Tf2AS transcripts was largely repressed during vegetative growth (Figure 6B, lane 1), Northern blot assays revealed a significant accumulation of Tf2AS RNA between 3 and 5 hr, with a peak at 5 hr, after the induction of meiosis (Figure 6B, lanes 4–6; quantification Figure 6C). The induction of Tf2AS RNA accumulation coincided with the horsetail and meiosis I stages of sexual differentiation (Figure 6A). A shorter form of Tf2 antisense transcript was also detected between 8 and 12 hr, peaking 12 hr after meiotic induction, which coincided with late stages of meiosis (Figure 6B, lanes 9–11). This short Tf2AS RNA was ∼790-nt-long with 5′ and 3′ ends mapping to nucleotides 4335 and 3546 of the Tf2 element, respectively, as determined by RACE experiments. Northern blot analysis also revealed accumulation of Tf2 mRNA at 1–2 and 8–12 hr after meiotic induction (Figure 6B, lanes 2–3 and 9–11). Interestingly, longer Tf2 transcripts were expressed in late stages of meiotic differentiation, 8–12 hr after meiotic induction (Figure 6B, lanes 8–11). Although we have not investigated whether the longer Tf2 RNAs accumulating in late meiosis had differential transcription start site as previously reported (Persson et al. 2016), RT-PCR analysis indicated 3′ extensions at different Tf2 loci suggestive of transcriptional read-through (data not shown). Tf2 mRNA expression was significantly repressed at 5 and 6 hr after meiotic induction (Figure 6D), and an inverse correlation (r = −0.984; P < 0.05) was established between Tf2 mRNA and Tf2AS expression during the early phase (2–5 hr) of meiotic differentiation (Figure 6, B–D). These results indicate that Tf2AS transcripts are expressed during specific stages of meiotic differentiation and contribute to Tf2 repression during the meiotic differentiation program.
Figure 6.
The expression of antisense Tf2 transcripts is developmentally regulated. (A) Meiosis progression in synchronized diploid h+/h+ pat1-114/pat1-114 cells. The data and error bars represent the average and SD from three independent experiments. At least 100 cells were counted by fluorescence microscopy for each time point. (B) Northern blot analysis of total RNA prepared from synchronized diploid h+/h+ pat1-114/pat1-114 cells at each hour from 0 to 8 hr (lanes 1–9), 10 hr (lane 10), and 12 hr (lane 11) after meiotic induction. The blot was hybridized using probes complementary to Tf2 antisense RNA, Tf2 mRNA, and 25s rRNA. The asterisk indicates a shorter antisense transcript mapping to the 3′ end of the Tf2 ORF. (C and D) Quantification of Northern blots data for antisense Tf2 RNA (C) and the Tf2 mRNA (D) levels. The data and error bars represent the average and SD from at least three independent experiments. * P < 0.05, ** P < 0.01; Student’s t-test.
Discussion
Although the nuclear poly(A)-binding protein Pab2 is known to contribute to Rrp6-dependent RNA degradation, the contribution of Pab2 to Tf2 regulation had remained elusive. In this study, we disclose a role for Pab2 in the repression of Tf2 mRNA expression. Our results also reveal that Pab2 and Abp1 function in redundant pathways that repress the accumulation of antisense Tf2 transcripts, which have the capacity to promote Tf2 repression via an RNAi-dependent mechanism. Our findings are consistent with the view that multiple transcriptional and post-transcriptional pathways collaborate to negatively control sense and antisense RNAs expressed from Tf2 retroelements.
