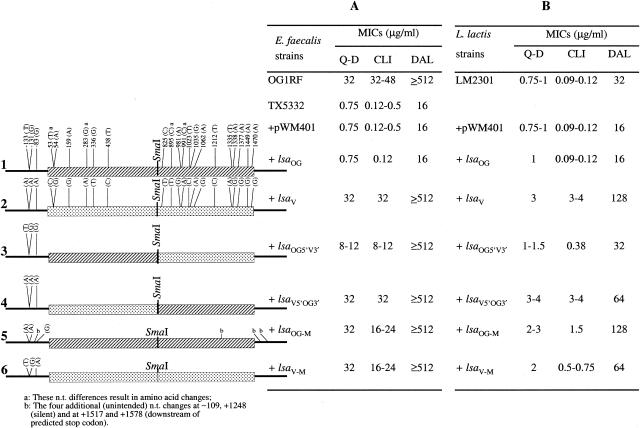
FIG. 1.
Susceptibility of E. faecalis and L. lactis derivatives to Q-D, DAL, and CLI. Rows 1 to 6 show the lsa gene constructs and nucleotide differences of lsaOG (ca. 2-kb lsa from OG1RF), lsaV (ca. 2-kb lsa from V583), lsaOG5′V3′ (hybrid lsa with 5′ half of OG1RF and 3′ half of V583), lsaV5′OG3′ (hybrid lsa with 5′ half of V583 and 3′ half of OG1RF), lsaOG-M (mutagenized lsaOG at positions −131 and −133 where G and T were replaced with A's [and four unintended changes were also found]), and lsaV-M (mutagenized lsaV at positions −131 and −133 where A's were replaced with G and T, respectively) cloned into pWM401 and their influence on susceptibility of TX5332 (E. faecalis OG1 lsa disruption mutant) (A) and L. lactis LM2301 (B) to Q-D, DAL, and CLI. n.t., nucleotide.