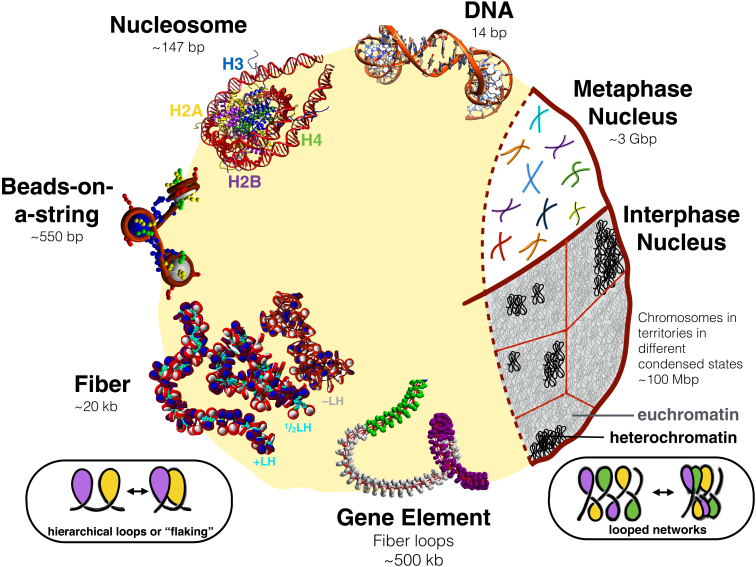
Figure 1.
A schematic view of the levels and structures involved as the chromatin fiber packs genomic DNA in the cell nucleus. Several levels of structural and functional states are known but not well understood. Double-stranded DNA binds to histone proteins to form complexes, known as nucleosomes, the building blocks of chromatin. Each nucleosome is composed of ∼147 bp of DNA coiled around eight histone proteins, two dimers each of H2A, H2B, H3, and H4. Many nucleosomes have been resolved at atomic resolution. Nucleosomes connect to one another to form the chromatin fiber, where an open state forms at low salt without linker histones called beads-on-a-string. A more condensed state forms at high salt with linker histones like H1 or H5. The long-assumed compact 30-nm fiber may adopt more variable forms in vivo, as shown for the subsaturated LH fibers (one LH per two nucleosomes, or ½LH) and LH-deficient (−LH) fibers shown in the bottom left. Relatively straight +LH fibers show little self-association, whereas ½LH fibers show moderate self-association, and −LH fibers show high levels of self-association. All three types show variable widths. Additionally, these self-associating fibers may involve hierarchical looping or flaking, as used in mountain climbing and rappelling, which is more pronounced in ½LH and −LH fibers. Flaking, in this context, refers to the lateral compaction of a long polymer due to loops of various size undergoing further folding in space, while avoiding tangles or knots. In the context of rope folding, flaking refers to packaging flexible elongated rope into neatly stacked loops (by winding the loops back and forth) so that the rope can be easily unraveled. A simple scheme for such rope stacking is shown at the bottom left. At bottom right, we show another possible variation of such compact networks of stacked loops. Active gene elements form large loose loops that bridge gene promoters to gene-transcribing regions. In cooperation with other epigenetic marks and scaffolding proteins, these gene elements organize into chromosomes. In interphase chromosomes, individual chromosomes are separated into different chromosomal territories. In the metaphase cell, individual chromosomes can be distinguished, as shown by different colors. Interphase chromosomes are also known to exist in different levels of condensation. The more loosely packed genomic state is known as “euchromatin”, while the more densely packed genomic material is denoted as “heterochromatin”. A possible internal organization for these two related states involves tight versus open networks of chromatin loops, which build upon the proposed flaking or hierarchical looping motif. To see this figure in color, go online.