Pab2 represses the expression of sense and antisense Tf2 transcripts
Analysis of gene expression changes at the genome-wide level in abp1-null cells indicates that Tf2 retroelements are transcribed in both the sense and antisense orientations (Lorenz et al. 2012). Interestingly, the data presented in this study reveal that Pab2 also contributes to the negative regulation of both sense Tf2 mRNA and antisense Tf2 transcripts. As levels of RNAPII at Tf2 loci were similar between wild-type and pab2Δ mutant cells (Figure 2D), the mechanism underlying Pab2-dependent control of Tf2 expression is likely to be at the post-transcriptional level. Consistent with this, Pab2 was shown to function as a key cofactor for exosome-mediated RNA decay of selective transcripts (St-Andre et al. 2010; Yamanaka et al. 2010; Chen et al. 2011; Lemieux et al. 2011). Accordingly, we found that cells deleted for rrp6, which encodes an exosome-associated 3′–5′ exonuclease, showed a slight increase in Tf2 mRNA as well as accumulated antisense Tf2 transcripts, consistent with previous data (Lee et al. 2013; Yamanaka et al. 2013) supporting that the nuclear exosome limits the accumulation of mRNA expressed from S. pombe Tf2 elements. Importantly, our results go beyond these findings by establishing the role of Pab2 in the post-transcriptional regulation of Tf2 expression, suggesting a mechanism that involves elimination of Pab2-bound polyadenylated Tf2 RNAs by the nuclear exosome (Figure 7). Although the detailed mechanism underlying the specificity of Pab2-dependent RNA turnover remains to be determined, the presence of cryptic, suboptimal introns in Tf2 (Lee et al. 2013) could prolong the nuclear residency of unspliced Tf2 transcripts, thereby increasing their susceptibility to Pab2-dependent pre-mRNA decay (Lemieux et al. 2011).
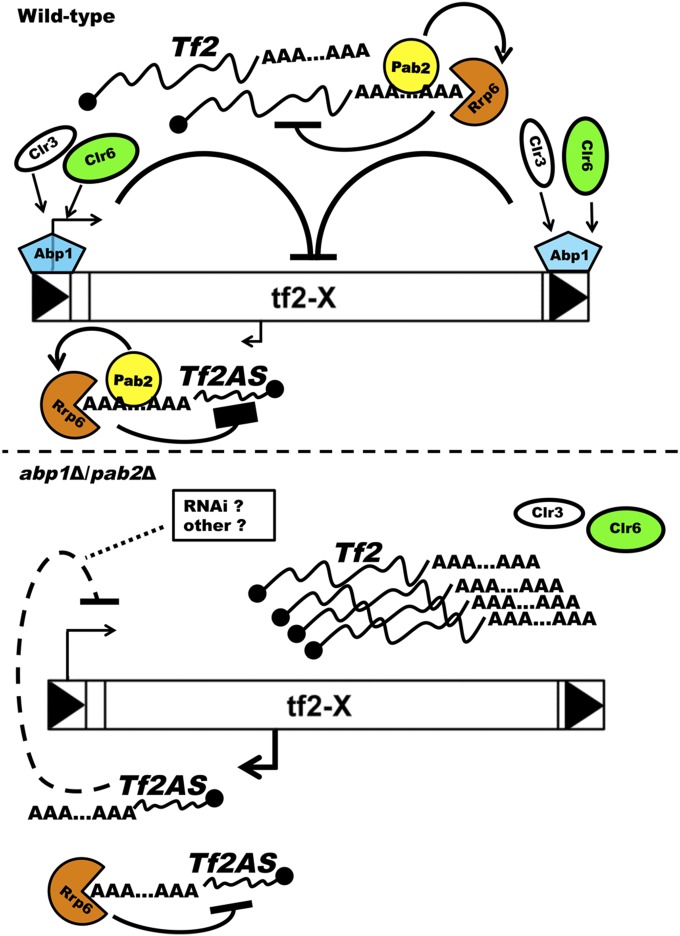
Figure 7.
