Fig. 7.
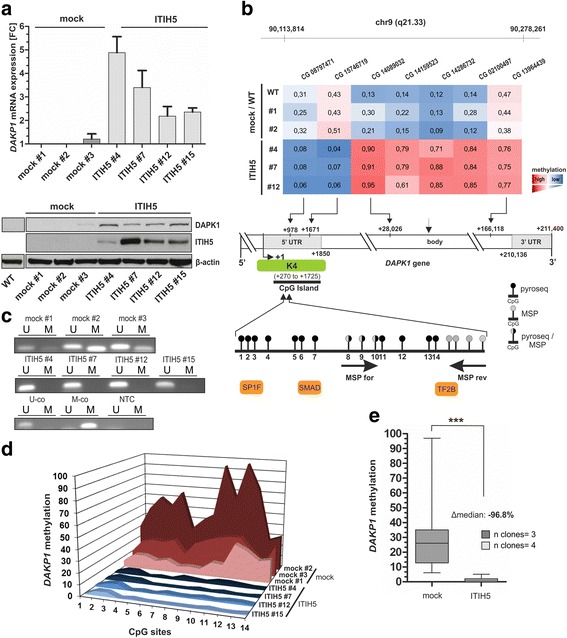
ITIH5 mediates demethylation of the DAPK1 promoter 5’UTR region leading to its re-expression in basal-type breast cancer cells. a DAPK1 re-expression was confirmed in ITIH5 clones (n = 4) by real-time PCR (upper graph) and western blot analysis (lower images) when compared with and mock clones (n = 3) and MDA-MB-231 WT. β-actin served as loading control. b Schematic map of the human DAPK1 gene including the relative positions and β-values of CpG dinucleotides measured by 450 K methylation array profiling in MDA-MB-231 WT cells, mock and ITIH5 single-cell clones. Red: high methylation, blue: low methylation. +1: DAPK1 transcription start site (TSS). A predicted CpG island is located between genomic positions 90,112,413 and 90,114,138 (+270 bp to +1725 bp relative to the expected TSS) within the 5’-UTR region. At this site a potential activating H3K4Me3 (K4) histone modification as mapped by Ku et al. [41] was described. The relative positions of 18 CpG sites analyzed either by MSP (used primer: black arrows) and / or pyrosequencing within the DAPK1 5’UTR region are indicated. Three putative transcription binding sites in this gene locus were statistically identified: SP1F (matrix similarity: 0.941), SMAD (matrix similarity: 0.963) and TF2B (matrix similarity: 1.0). c DNA methylation of the DAPK1 5’UTR locus verified in mock and ITIH5 single-cell clones by using MSP. Band labels with U and M represent an unmethylated and methylated DNA region, respectively. Bisulfite-converted unmethylated, (U-co) and polymethylated, genomic (M-co) DNA were used as controls. NTC: non-template control. d-e Quantification of DAPK1 5’UTR DNA methylation frequency by using pyrosequencing. d 3D graph illustrates methylation level for each analyzed CpG site (overall 14 CpGs) within the DAPK1 5’UTR locus in mock (n = 3) and ITIH5 (n = 4) single-cell clones. e Box plot analysis demonstrates significant reduction of the median methylation ratio within the DAPK1 5’UTR region in ΔpBK-ITIH5 compared to ΔpBK-mock clones. Horizontal lines: grouped medians. Boxes: 25–75% quartiles. Vertical lines: range, peak and minimum; *p < 0.05, **p < 0.01, ***p < 0.001
