Fig. 1.
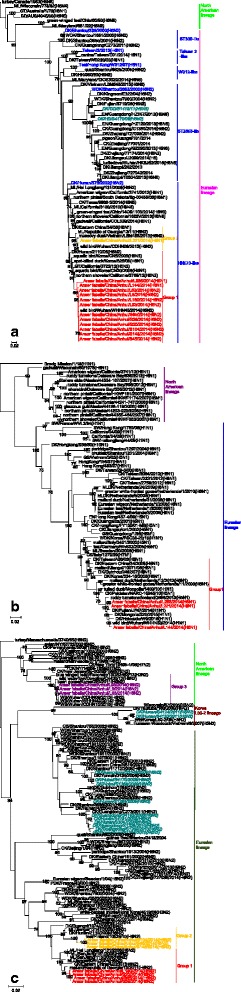
Phylogenetic analysis based on maximum likelihood of surface genes H6 (a), N1 (b) and N2 (c) of AIVs isolated in Anhui Province, 2014. Phylogenetic trees were generated with the MAGE 6.0 software package. The phylogenetic trees for HA (1A) were rooted to A/Turkey/Canada/1/1963 (H6N8). The phylogenetic tree for the N1 genes were rooted to A/Brevig_Mission/1/18(H1N1) and N2 genes were rooted to turkey/Massachussetts/3740/65(H6N2). The genomic sequences of the viruses listed in black and blue were downloaded from available databases; the viruses listed in red, yellow, and/or purple was sequenced in this study. The sequences shown in cyan and italics in Fig 1c were obtained from a paper by Wang GJ. Abbreviations: CK, chicken; DK, duck; GS, goose; SW, swine; ML, mallard; GT, green-winged teal; PD, Pacific black duck; WDK, wild duck; MLDK, mallard duck; WB, wild bird; EN, environment