Regulation of Tf2 expression by Pab2 and Abpl in fission yeast. Suggested model for how the absence of Pab2/Rrp6 attenuates the derepression of Tf2 transcription detected in the abplΔ single mutant. In wild-type cells (top panel), Abpl facilitates the recruitment of histone deacetylases such as Clr3 and Clr6 at Tf2 loci, repressing transcriptional activity at LTRs and Tf2 internal cryptic promoters. Concurrently, Pab2 collaborates with Rrp6 to promote degradation of Tf2 mRNA and antisense (Tf2AS) transcripts at the site of transcription. In the abplΔ/pab2Δ double mutant (bottom panel), the absence of Abp1 allows increased transcriptional activity from LTRs and internal cryptic promoters, whereas the absence of Pab2 reduces the capacity of Rrp6 to degrade Tf2 mRNA and Tf2AS transcripts. The accumulating levels Tf2AS at the site of transcription may act in cis to promote Tf2 repression via RNAi or redundant mechanisms that prevent the accumulation of dsRNA.
Pab2/Rrp6 and Abp1 function in overlapping pathways to repress the accumulation of Tf2 antisense transcripts
Given the established role of Abp1 in Tf2 transcriptional silencing (Cam et al. 2008; Lorenz et al. 2012) and the post-transcriptional control of Tf2 mRNA accumulation by Pab2/Rrp6-mediated RNA elimination (this study and Yamanaka et al. 2013), we expected an additive accumulation of Tf2 mRNA in abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants. In contrast, we found that the absence of Pab2 and Rrp6 attenuated the derepression of Tf2 silencing observed in the abp1Δ mutant by the restoration of transcriptional repression at Tf2 loci (Figure 2). Given that the deletion of ago1 in the rrp6Δ mutant results in the derepression of Tf2 (Figure S3 and Yamanaka et al. 2013), this indicates that rrp6-null cells are dependent on the RNAi machinery for Tf2 silencing. On the basis of these observations, we tested for the presence of endogenous antisense Tf2 transcripts that could activate the RNAi pathway via the formation of double-stranded (ds) RNA. Notably, our data revealed greater levels of antisense Tf2 RNAs in the abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants compared to either single mutant (Figure 4), supporting a model in which Pab2/Rrp6-mediated RNA decay and Abp1-dependent transcriptional repression act redundantly to suppress the accumulation of antisense Tf2 transcripts (see Figure 7). The increased levels of Tf2AS RNA in abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants led to the possibility that dsRNA generation could trigger RNAi-dependent repression of Tf2 transcription (Figure 7), which would explain the attenuation of Tf2 mRNA accumulation observed after the deletion of pab2 or rrp6 in the abp1Δ mutant (Figure 2). However, it was impossible to test whether the restoration of transcriptional repression in abp1Δ/pab2Δ and abp1Δ/rrp6Δ double mutants was due to RNAi, as the deletion of genes encoding RNAi components in abp1Δ/pab2Δ or abp1Δ/rrp6Δ backgrounds was lethal. The fact that the expression of exogenous Tf2AS in trans was unable to attenuate the Tf2 derepression detected in the absence of Abp1 (Figure S2) suggests that Pab2/Rrp6 promote the degradation of endogenous Tf2AS at the site of transcription and that endogenous Tf2AS mainly acts in cis to induce Tf2 transcriptional repression. Consistent with this view, RNAi-dependent gene silencing primarily acts in cis in S. pombe (Buhler et al. 2006). Interestingly, we found that expression of exogenous Tf2AS in trans suppressed the accumulation of Tf2 mRNA in pab2Δ cells, and that this effect was dependent on the presence of Ago1 (Figure 5, A and B). Although the detailed mechanism by which exogenous Tf2AS attenuated Tf2 mRNA accumulation by acting in trans in the pab2Δ single mutant remains to be determined, this repressive effect could potentially involve post-transcriptional gene regulation by the RNAi machinery, which was previously reported in S. pombe (Smialowska et al. 2014).
The absence of Pab2 reduces Tf2 mobilization in abp1-null cells
The deletion of S. pombe abp1 is known to relieve silencing and clustering at Tf2 loci, which allows Tf1 retroelements to recombine with preexisting Tf2 (Cam et al. 2008) and causes increased levels of Tf2 mobilization (Murton et al. 2016). Consistent with these findings, our data have shown that the increased in RNAPII occupancy at Tf2 in the absence of Abp1 (Figure 2D) results in Tf2 mRNA accumulation (Figure 2A) and greater mobilization (Figure 3). In contrast, our analyses did not find increased levels of Tf2 mobilization in the pab2Δ mutant despite Tf2 mRNA up-regulation. One possibility is that a twofold increase in Tf2 mRNA is not sufficient to cause a concomitant increase in Tf2 protein expression, as the Tf2 PR-RT fusion protein was not detected in the pab2Δ mutant (Figure 3). Alternatively, recent findings indicate that Tf2 mRNA accumulation is not sufficient to cause an increase in mobilization, since the structural organization of Tf2 into clusters (referred to as Tf bodies) restricts retrotransposition (Murton et al. 2016). Accordingly, the organization of Tf bodies is unlikely to be impaired by the absence of Pab2, thereby restricting the increased number of Tf2 mRNA molecules to induce mobilization by homologous recombination. In the absence of Abp1, however, which results in declustering of Tf2 elements (Cam et al. 2008; Lorenz et al. 2012), we found that Tf2 mRNA levels correlated with mobilization frequency. Specifically, we found that the attenuation of Tf2 mRNA accumulation induced by the deletion of pab2 in the abp1Δ mutant resulted in a significant reduction in the frequency of Tf2-12 mobilization (Figure 3). Although it was previously reported that expression of antisense transcripts to S. cerevisiae Ty1 retrotransposons induces defects in retrotransposon protein maturation that lead to a decrease in mobilization frequency (Matsuda and Garfinkel 2009), we have found no evidence of PR-RT maturation defects in the abp1Δ/pab2Δ strain (data not shown); however, there was a decrease in the level of PR-RT protein (Figure 3, E and F) that correlated with lower Tf2 mRNA levels (Figure 3, C and D). Yet, the possibility that RNAi-dependent heterochromatin formation at Tf2 in the abp1Δ/pab2Δ double mutant impairs homologous recombination, thereby restricting Tf2 mobilization, cannot be excluded.
The expression of antisense Tf2 transcripts is developmentally regulated
Whereas siRNAs mapping to Tf2 elements are not readily detected in haploid S. pombe grown in optimal conditions (Hansen et al. 2005; Woolcock et al. 2011), Tf2-specific siRNAs have been detected after induction of meiotic differentiation (Yamanaka et al. 2013). Consistent with the detection of siRNAs mapping to Tf2 during meiosis, we show here that antisense Tf2 transcripts are detected at specific stages of the fission yeast sexual differentiation program (Figure 6). Specifically, we found maximal expression of Tf2AS transcripts during the horsetail and meiosis I stages of the meiotic division cycle (Figure 6, A and B). During the horsetail-stage nuclei of S. pombe meiosis, homologous chromosomes align and recombination takes place (Wells et al. 2006). It is therefore reasonable to suggest that there is declustering of Tf bodies during this stage of meiosis, providing a window of opportunity to express sense and antisense Tf2 transcripts, and produce siRNAs. Interestingly, we observed a negative correlation between the expression of Tf2AS transcripts and Tf2 mRNA during the horsetail and meiosis I stages (Figure 6). However, siRNA production may not necessarily be the functional outcome of Tf2AS expression during meiosis, as an inverse correlation was observed between the expression of Tf2 mRNA and Tf2AS in the middle phase (6–8 hr) of meiotic differentiation in the ago1Δ mutant (Figure S4), raising the possibility that a mechanism of Tf2AS-dependent repression of Tf2 expression during meiosis may not require RNAi. Although future studies will be required to define the functional role of Tf2 antisense RNAs during meiosis, it remains tempting to speculate about the use of Tf2AS during meiosis to prevent DNA double-strand breaks and minimize recombination between different Tf2 loci during the cross-over stage of meiosis, which coincided with the accumulation of Tf2AS. Accordingly, RNAi-dependent heterochromatin formation was shown to suppress DNA double-strand breaks and recombination frequency at centromeres during fission yeast meiosis (Ellermeier et al. 2010).
In summary, our study suggests that an RNA elimination pathway that involves Pab2 and the RNA exosome controls Tf2 mRNAs. Furthermore, our findings indicate that Pab2/Rrp6-dependent RNA decay and Abp1-mediated transcriptional silencing function redundantly to negatively regulate the expression of antisense Tf2 RNAs, which have the capacity to suppress Tf2 mRNA accumulation by a mechanism that appears to involve RNA interference. Given the importance of the 3′ poly(A) tail in the mobilization of human retrotransposons (Doucet et al. 2015), it will be interesting to examine whether the Pab2-dependent RNA elimination pathway of retrotransposon surveillance described in this study is conserved in humans.
Supplementary Material
Supplemental material is available online at www.genetics.org/lookup/suppl/doi:10.1534/genetics.116.193870/-/DC1.
Acknowledgments
We thank P. Espenshade for the neoAI-tagged Tf2-12 strain, S. Whitehall for the individually tagged ura4+ Tf2 strains, and H. Levin for the anti-RT antibody. We also thank Jean-François Lemay for the generation of some DNA constructs. This work was supported by a grant from the Canadian Institutes of Health Research (CIHR) to F.B. (MOP-106595). P.-L.M. was supported by a scholarship from the Fonds de la recherche du Québec – Santé. F.B. is the recipient of a Canada Research Chair.
Footnotes
Communicating editor: A. Hinnebusch
Literature Cited
- Allmang C., Petfalski E., Podtelejnikov A., Mann M., Tollervey D., et al. , 1999. The yeast exosome and human PM-Scl are related complexes of 3′ → 5′ exonucleases. Genes Dev. 13: 2148–2158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anderson H. E., Wardle J., Korkut S. V., Murton H. E., Lopez-Maury L., et al. , 2009. The fission yeast HIRA histone chaperone is required for promoter silencing and the suppression of cryptic antisense transcripts. Mol. Cell. Biol. 29: 5158–5167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bähler J., Schuchert P., Grimm C., Kohli J., 1991. Synchronized meiosis and recombination in fission yeast: observations with pat1–114 diploid cells. Curr. Genet. 19: 445–451. [DOI] [PubMed] [Google Scholar]
- Bähler J., Wu J. Q., Longtine M. S., Shah N. G., McKenzie A., 3rd, et al. , 1998. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14: 943–951. [DOI] [PubMed] [Google Scholar]
- Beaudoin J., Labbe S., 2006. Copper induces cytoplasmic retention of fission yeast transcription factor cuf1. Eukaryot. Cell 5: 277–292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bitton D. A., Grallert A., Scutt P. J., Yates T., Li Y., et al. , 2011. Programmed fluctuations in sense/antisense transcript ratios drive sexual differentiation in S. pombe. Mol. Syst. Biol. 7: 559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowen N. J., Jordan I. K., Epstein J. A., Wood V., Levin H. L., 2003. Retrotransposons and their recognition of pol II promoters: a comprehensive survey of the transposable elements from the complete genome sequence of Schizosaccharomyces pombe. Genome Res. 13: 1984–1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buhler M., Verdel A., Moazed D., 2006. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125: 873–886. [DOI] [PubMed] [Google Scholar]
- Cam H. P., Sugiyama T., Chen E. S., Chen X., FitzGerald P. C., et al. , 2005. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37: 809–819. [DOI] [PubMed] [Google Scholar]
- Cam H. P., Noma K., Ebina H., Levin H. L., Grewal S. I., 2008. Host genome surveillance for retrotransposons by transposon-derived proteins. Nature 451: 431–436. [DOI] [PubMed] [Google Scholar]
- Chen H. M., Futcher B., Leatherwood J., 2011. The fission yeast RNA binding protein Mmi1 regulates meiotic genes by controlling intron specific splicing and polyadenylation coupled RNA turnover. PLoS One 6: e26804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cordaux R., Batzer M. A., 2009. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10: 691–703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doucet A. J., Wilusz J. E., Miyoshi T., Liu Y., Moran J. V., 2015. A 3′ poly(A) tract is required for LINE-1 retrotransposition. Mol. Cell 60: 728–741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dumesic P. A., Madhani H. D., 2014. Recognizing the enemy within: licensing RNA-guided genome defense. Trends Biochem. Sci. 39: 25–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Egan E. D., Braun C. R., Gygi S. P., Moazed D., 2014. Post-transcriptional regulation of meiotic genes by a nuclear RNA silencing complex. RNA 20: 867–881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellermeier C., Higuchi E. C., Phadnis N., Holm L., Geelhood J. L., et al. , 2010. RNAi and heterochromatin repress centromeric meiotic recombination. Proc. Natl. Acad. Sci. USA 107: 8701–8705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esnault C., Levin H. L., 2015. The long terminal repeat retrotransposons Tf1 and Tf2 of Schizosaccharomyces pombe. Microbiol. Spectr. 3(4). Available at: http://www.asmscience.org/content/journal/microbiolspec/10.1128/microbiolspec.MDNA3-0040-2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Girard A., Hannon G. J., 2008. Conserved themes in small-RNA-mediated transposon control. Trends Cell Biol. 18: 136–148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goto D. B., Nakayama J., 2012. RNA and epigenetic silencing: insight from fission yeast. Dev. Growth Differ. 54: 129–141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall B. M., Ma C. X., Liang P., Singh K. K., 2009. Fluctuation analysis CalculatOR: a web tool for the determination of mutation rate using Luria-Delbruck fluctuation analysis. Bioinformatics 25: 1564–1565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen K. R., Burns G., Mata J., Volpe T. A., Martienssen R. A., et al. , 2005. Global effects on gene expression in fission yeast by silencing and RNA interference machineries. Mol. Cell. Biol. 25: 590–601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoff E. F., Levin H. L., Boeke J. D., 1998. Schizosaccharomyces pombe retrotransposon Tf2 mobilizes primarily through homologous cDNA recombination. Mol. Cell. Biol. 18: 6839–6852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johansen P., Cam H. P., 2015. Suppression of meiotic recombination by CENP-B homologs in Schizosaccharomyces pombe. Genetics 201: 897–904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larochelle M., Lemay J. F., Bachand F., 2012. The THO complex cooperates with the nuclear RNA surveillance machinery to control small nucleolar RNA expression. Nucleic Acids Res. 40: 10240–10253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee N. N., Chalamcharla V. R., Reyes-Turcu F., Mehta S., Zofall M., et al. , 2013. Mtr4-like protein coordinates nuclear RNA processing for heterochromatin assembly and for telomere maintenance. Cell 155: 1061–1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemay J. F., D’Amours A., Lemieux C., Lackner D. H., St-Sauver V. G., et al. , 2010. The nuclear poly(A)-binding protein interacts with the exosome to promote synthesis of noncoding small nucleolar RNAs. Mol. Cell 37: 34–45. [DOI] [PubMed] [Google Scholar]
- Lemieux C., Marguerat S., Lafontaine J., Barbezier N., Bahler J., et al. , 2011. A Pre-mRNA degradation pathway that selectively targets intron-containing genes requires the nuclear poly(A)-binding protein. Mol. Cell 44: 108–119. [DOI] [PubMed] [Google Scholar]
- Lorenz D. R., Mikheyeva I. V., Johansen P., Meyer L., Berg A., et al. , 2012. CENP-B cooperates with Set1 in bidirectional transcriptional silencing and genome organization of retrotransposons. Mol. Cell. Biol. 32: 4215–4225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mata J., Lyne R., Burns G., Bahler J., 2002. The transcriptional program of meiosis and sporulation in fission yeast. Nat. Genet. 32: 143–147. [DOI] [PubMed] [Google Scholar]
- Matsuda E., Garfinkel D. J., 2009. Posttranslational interference of Ty1 retrotransposition by antisense RNAs. Proc. Natl. Acad. Sci. USA 106: 15657–15662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mita P., Boeke J. D., 2016. How retrotransposons shape genome regulation. Curr. Opin. Genet. Dev. 37: 90–100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murton H. E., Grady P. J., Chan T. H., Cam H. P., Whitehall S. K., 2016. Restriction of retrotransposon mobilization in Schizosaccharomyces pombe by transcriptional silencing and higher-order chromatin organization. Genetics 203: 1669–1678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Persson J., Steglich B., Smialowska A., Boyd M., Bornholdt J., et al. , 2016. Regulating retrotransposon activity through the use of alternative transcription start sites. EMBO Rep. 17: 753–768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhind N., Chen Z., Yassour M., Thompson D. A., Haas B. J., et al. , 2011. Comparative functional genomics of the fission yeasts. Science 332: 930–936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roguev A., Bandyopadhyay S., Zofall M., Zhang K., Fischer T., et al. , 2008. Conservation and rewiring of functional modules revealed by an epistasis map in fission yeast. Science 322: 405–410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sehgal A., Lee C. Y., Espenshade P. J., 2007. SREBP controls oxygen-dependent mobilization of retrotransposons in fission yeast. PLoS Genet. 3: e131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smialowska A., Djupedal I., Wang J., Kylsten P., Swoboda P., et al. , 2014. RNAi mediates post-transcriptional repression of gene expression in fission yeast Schizosaccharomyces pombe. Biochem. Biophys. Res. Commun. 444: 254–259. [DOI] [PubMed] [Google Scholar]
- St-Andre O., Lemieux C., Perreault A., Lackner D. H., Bahler J., et al. , 2010. Negative regulation of meiotic gene expression by the nuclear poly(a)-binding protein in fission yeast. J. Biol. Chem. 285: 27859–27868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verdel A., Jia S., Gerber S., Sugiyama T., Gygi S., et al. , 2004. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303: 672–676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Volpe T. A., Kidner C., Hall I. M., Teng G., Grewal S. I., et al. , 2002. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297: 1833–1837. [DOI] [PubMed] [Google Scholar]
- Wells J. L., Pryce D. W., McFarlane R. J., 2006. Homologous chromosome pairing in Schizosaccharomyces pombe. Yeast 23: 977–989. [DOI] [PubMed] [Google Scholar]
- Woolcock K. J., Gaidatzis D., Punga T., Buhler M., 2011. Dicer associates with chromatin to repress genome activity in Schizosaccharomyces pombe. Nat. Struct. Mol. Biol. 18: 94–99. [DOI] [PubMed] [Google Scholar]
- Yamanaka S., Yamashita A., Harigaya Y., Iwata R., Yamamoto M., 2010. Importance of polyadenylation in the selective elimination of meiotic mRNAs in growing S. pombe cells. EMBO J. 29: 2173–2181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamanaka S., Mehta S., Reyes-Turcu F. E., Zhuang F., Fuchs R. T., et al. , 2013. RNAi triggered by specialized machinery silences developmental genes and retrotransposons. Nature 493: 557–560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou Y., Zhu J., Schermann G., Ohle C., Bendrin K., et al. , 2015. The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome. Nat. Commun. 6: 7050. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Strains and plasmids are available upon request. Table S1 contains all strains used in this study.